The Molecular Biology of Genes and Gene Expression
The Molecular Biology of Genes and Gene Expression

Central Dogma • First described by Francis Crick • Information only flows from DNA → RNA → protein • Transcription = DNA → RNA • Translation = RNA → protein • Retroviruses violate this order using reverse transcriptase to convert their RNA genome into DNA 2
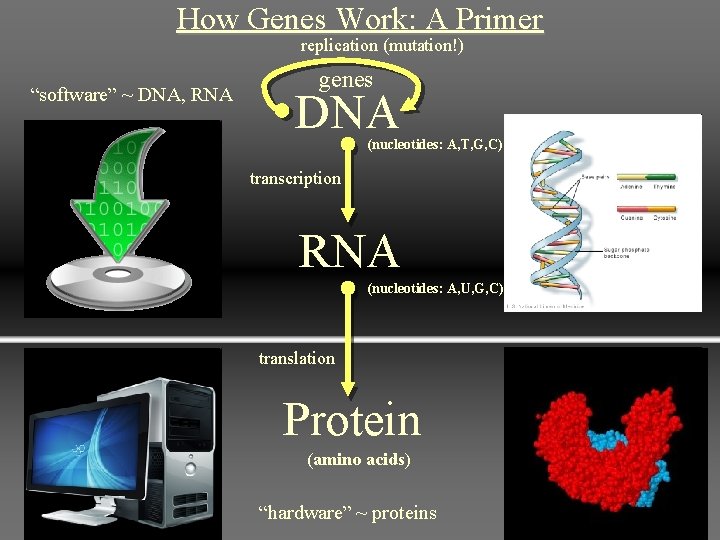
How Genes Work: A Primer replication (mutation!) “software” ~ DNA, RNA genes DNA (nucleotides: A, T, G, C) transcription RNA (nucleotides: A, U, G, C) translation Protein (amino acids) “hardware” ~ proteins

The Nature of Genes • Early ideas to explain how genes work came from studying human diseases • Archibald Garrod – 1902 – Recognized that alkaptonuria is inherited via a recessive allele – Proposed that patients with the disease lacked a particular enzyme • These ideas connected genes to enzymes 4

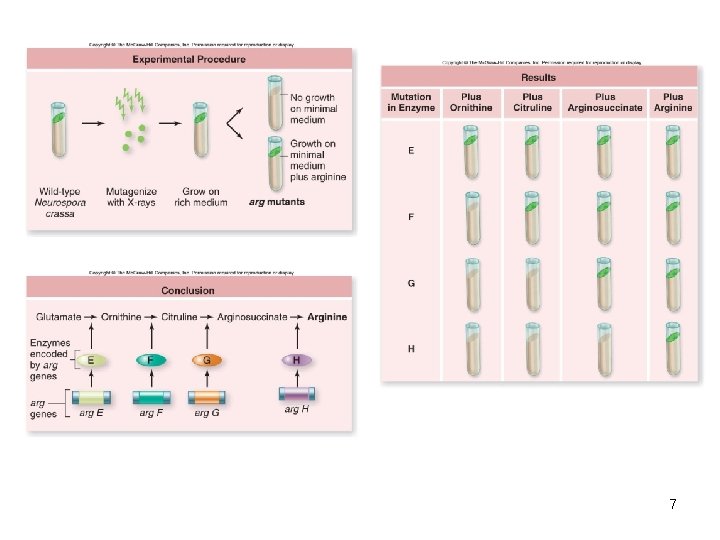
Beadle and Tatum – 1941 • Deliberately set out to create mutations in chromosomes and verify that they behaved in a Mendelian fashion in crosses • Studied Neurospora crassa – Used X-rays to damage DNA – Looked for nutritional mutations • Had to have minimal media supplemented to grow 5

• Beadle and Tatum looked for fungal cells lacking specific enzymes – The enzymes were required for the biochemical pathway producing the amino acid arginine – They identified mutants deficient in each enzyme of the pathway • One-gene/one-enzyme hypothesis has been modified to one-gene/one-polypeptide hypothesis 6
7
8

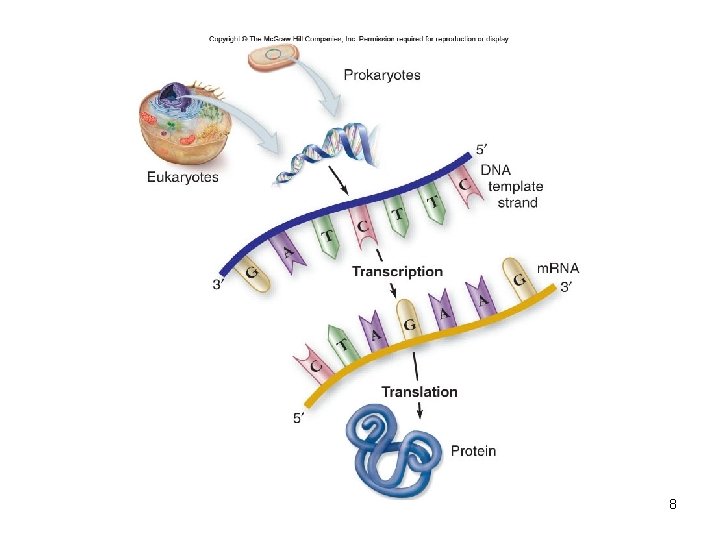
• Transcription – – DNA-directed synthesis of RNA Only template strand of DNA used U (uracil) in DNA replaced by T (thymine) in RNA m. RNA used to direct synthesis of polypeptides • Translation – Synthesis of polypeptides – Takes place at ribosome – Requires several kinds of RNA 9

RNA • All synthesized from DNA template by transcription • Messenger RNA (m. RNA) • Ribosomal RNA (r. RNA) • Transfer RNA (t. RNA) • Small nuclear RNA (sn. RNA) • Signal recognition particle RNA • Micro-RNA (mi. RNA) 10

Genetic Code • Francis Crick and Sydney Brenner determined how the order of nucleotides in DNA encoded amino acid order • Codon – block of 3 DNA nucleotides corresponding to an amino acid • Introduced single nulcleotide insertions or deletions and looked for mutations – Frameshift mutations • Indicates importance of reading frame 11
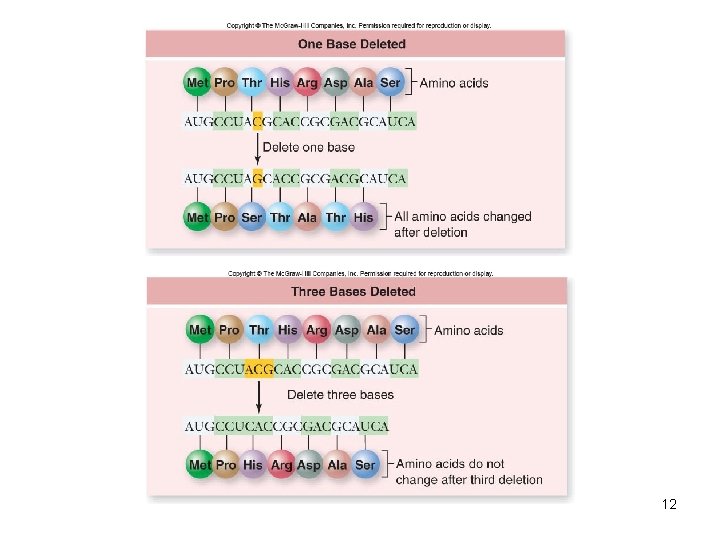
12

• Marshall Nirenberg identified the codons that specify each amino acid • Stop codons – 3 codons (UUA, UGA, UAG) used to terminate translation • Start codon – Codon (AUG) used to signify the start of translation • Code is degenerate, meaning that some amino acids are specified by more than one codon 13
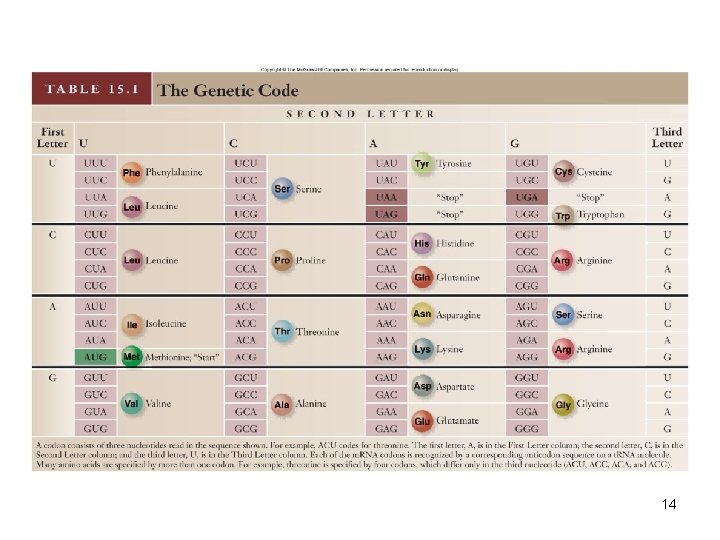
14

