The Molecular Basis of Inheritance Chapter 16 HISTORY

The Molecular Basis of Inheritance Chapter 16

HISTORY OF DNA STRUCTURE n How did we learn that DNA is the key to coding for all characteristics of living things? ¨ Once Morgan’s group showed that genes are located on chromosomes, the two chemical components of chromosomes – DNA & PROTEIN, became the candidates for the genetic material.

TIMELINE n 1928: British scientist -- Frederick Griffith studies bacteria looking for cause of pneumonia: ¨ found two specific strains or cultures of bacteria that looked different when growing on petri dishes: ¨ n n n one grew in smooth-edged groups other one produced colonies that were rough and ragged around the edges IMPORTANCE: Visual differences made it easy to recognize and distinguish between the strains of bacteria ¨ Also, Griffith found that: ¨ n n smooth-edged colonies of bacteria caused disease rough-edged colonies were harmless
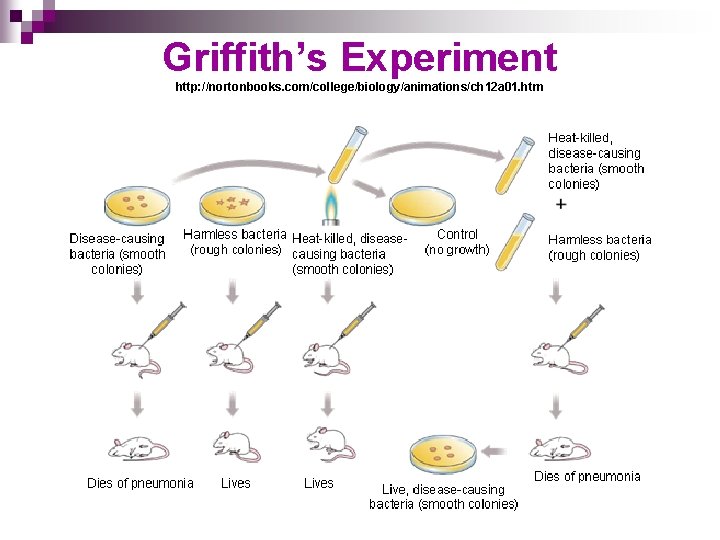
Griffith’s Experiment http: //nortonbooks. com/college/biology/animations/ch 12 a 01. htm

RESULTS OF GRIFFITH’S 1928 EXPERIMENT n n Discovery of process of transformation Somehow the heat-killed bacteria had passed their disease-causing ability to the harmless strain The harmless strain had been “transformed” into a disease-causing strain Hypothesized that some “factor” was responsible for this change…

TIMELINE CONTINUED n 1944: American, Oswald Avery, continued bacteria research of Griffith ¨ Knew were 4 types of organic compounds that make up all life… ¨ used enzymes to destroy lipids, carbohydrates, proteins, and RNA in an extract from the disease causing bacteria. ¨ n IMPORTANCE: ¨ Transformation still occurred, so obviously the molecules they had destroyed were not responsible for transformation. n n Only organic molecule left that had not been destroyed was DNA When repeated experiment with DNA-destroying enzymes, no transformation occurred…. DNA was the key to heredity