Code practically universal • Strongest evidence that all living things share common ancestry • Advances in genetic engineering • Mitochondria and chloroplasts have some differences in “stop” signals 15
Prokaryotic transcription • Single RNA polymerase • Initiation of m. RNA synthesis does not require a primer • Requires – Promoter – Start site – Termination site Transcription unit 16

• Promoter – Forms a recognition and binding site for the RNA polymerase – Found upstream of the start site – Not transcribed – Asymmetrical – indicate site of initiation and direction of transcription 17
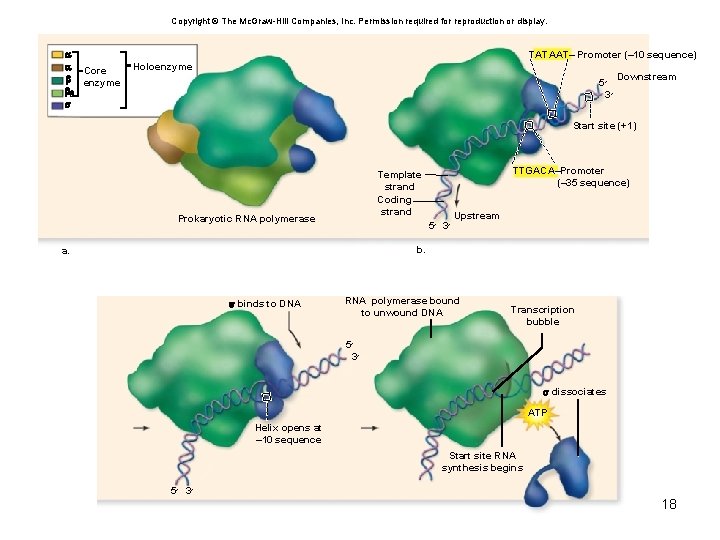
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Core enzyme 9 TATAAT– Promoter (– 10 sequence) Holoenzyme 5 ׳ 3 ׳ Downstream Start site (+1) TTGACA–Promoter (– 35 sequence) Template strand Coding strand Prokaryotic RNA polymerase 5 ׳ 3 ׳ Upstream b. a. binds to DNA RNA polymerase bound to unwound DNA Transcription bubble 5 ׳ 3 ׳ dissociates ATP Helix opens at – 10 sequence Start site RNA synthesis begins 5 ׳ 3 ׳ 18

• Elongation – Grows in the 5′-to-3′ direction as ribonucleotides are added – Transcription bubble – contains RNA polymerase, DNA template, and growing RNA transcript – After the transcription bubble passes, the nowtranscribed DNA is rewound as it leaves the bubble 19
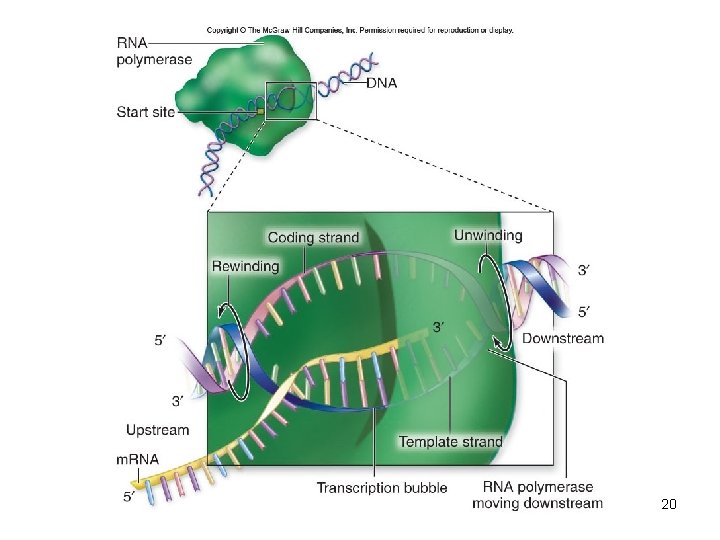
20

• Termination – Marked by sequence that signals “stop” to polymerase • Causes the formation of phosphodiester bonds to cease • RNA–DNA hybrid within the transcription bubble dissociates • RNA polymerase releases the DNA • DNA rewinds – Hairpin 21
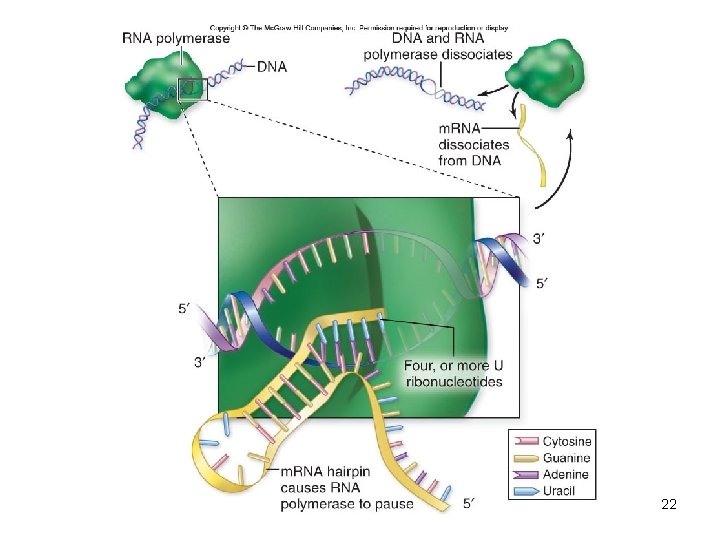
22

• Prokaryotic transcription is coupled to translation – m. RNA begins to be translated before transcription is finished – Operon • Grouping of functionally related genes • Multiple enzymes for a pathway • Can be regulated together 23
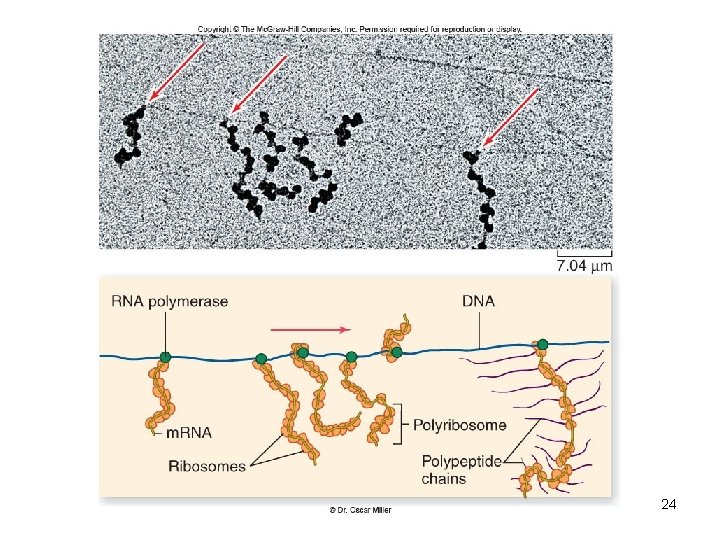
24

Eukaryotic Transcription • 3 different RNA polymerases – RNA polymerase I transcribes r. RNA – RNA polymerase II transcribes m. RNA and some sn. RNA – RNA polymerase III transcribes t. RNA and some other small RNAs • Each RNA polymerase recognizes its own promoter 25

• Initiation of transcription – Requires a series of transcription factors • Necessary to get the RNA polymerase II enzyme to a promoter and to initiate gene expression • Interact with RNA polymerase to form initiation complex at promoter • Termination – Termination sites not as well defined 26
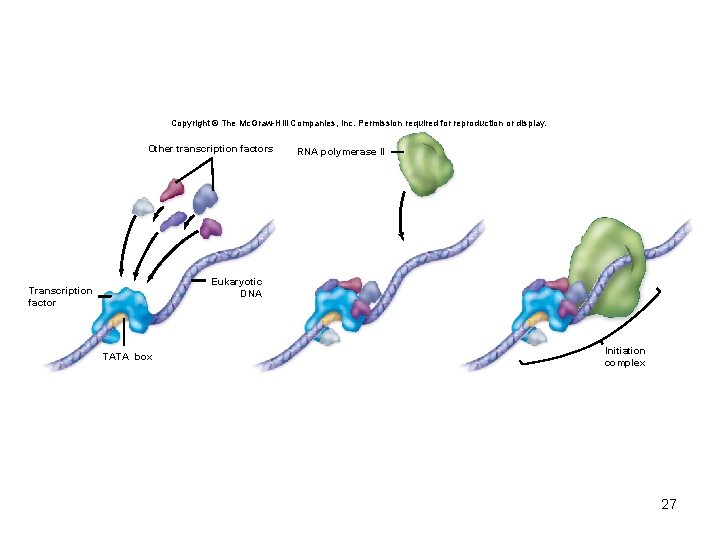
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Other transcription factors RNA polymerase II Eukaryotic DNA Transcription factor TATA box Initiation complex 27
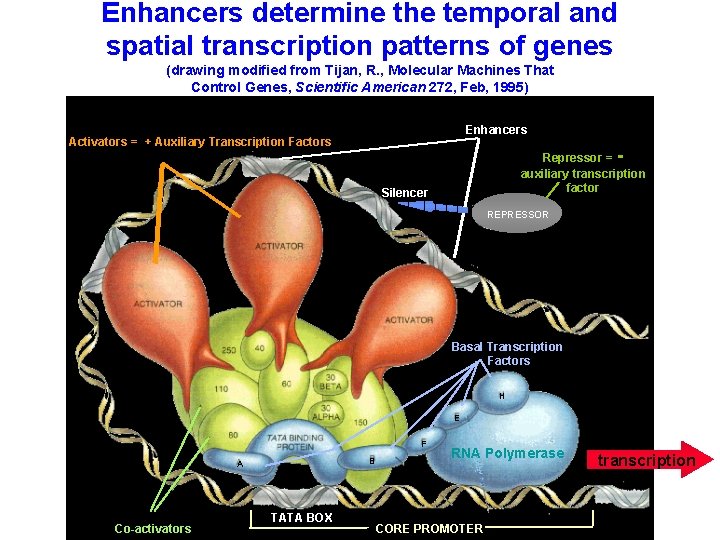
Enhancers determine the temporal and spatial transcription patterns of genes (drawing modified from Tijan, R. , Molecular Machines That Control Genes, Scientific American 272, Feb, 1995) Enhancers Activators = + Auxiliary Transcription Factors - Repressor = auxiliary transcription factor Silencer REPRESSOR Basal Transcription Factors H E F B A Co-activators TATA BOX RNA Polymerase CORE PROMOTER transcription

m. RNA modifications • In eukaryotes, the primary transcript must be modified to become mature m. RNA – Addition of a 5′ cap • Protects from degradation; involved in translation initiation – Addition of a 3′ poly-A tail • Created by poly-A polymerase; protection from degradation – Removal of non-coding sequences (introns) • Pre-m. RNA splicing done by spliceosome 29
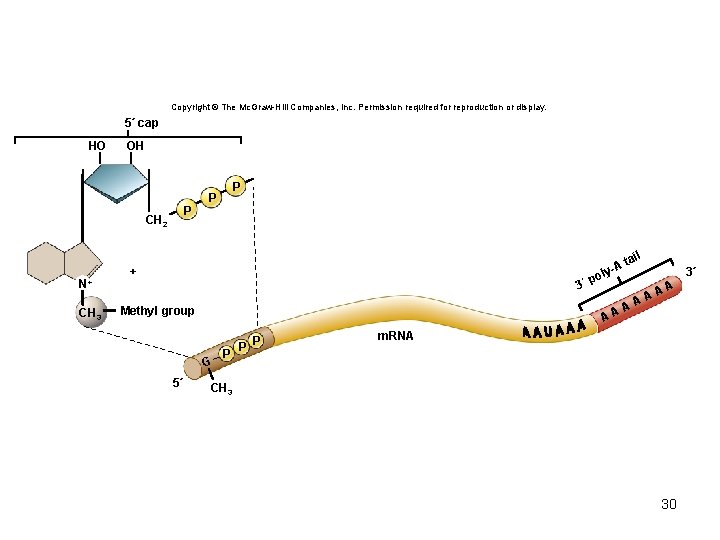
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. 5´ cap HO OH P CH 2 N+ CH 3 P P + 3´ Methyl group P P P 3´ l po A G 5´ l tai A y- A AA m. RNA CH 3 30

Eukaryotic pre-m. RNA splicing • Introns – non-coding sequences • Exons – sequences that will be translated • Small ribonucleoprotein particles (sn. RNPs) recognize the intron–exon boundaries • sn. RNPs cluster with other proteins to form spliceosome – Responsible for removing introns 31
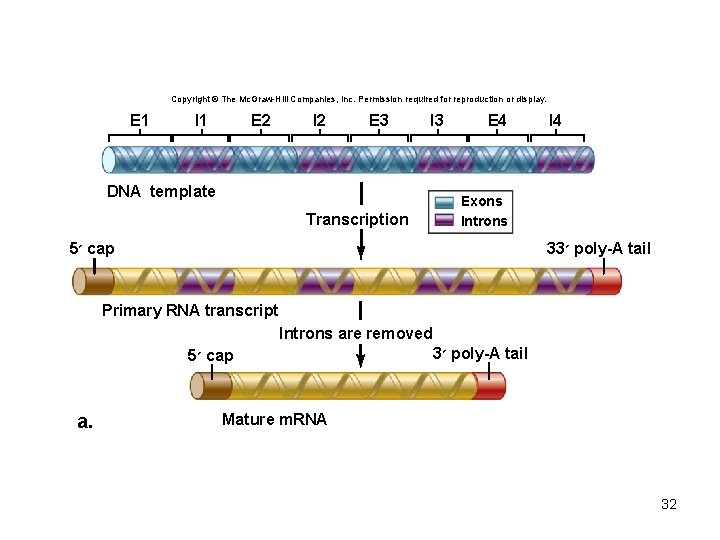
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. E 1 I 1 E 2 I 2 E 3 I 3 DNA template Transcription E 4 I 4 Exons Introns 5 ׳ cap 33 ׳ poly-A tail Primary RNA transcript Introns are removed 5 ׳ cap a. 3 ׳ poly-A tail Mature m. RNA 32
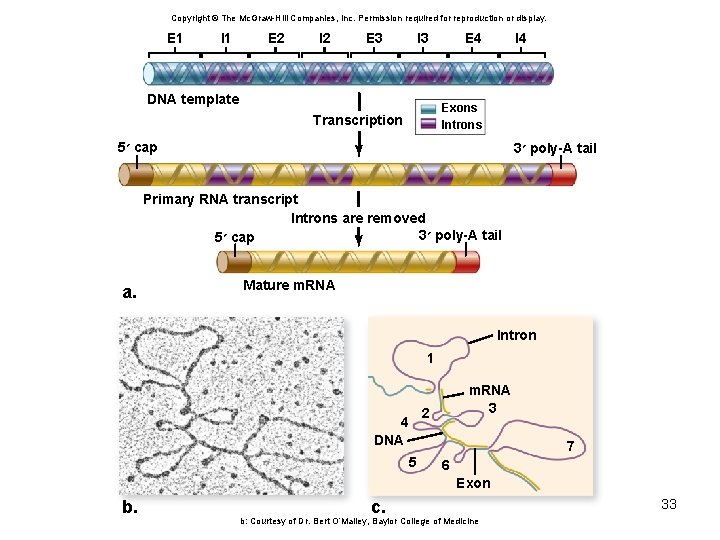
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. E 1 I 1 E 2 I 2 E 3 I 3 DNA template E 4 I 4 Exons Introns Transcription 5 ׳ cap 3 ׳ poly-A tail Primary RNA transcript Introns are removed 3 ׳ poly-A tail 5 ׳ cap a. Mature m. RNA Intron 1 m. RNA 3 2 4 DNA 7 5 6 Exon b. c. b: Courtesy of Dr. Bert O’Malley, Baylor College of Medicine 33
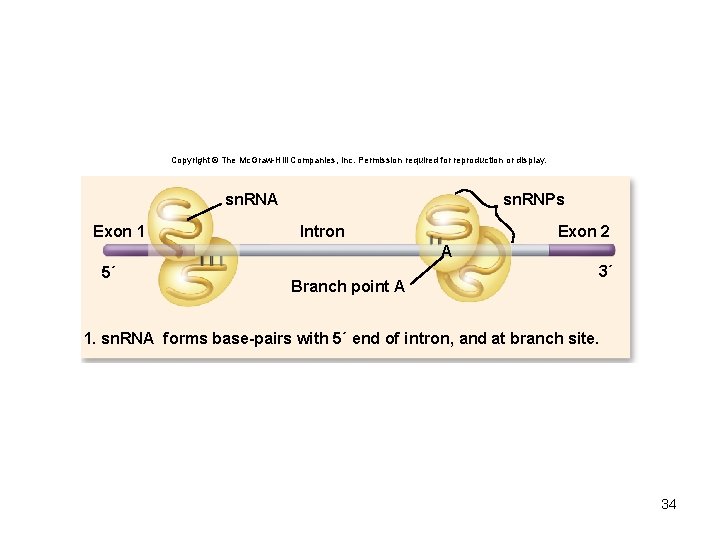
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. sn. RNA Exon 1 sn. RNPs Intron Exon 2 A 5´ Branch point A 3´ 1. sn. RNA forms base-pairs with 5´ end of intron, and at branch site. 34
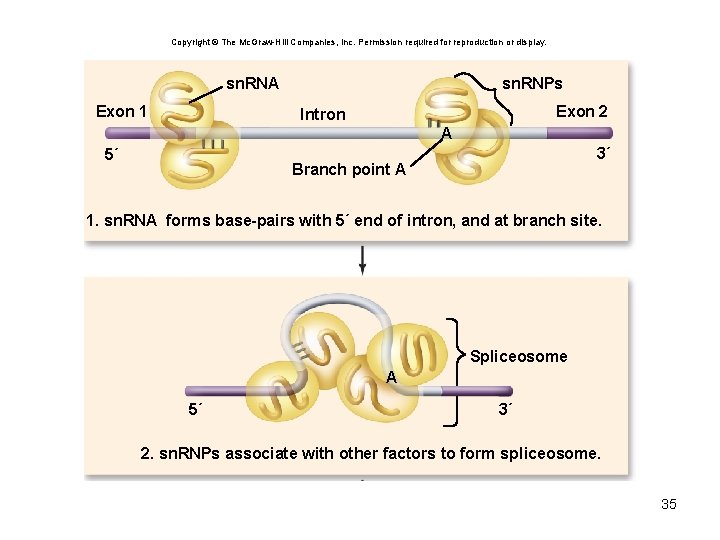
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. sn. RNA Exon 1 sn. RNPs Exon 2 Intron A 5´ 3´ Branch point A 1. sn. RNA forms base-pairs with 5´ end of intron, and at branch site. Spliceosome A 5´ 3´ 2. sn. RNPs associate with other factors to form spliceosome. 35
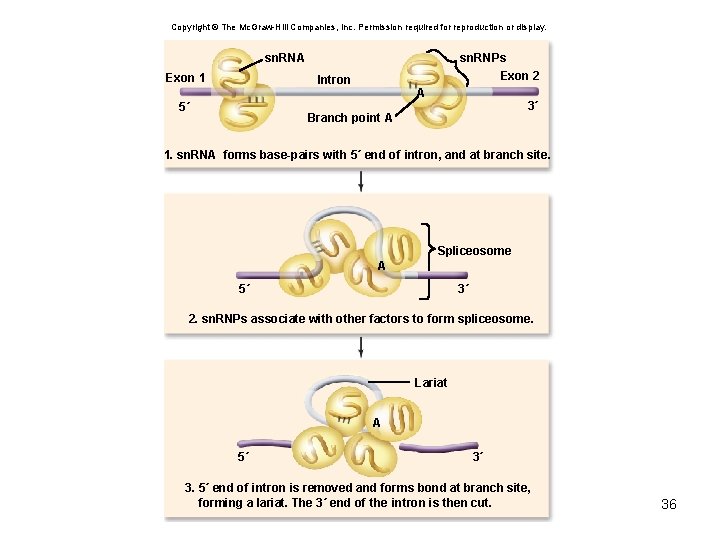
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. sn. RNA Exon 1 sn. RNPs Exon 2 Intron A 5´ 3´ Branch point A 1. sn. RNA forms base-pairs with 5´ end of intron, and at branch site. Spliceosome A 5´ 3´ 2. sn. RNPs associate with other factors to form spliceosome. Lariat A 5´ 3´ 3. 5´ end of intron is removed and forms bond at branch site, forming a lariat. The 3´ end of the intron is then cut. 36
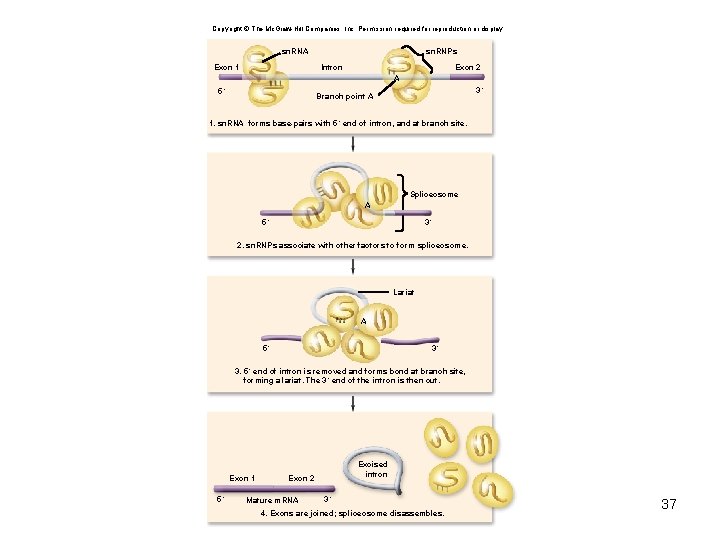
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. sn. RNA Exon 1 sn. RNPs Intron Exon 2 A 5´ 3´ Branch point A 1. sn. RNA forms base-pairs with 5´ end of intron, and at branch site. Spliceosome A 5´ 3´ 2. sn. RNPs associate with other factors to form spliceosome. Lariat A 5´ 3´ 3. 5´ end of intron is removed and forms bond at branch site, forming a lariat. The 3´ end of the intron is then cut. Exon 1 5´ Excised intron Exon 2 Mature m. RNA 3´ 4. Exons are joined; spliceosome disassembles. 37