TIMELINE CONTINUED n 1952: ¨ Americans n Alfred Hershey and Martha Chase worked with viruses called bacteriophages ¨ viruses are simple – DNA or RNA core and a protein coat around them ¨ when they infect, bacteriophages inject DNA or RNA into cell and protein coat is left outside ¨ used radioactive markers to trace – n n phosphorus-32 (32 P) for DNA sulfur-35 (35 S) for protein coat
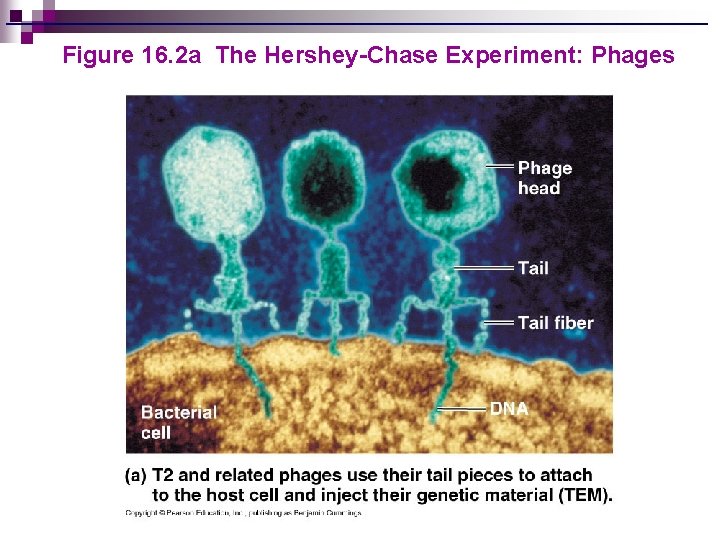
Figure 16. 2 a The Hershey-Chase Experiment: Phages

Hershey-Chase Experiment http: //nortonbooks. com/college/biology/animations/ch 12 a 02. htm Go to Section: Bacteriophage with phosphorus-32 in DNA Phage infects bacterium Radioactivity inside bacterium Bacteriophage with sulfur-35 in protein coat Phage infects bacterium No radioactivity inside bacterium

RESULTS OF HERSHEY-CHASE When viruses were separated from the bacteria and tested for radioactivity, all of the radioactivity from the bacteria was found to be 32 P – n Conclusion: n ¨ genetic material of the bacteriophage that was transferred was DNA – NOT PROTEIN!

RACE FOR STRUCTURE OF DNA n 1940: ¨ Erwin Chargaff discovers that percentages of A and T are equal in any sample of DNA; same is true for C and G n 1944: ¨ Linus Pauling discovers that proteins can have a helical shape n 1952: ¨ Rosalind Franklin takes pictures of DNA molecule using technique called X-ray diffraction, shows that DNA has helical shape
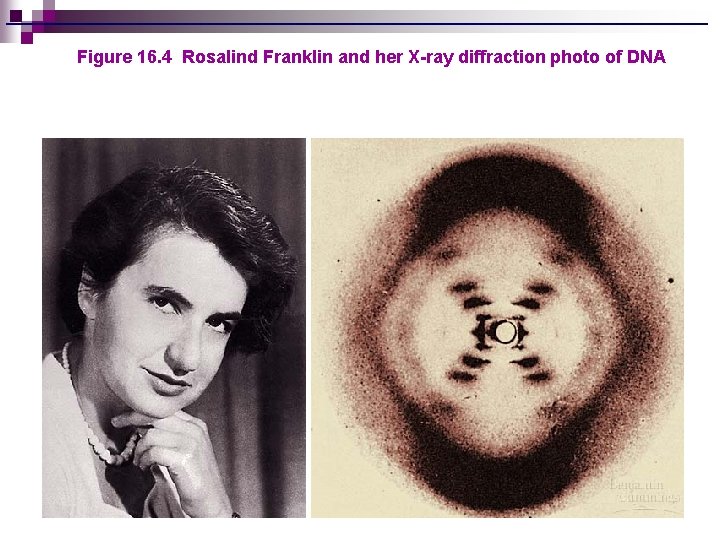
Figure 16. 4 Rosalind Franklin and her X-ray diffraction photo of DNA

RACE FOR STRUCTURE OF DNA n 1951 -1952: ¨ Maurice Wilkins works with X- ray diffraction and sees same pattern as Franklin, shares info with James Watson n April, 1953: ¨ James Watson and Francis Crick build first model of DNA (are awarded Nobel Prize in 1960’s)

Figure 5. x 3 James Watson and Francis Crick

Meselson-Stahl Experiment n Late 1950’s – experiments by Meselson & Stahl support the semi-conservative replication of DNA ¨ Data reject conservative and dispersal hypotheses!