Alternative splicing • Single primary transcript can be spliced into different m. RNAs by the inclusion of different sets of exons • 15% of known human genetic disorders are due to altered splicing • 35 to 59% of human genes exhibit some form of alternative splicing • Explains how 25, 000 genes of the human genome can encode the more than 80, 000 different m. RNAs 38
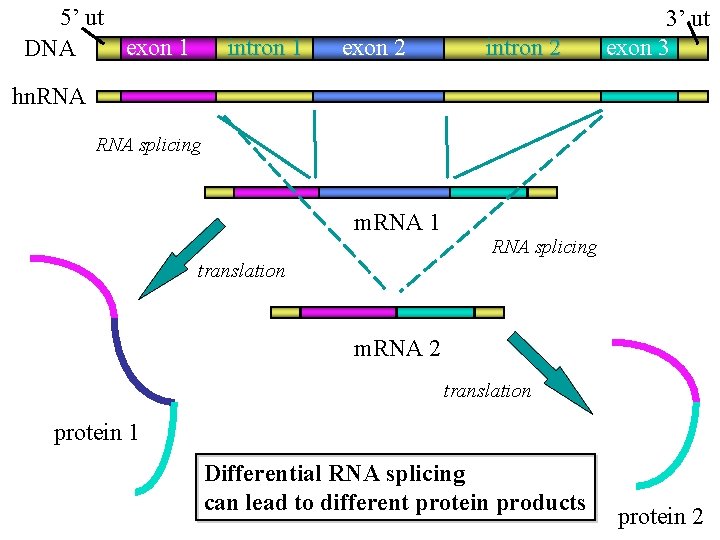
5’ ut exon 1 DNA intron 1 exon 2 intron 2 3’ ut exon 3 hn. RNA splicing m. RNA 1 RNA splicing translation m. RNA 2 translation protein 1 Differential RNA splicing can lead to different protein products protein 2

t. RNA and Ribosomes • t. RNA molecules carry amino acids to the ribosome for incorporation into a polypeptide – Aminoacyl-t. RNA synthetases add amino acids to the acceptor stem of t. RNA – Anticodon loop contains 3 nucleotides complementary to m. RNA codons 40
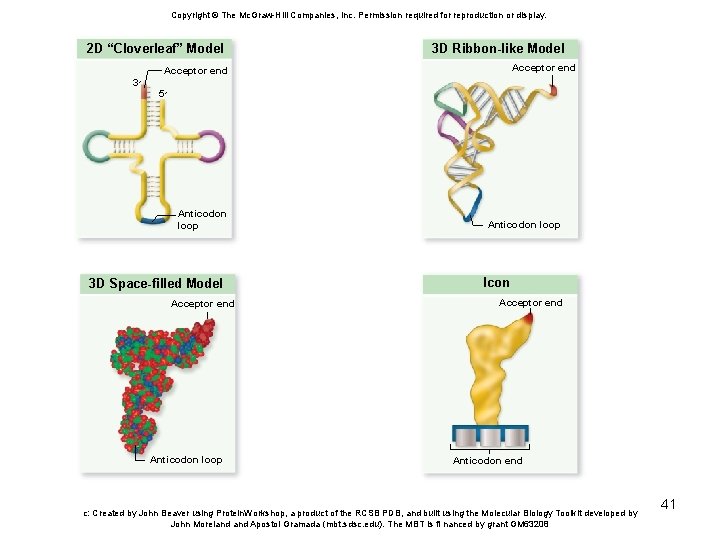
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. 2 D “Cloverleaf” Model 3 D Ribbon-like Model Acceptor end 3 ׳ 5 ׳ Anticodon loop 3 D Space-filled Model Acceptor end Anticodon loop Icon Acceptor end Anticodon end c: Created by John Beaver using Protein. Workshop, a product of the RCSB PDB, and built using the Molecular Biology Toolkit developed by John Moreland Apostol Gramada (mbt. sdsc. edu). The MBT is fi nanced by grant GM 63208 41

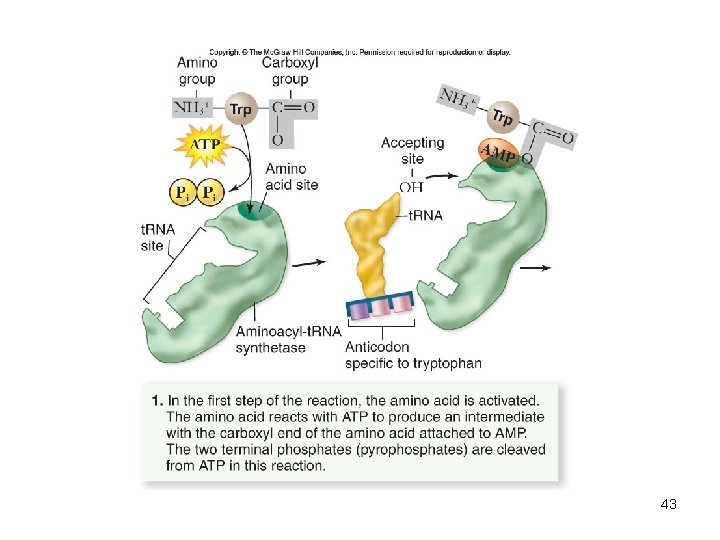
t. RNA charging reaction • Each aminoacyl-t. RNA synthetase recognizes only 1 amino acid but several t. RNAs • Charged t. RNA – has an amino acid added using the energy from ATP – Can undergo peptide bond formation without additional energy • Ribosomes do not verify amino acid attached to t. RNA 42
43
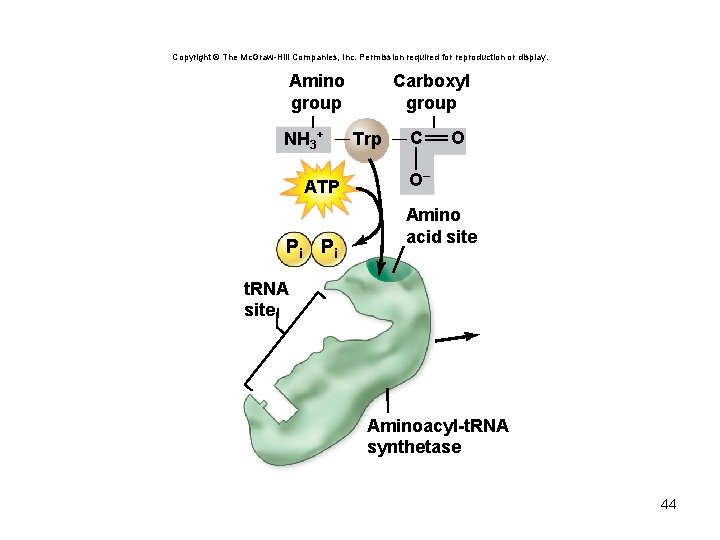
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Amino group NH 3+ ATP Pi Pi Carboxyl group Trp C O O– Amino acid site t. RNA site Aminoacyl-t. RNA synthetase 44
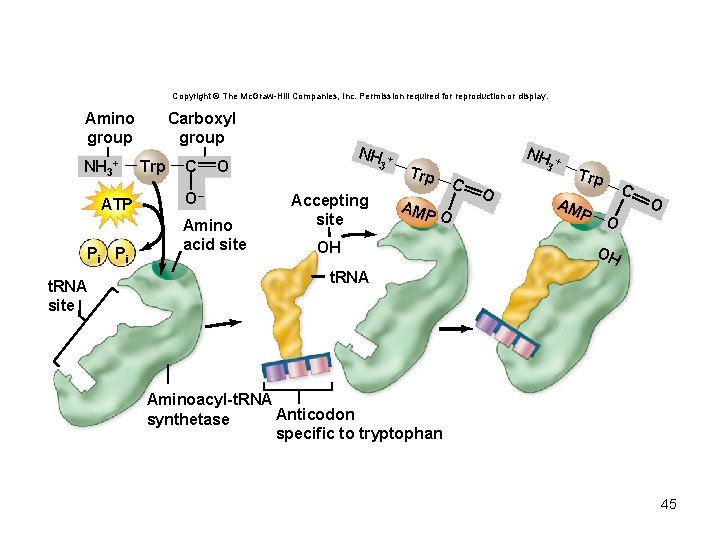
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Amino group NH 3+ ATP Pi Pi t. RNA site Carboxyl group Trp C NH O O– Amino acid site 3 Accepting site + Trp AM NH 3 C PO OH O + Trp AM P O C O OH t. RNA Aminoacyl-t. RNA Anticodon synthetase specific to tryptophan 45
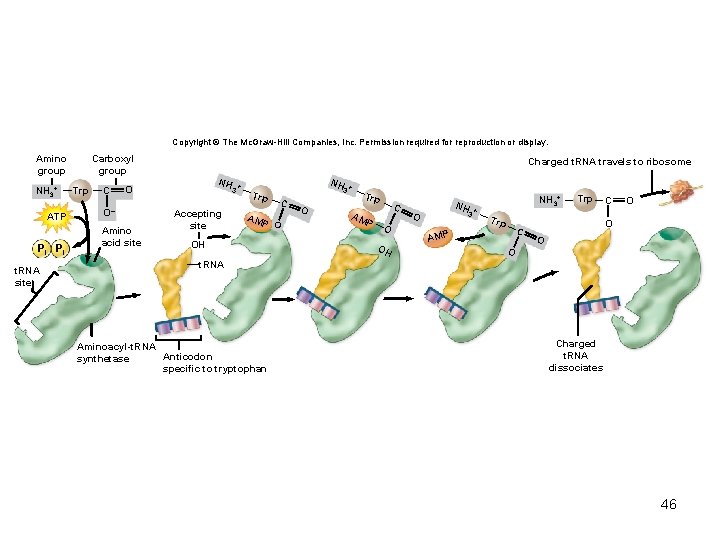
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Amino group NH 3+ ATP Pi Pi t. RNA site Carboxyl group Trp C Charged t. RNA travels to ribosome NH O O– Amino acid site 3 Accepting site NH + Trp AM P O OH 3 C O + Trp AM P C O OH NH + 3 O NH 3+ Trp C AMP Trp C O O t. RNA Aminoacyl-t. RNA Anticodon synthetase specific to tryptophan Charged t. RNA dissociates 46

• The ribosome has multiple t. RNA binding sites – P site – binds the t. RNA attached to the growing peptide chain – A site – binds the t. RNA carrying the next amino acid – E site – binds the t. RNA that carried the last amino acid 47
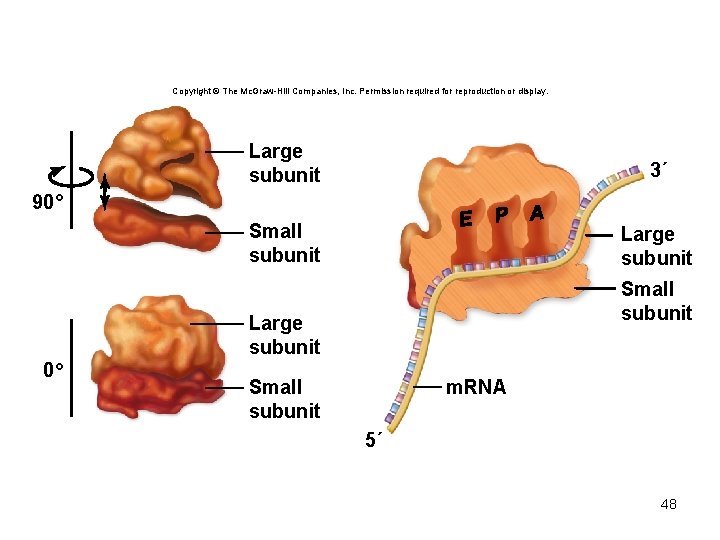
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Large subunit 3´ Small subunit Large subunit 90° Small subunit Large subunit 0° m. RNA Small subunit 5´ 48

• The ribosome has two primary functions – Decode the m. RNA – Form peptide bonds • Peptidyl transferase – Enzymatic component of the ribosome – Forms peptide bonds between amino acids 49

Translation • In prokaryotes, initiation complex includes – Initiator t. RNA charged with N-formylmethionine – Small ribosomal subunit – m. RNA strand • Ribosome binding sequence (RBS) of m. RNA positions small subunit correctly • Large subunit now added • Initiator t. RNA bound to P site with A site empty 50
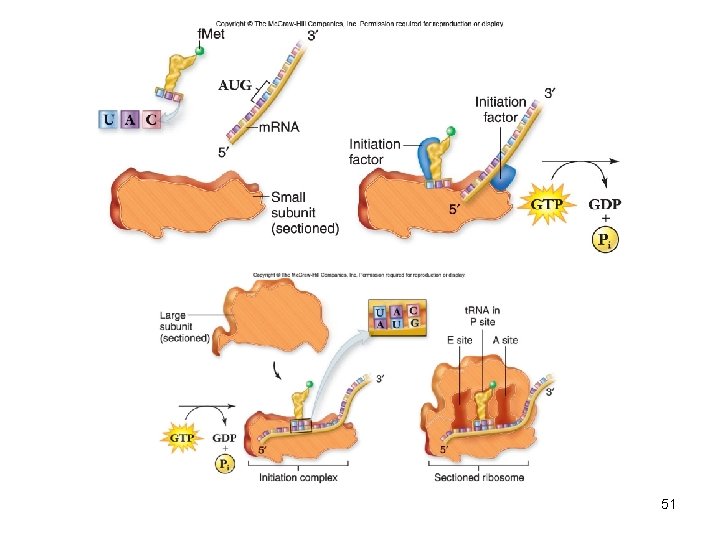
51