Practice Questions n Which of the following is LEAST related to the others? ¨ Transformation ¨ Phage ¨ DNA ¨ Griffeth ¨ Avery n Answer: phage

Practice Questions n What was the contribution of the following scientists regarding the discovery of our present knowledge of the nature of genes and/or the shape of the DNA? ¨ Griffith ¨ Hershey & Chase ¨ Avery, Mac. Leod, and Mc. Carty ¨ Chargaff ¨ Meselson & Stahl

BASIC DNA STRUCTURE n n n Exists as a double helix Backbone of DNA structure made up of alternating sugar (deoxyribose) and phosphate groups Bases are attached to the sugars ¨ Bases in DNA are adenine, thymine, cytosine, and guanine n A pairs with T, C pairs with G and vice-versa ¨A and G are purines – larger, double rings ¨ T and C are pyrimidines – smaller, single rings
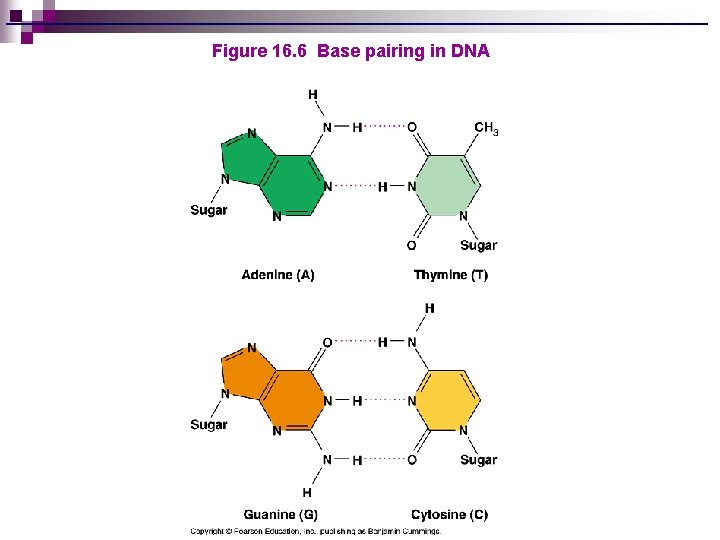
Figure 16. 6 Base pairing in DNA
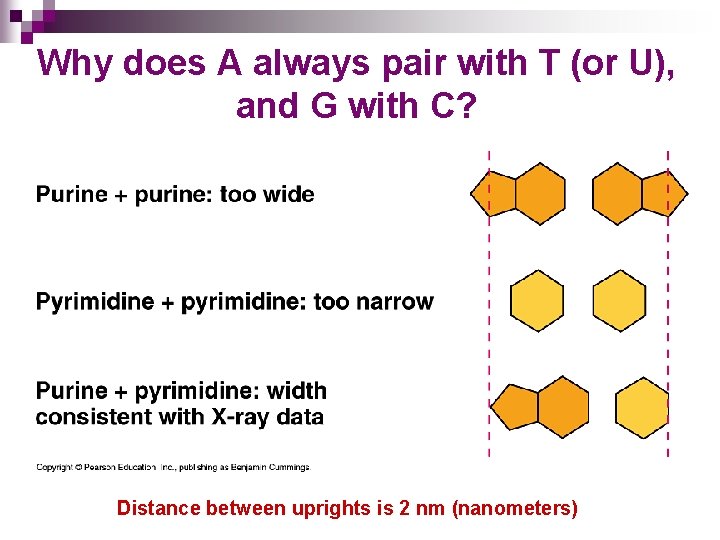
Why does A always pair with T (or U), and G with C? Distance between uprights is 2 nm (nanometers)
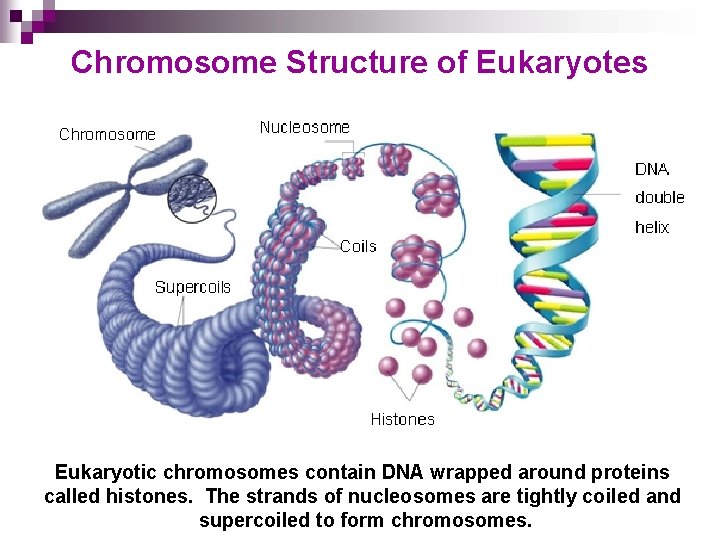
Chromosome Structure of Eukaryotes Eukaryotic chromosomes contain DNA wrapped around proteins Go tocalled histones. The strands of nucleosomes are tightly coiled and Section: supercoiled to form chromosomes.