• Initiations in eukaryotes similar except – Initiating amino acid is methionine – More complicated initiation complex – Lack of an RBS – small subunit binds to 5′ cap of m. RNA 52

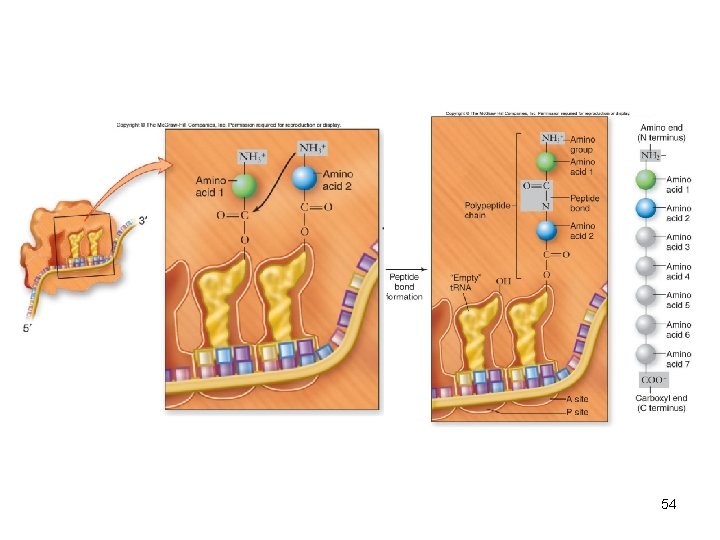
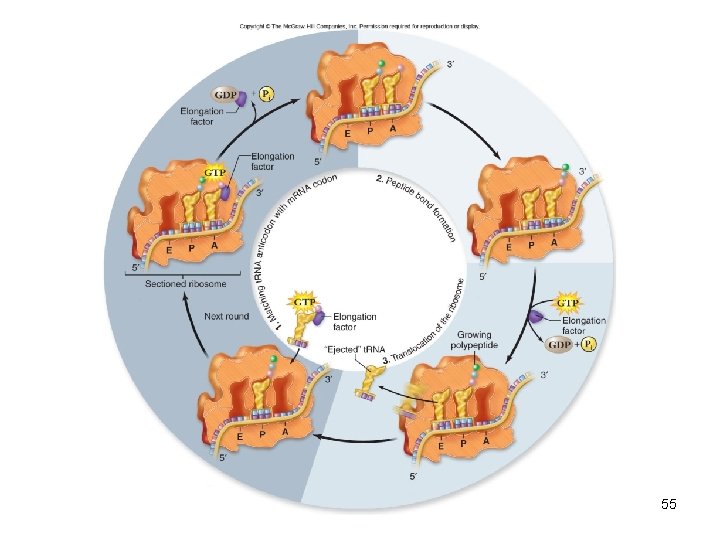
• Elongation adds amino acids – 2 nd charged t. RNA can bind to empty A site – Requires elongation factor called EF-Tu to bind to t. RNA and GTP – Peptide bond can then form – Addition of successive amino acids occurs as a cycle 53
54
55

• There are fewer t. RNAs than codons • Wobble pairing allows less stringent pairing between the 3′ base of the codon and the 5′ base of the anticodon • This allows fewer t. RNAs to accommodate all codons 56

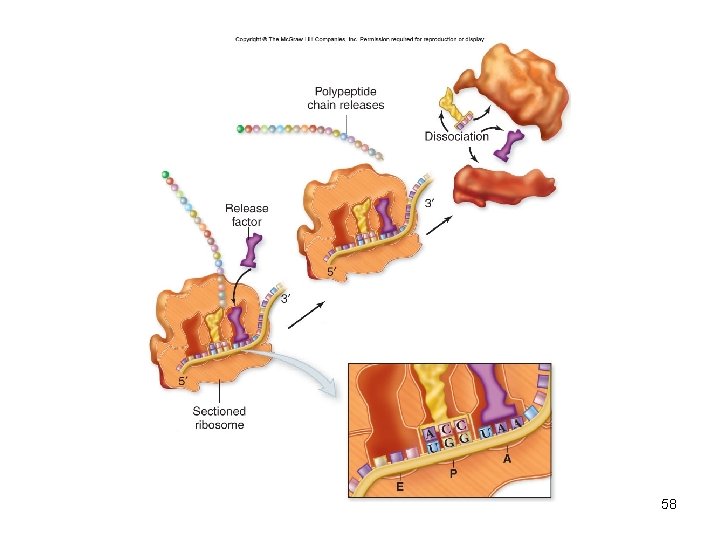
• Termination – Elongation continues until the ribosome encounters a stop codon – Stop codons are recognized by release factors which release the polypeptide from the ribosome 57
58