3 Important Chromosome Functions Carry information from one generation to the next. 2. Put that information to work by determining the heritable characteristics of organisms. 3. Have to be easily copied each time a cell divides. 1.

Practice Question n All of the following elements are present in DNA except: ¨ Oxygen ¨ Nitrogen ¨ Carbon ¨ Sulfur ¨ Phosphorus n Answer: sulfur

Practice Questions n Cytosine makes up 38% of the nucleotides in a sample of DNA from an organism. What percent of the nucleotides in this sample will be thymine? ¨ 12 % ¨ 24 % ¨ 31 % ¨ 38 % ¨ Cannot be determined with information provided n Answer: 12 %

Practice Questions n In an analysis of the nucleotide composition of DNA, which of the following is TRUE? ¨A =C ¨ A = G and C = T ¨A + C = G + T ¨A + T = G + C ¨ Both B and C are true n Answer: A + C = G + T

Practice Question n All of the following were determined directly from X-ray diffraction photographs of crystallized DNA except: ¨ The diameter of the double helix ¨ The helical shape of DNA ¨ The sequence of nucleotides ¨ The linear distance required for one full turn of the double helix ¨ The width of the double helix n Answer: the sequence of nucleotides

DNA REPLICATION n n n Open up the helix to promote easier copying – HELICASE is the enzyme that facilitates this There are MANY helicases that work together to unwind the double helix efficiently…. WATCH THIS ANIMATION SEVERAL TIMES: ¨ http: //www. wiley. com/college/pratt/0471393878/stude nt/animations/dna_replication/index. html

Figure 16. 7 A model for DNA replication: the basic concept (Layer 1) REMEMBER: this all occurs during the S phase of interphase in mitosis –or- during MEIOSIS I ONLY! http: //bcs. whfreeman. com/thelifewire/content/chp 11/11020 02. html http: //bcs. whfreeman. com/thelifewire/content/chp 11/11020 03. html

Figure 16. 7 A model for DNA replication: the basic concept (Layer 2) Hydrogen bonds are broken here!!!
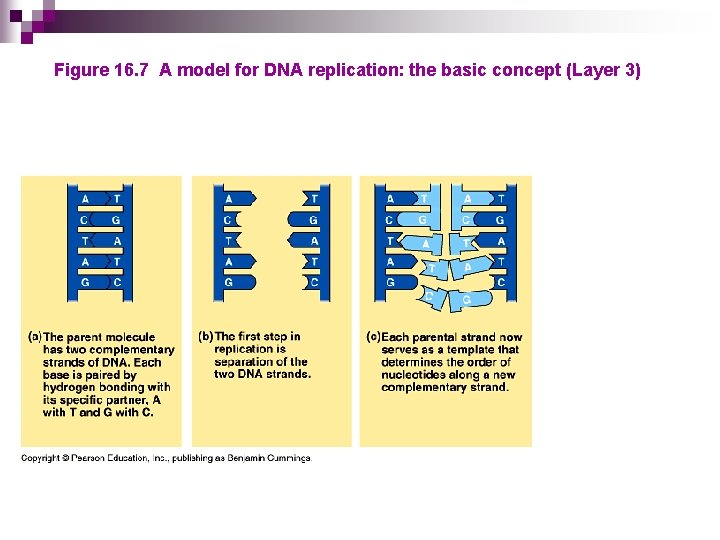
Figure 16. 7 A model for DNA replication: the basic concept (Layer 3)
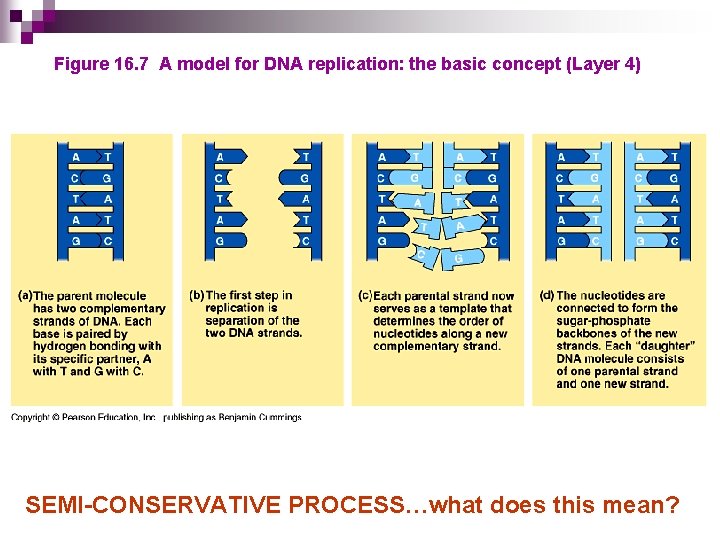
Figure 16. 7 A model for DNA replication: the basic concept (Layer 4) SEMI-CONSERVATIVE PROCESS…what does this mean?
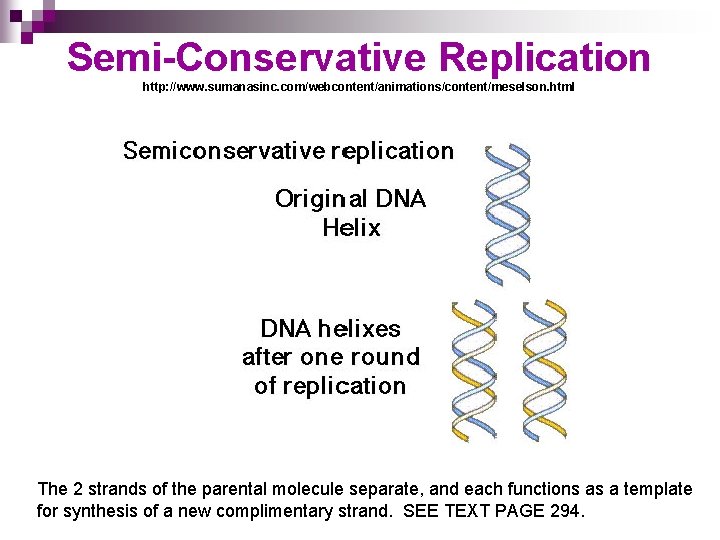
Semi-Conservative Replication http: //www. sumanasinc. com/webcontent/animations/content/meselson. html The 2 strands of the parental molecule separate, and each functions as a template for synthesis of a new complimentary strand. SEE TEXT PAGE 294.