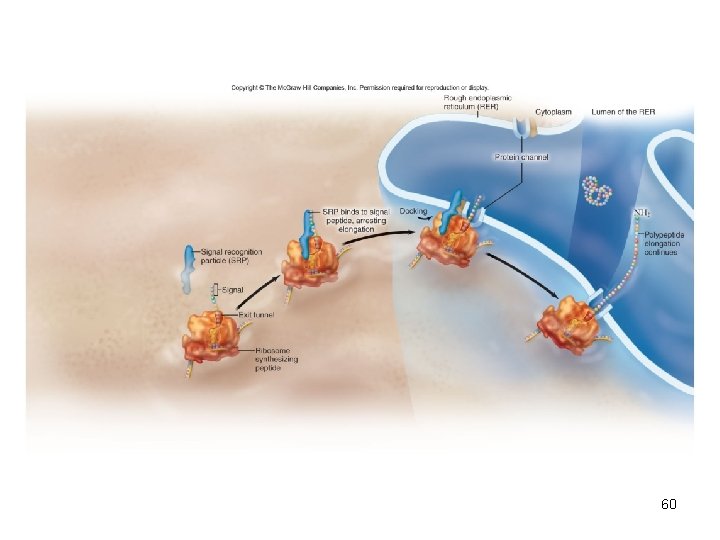
Protein targeting • In eukaryotes, translation may occur in the cytoplasm or the rough endoplasmic reticulum (RER) • Signal sequences at the beginning of the polypeptide sequence bind to the signal recognition particle (SRP) • The signal sequence and SRP are recognized by RER receptor proteins • Docking holds ribosome to RER • Beginning of the protein-trafficking pathway 59
60
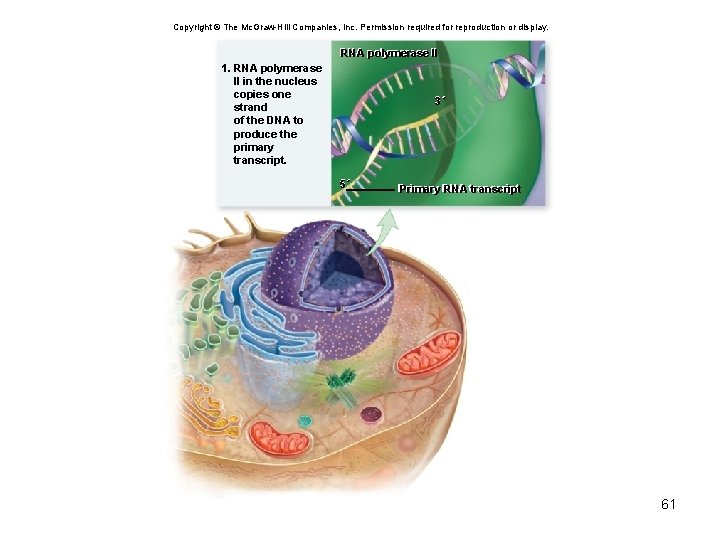
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 61
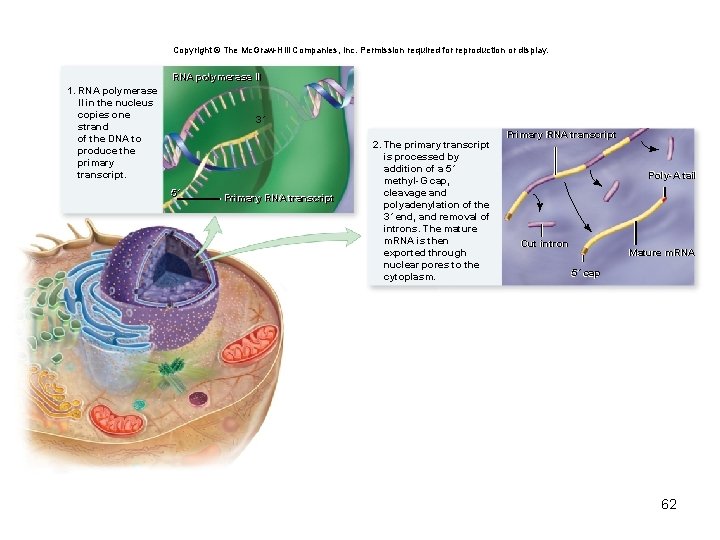
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 2. The primary transcript is processed by addition of a 5´ methyl-G cap, cleavage and polyadenylation of the 3´ end, and removal of introns. The mature m. RNA is then exported through nuclear pores to the cytoplasm. Primary RNA transcript Poly-A tail Poly-A Cut intron Cut Mature m. RNA 5´ cap 5´ 62
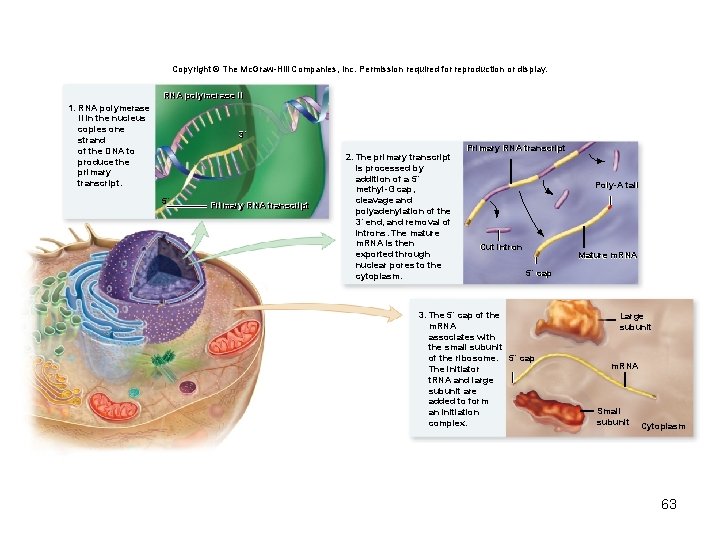
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 2. The primary transcript is processed by addition of a 5´ methyl-G cap, cleavage and polyadenylation of the 3´ end, and removal of introns. The mature m. RNA is then exported through nuclear pores to the cytoplasm. Primary RNA transcript Poly-A tail Cut intron Mature m. RNA 5´ cap 3. The 5´ cap of the m. RNA associates with the small subunit of the ribosome. 5´ cap The initiator t. RNA and large subunit are added to form an initiation complex. Large subunit m. RNA Small subunit Cytoplasm 63
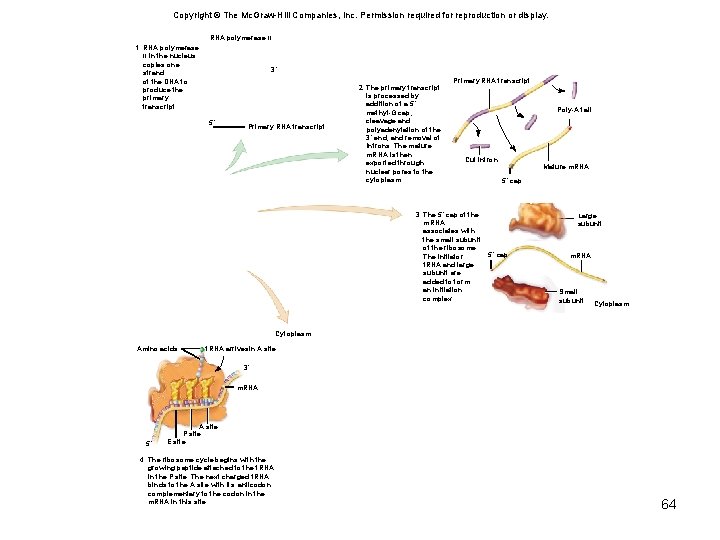
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 2. The primary transcript is processed by addition of a 5´ methyl-G cap, cleavage and polyadenylation of the 3´ end, and removal of introns. The mature m. RNA is then exported through nuclear pores to the cytoplasm. Primary RNA transcript Poly-A tail Cut intron 3. The 5´ cap of the m. RNA associates with the small subunit of the ribosome. The initiator t. RNA and large subunit are added to form an initiation complex. Mature m. RNA 5´ cap Large subunit 5´ cap m. RNA Small subunit Cytoplasm Amino acids t. RNA arrivesin A site 3´ m. RNA 5´ A site P site E site 4. The ribosome cycle begins with the growing peptide attached to the t. RNA in the P site. The next charged t. RNA binds to the A site with its anticodon complementary to the codon in the m. RNA in this site. 64
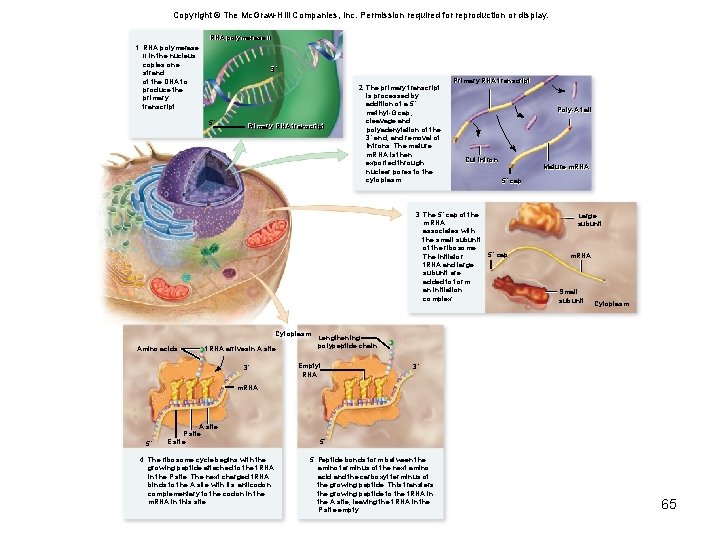
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 2. The primary transcript is processed by addition of a 5´ methyl-G cap, cleavage and polyadenylation of the 3´ end, and removal of introns. The mature m. RNA is then exported through nuclear pores to the cytoplasm. Primary RNA transcript Poly-A tail Cut intron 3. The 5´ cap of the m. RNA associates with the small subunit of the ribosome. The initiator t. RNA and large subunit are added to form an initiation complex. Cytoplasm Amino acids t. RNA arrivesin A site 3´ Mature m. RNA 5´ cap Large subunit 5´ cap m. RNA Small subunit Cytoplasm Lengthening polypeptide chain Emptyt RNA 3´ m. RNA 5´ A site P site E site 4. The ribosome cycle begins with the growing peptide attached to the t. RNA in the P site. The next charged t. RNA binds to the A site with its anticodon complementary to the codon in the m. RNA in this site. 5´ 5. Peptide bonds form between the amino terminus of the next amino acid and the carboxyl terminus of the growing peptide. This transfers the growing peptide to the t. RNA in the A site, leaving the t. RNA in the P site empty. 65
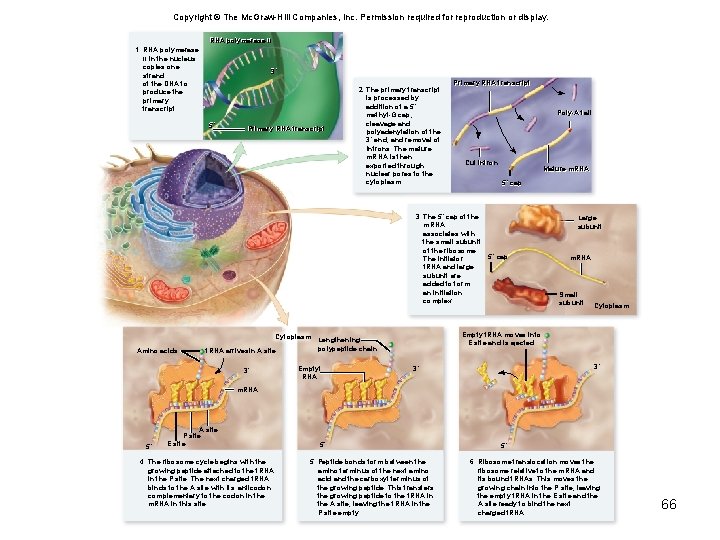
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. RNA polymerase IIII 1. RNA polymerase II in the nucleus copies one strand of the DNA to produce the primary transcript. 3´ 5´ Primary RNA transcript 2. The primary transcript is processed by addition of a 5´ methyl-G cap, cleavage and polyadenylation of the 3´ end, and removal of introns. The mature m. RNA is then exported through nuclear pores to the cytoplasm. Primary RNA transcript Poly-A tail Cut intron 5´ cap 3. The 5´ cap of the m. RNA associates with the small subunit of the ribosome. The initiator t. RNA and large subunit are added to form an initiation complex. Cytoplasm Amino acids t. RNA arrivesin A site 3´ Large subunit 5´ cap m. RNA Small subunit Cytoplasm Empty t. RNA moves into E site and is ejected Lengthening polypeptide chain Emptyt RNA Mature m. RNA 3´ 3´ m. RNA 5´ A site P site E site 4. The ribosome cycle begins with the growing peptide attached to the t. RNA in the P site. The next charged t. RNA binds to the A site with its anticodon complementary to the codon in the m. RNA in this site. 5´ 5. Peptide bonds form between the amino terminus of the next amino acid and the carboxyl terminus of the growing peptide. This transfers the growing peptide to the t. RNA in the A site, leaving the t. RNA in the P site empty. 5´ 6. Ribosome translocation moves the ribosome relative to the m. RNA and its bound t. RNAs. This moves the growing chain into the P site, leaving the empty t. RNA in the E site and the A site ready to bind the next charged t. RNA. 66
67