Replication n The replication of a DNA molecule begins at an “origin of replication”. Bacterial chromosomes are circular and have a SINGLE origin of replication. ¨ Eukaryotic chromosomes are linear and may have hundreds of origins of replication. ¨ Proteins that initiate DNA replication recognize these sequences and attach to the DNA, separating the two strands and opening up a “replication bubble”. ¨ n n DNA replication occurs in a Replication Bubble; and each end is called a Replication fork Replication is catalyzed by the enzyme DNA polymerase – makes sure that correct bases pair. ¨ n Llike a “spell check” for the process of replication. DNA Ligase is the enzyme that forms the new covalent bonds between the DNA nucleotides.
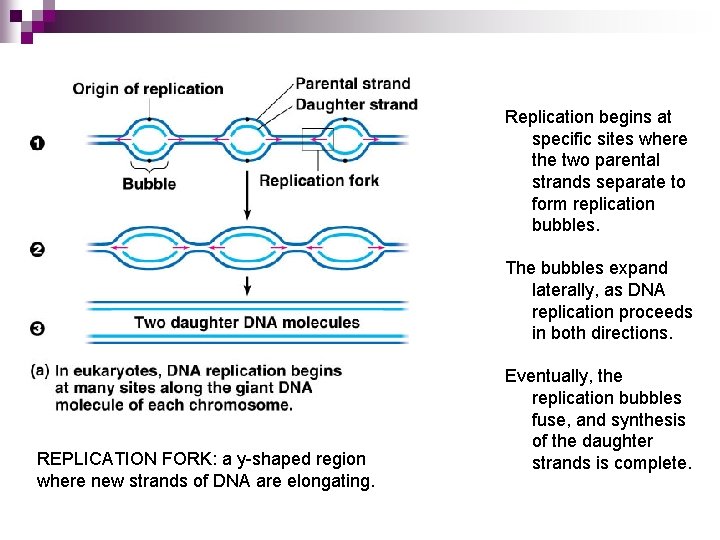
Replication begins at specific sites where the two parental strands separate to form replication bubbles. The bubbles expand laterally, as DNA replication proceeds in both directions. REPLICATION FORK: a y-shaped region where new strands of DNA are elongating. Eventually, the replication bubbles fuse, and synthesis of the daughter strands is complete.
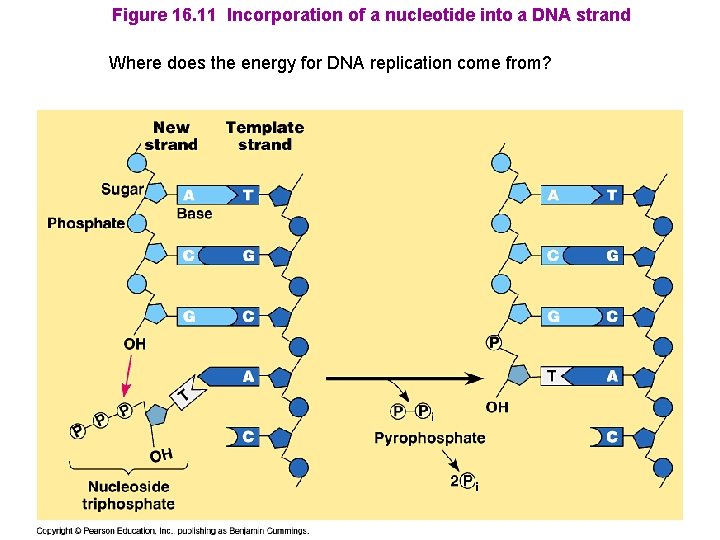
Figure 16. 11 Incorporation of a nucleotide into a DNA strand Where does the energy for DNA replication come from?
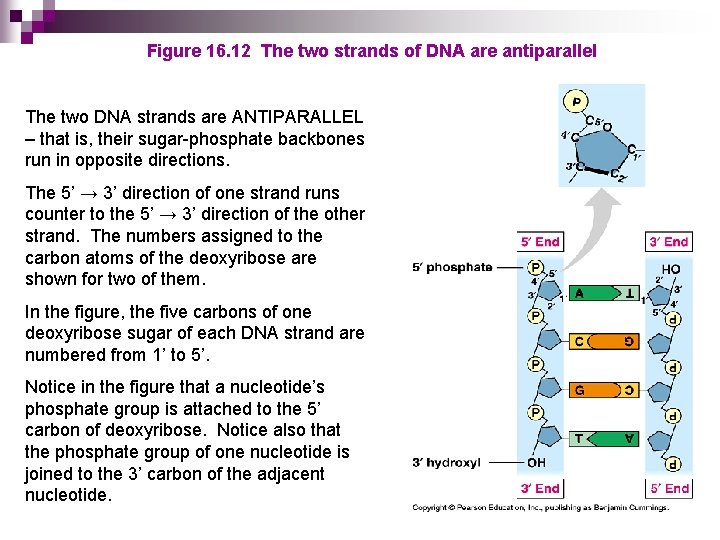
Figure 16. 12 The two strands of DNA are antiparallel The two DNA strands are ANTIPARALLEL – that is, their sugar-phosphate backbones run in opposite directions. The 5’ → 3’ direction of one strand runs counter to the 5’ → 3’ direction of the other strand. The numbers assigned to the carbon atoms of the deoxyribose are shown for two of them. In the figure, the five carbons of one deoxyribose sugar of each DNA strand are numbered from 1’ to 5’. Notice in the figure that a nucleotide’s phosphate group is attached to the 5’ carbon of deoxyribose. Notice also that the phosphate group of one nucleotide is joined to the 3’ carbon of the adjacent nucleotide.

Antiparallel Arrangement of DNA n Replicating the LEADING STRAND: ¨ Antiparallel – sugar-phosphate backbones run in opposite directions: n See page 296 in text ¨ DNA polymerases add nucleotides ONLY to the free 3’ end of a growing DNA strand, never to the 5’ end: n SO, a new DNA strand can only elongate in the 5’ 3’ direction! ¨ Polymerase nestles in the replication fork on the template strand that is read 5’ 3’ (leading strand) ¨ This strand is replicated continuously in the 5’ to 3’ direction.

Antiparallel Arrangement of DNA n Replicating the LAGGING STRAND: ¨ To elongate the other new DNA strand, polymerase must work along the other template strand in the direction AWAY FROM the replication fork. ¨ As a replication bubble opens, a polymerase molecule can work its way away from a replication fork and synthesize a short segment of DNA. ¨ As the bubble grows, another short segment of the lagging strand can be made in a similar way. ¨ This is known as DISCONTINUOUS replication – because the DNA is synthesized in short fragments called OKAZAKI fragments.

Practice Question n The strands that make up DNA are antiparallel. What does this mean? n Answer: the 5’ to 3’ direction of one strand runs counter to the 5’ to 3’ direction of the other strand
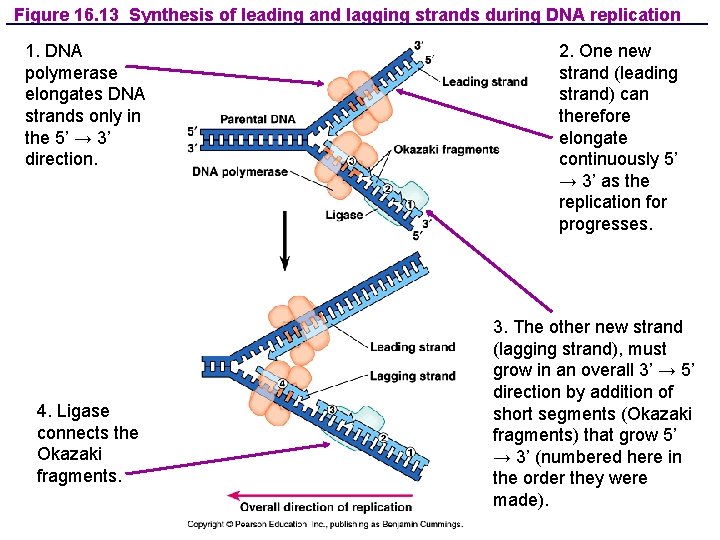
Figure 16. 13 Synthesis of leading and lagging strands during DNA replication 1. DNA polymerase elongates DNA strands only in the 5’ → 3’ direction. 4. Ligase connects the Okazaki fragments. 2. One new strand (leading strand) can therefore elongate continuously 5’ → 3’ as the replication for progresses. 3. The other new strand (lagging strand), must grow in an overall 3’ → 5’ direction by addition of short segments (Okazaki fragments) that grow 5’ → 3’ (numbered here in the order they were made).