Please note that due to differing operating systems, some animations will not appear until the presentation is viewed in Presentation Mode (Slide Show view). You may see blank slides in the “Normal” or “Slide Sorter” views. All animations will appear after viewing in Presentation Mode and playing each animation. Most animations will require the latest version of the Flash Player, which is available at http: //get. adobe. com/flashplayer. 68

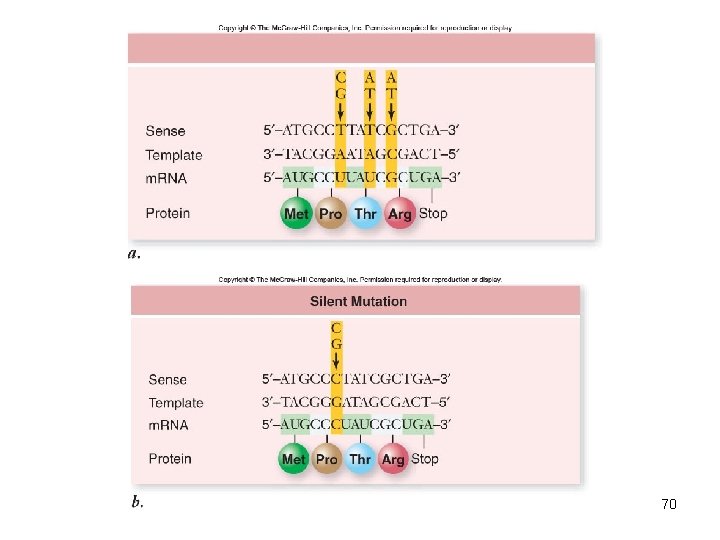
Mutation: Altered Genes • Point mutations alter a single base • Base substitution – substitute one base for another – Silent mutation – same amino acid inserted – Missense mutation – changes amino acid inserted • Transitions • Transversions – Nonsense mutations – changed to stop codon 69
70
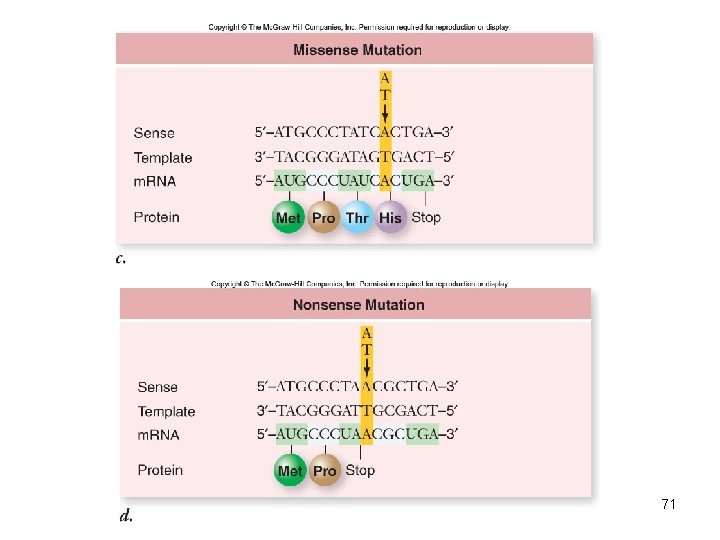
71
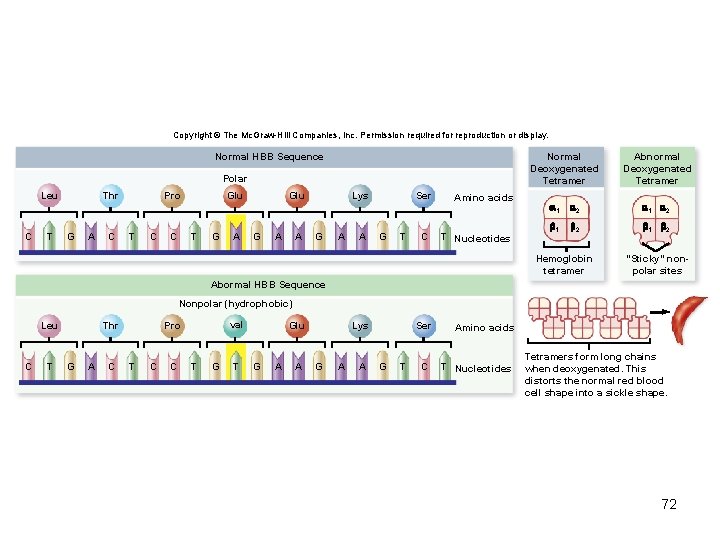
Copyright © The Mc. Graw-Hill Companies, Inc. Permission required for reproduction or display. Normal Deoxygenated Tetramer Normal HBB Sequence Polar Leu C T Thr G A C Pro T C C Glu T G A Glu G A A Lys G A A Ser G T C Amino acids T Nucleotides Abnormal Deoxygenated Tetramer 1 2 1 2 Hemoglobin tetramer "Sticky" nonpolar sites Abormal HBB Sequence Nonpolar (hydrophobic) Leu C T Thr G A C val Pro T C C T Glu G A A Lys G A A Ser G T C Amino acids T Nucleotides Tetramers form long chains when deoxygenated. This distorts the normal red blood cell shape into a sickle shape. 72

• Frameshift mutations – Addition or deletion of a single base – Much more profound consequences – Alter reading frame downstream – Triplet repeat expansion mutation • Huntington disease • Repeat unit is expanded in the disease allele relative to the normal 73

Chromosomal mutations • Change the structure of a chromosome – Deletions – part of chromosome is lost – Duplication – part of chromosome is copied – Inversion – part of chromosome in reverse order – Translocation – part of chromosome is moved to a new location 74
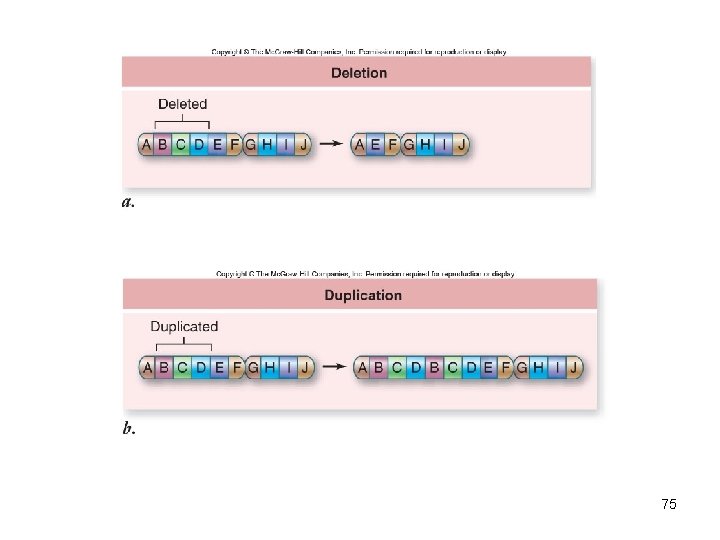
75
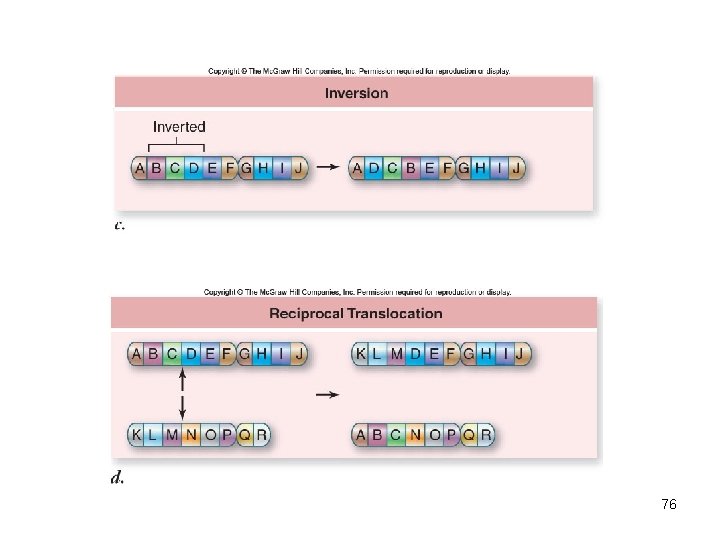
76

• Mutations are the starting point for evolution • Too much change, however, is harmful to the individual with a greatly altered genome • Balance must exist between amount of new variation and health of species 77
- Slides: 77