Priming DNA Synthesis n n DNA polymerases cannot initiate synthesis of a polynucleotide – it can only add nucleotides to already existing chain! The start of a NEW chain is not DNA, but a short stretch of RNA called a primer. ¨ The enzyme primase joins this RNA to the DNA template to make the primer. ¨ Primase can start an RNA chain from scratch!
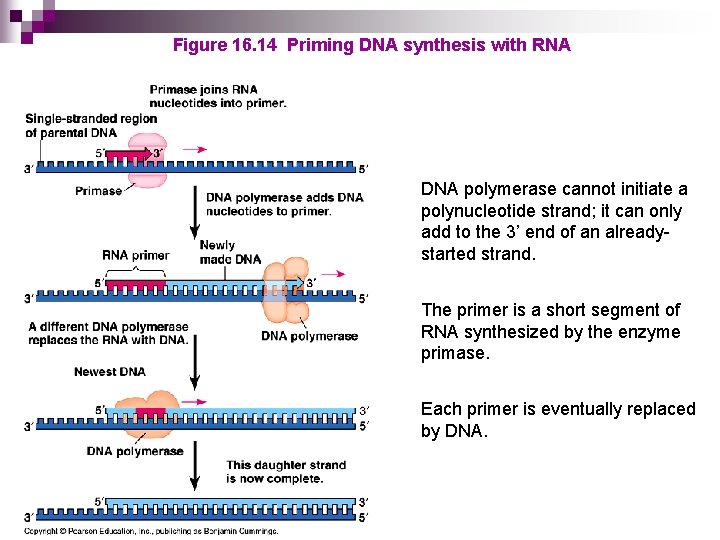
Figure 16. 14 Priming DNA synthesis with RNA DNA polymerase cannot initiate a polynucleotide strand; it can only add to the 3’ end of an alreadystarted strand. The primer is a short segment of RNA synthesized by the enzyme primase. Each primer is eventually replaced by DNA.
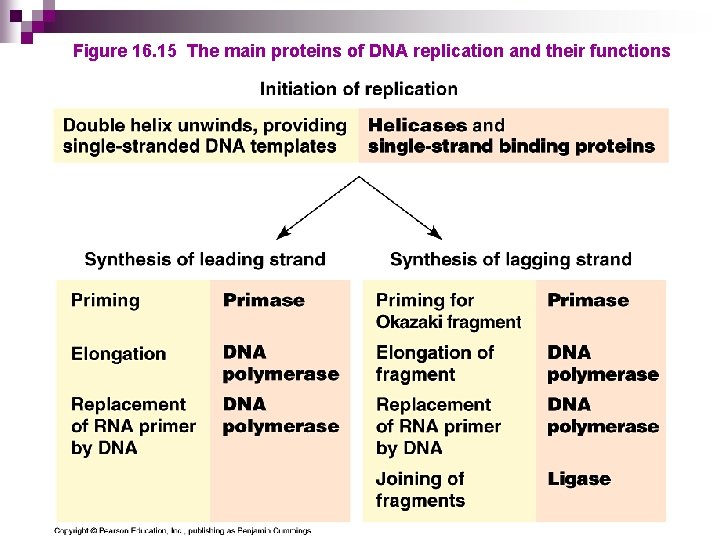
Figure 16. 15 The main proteins of DNA replication and their functions
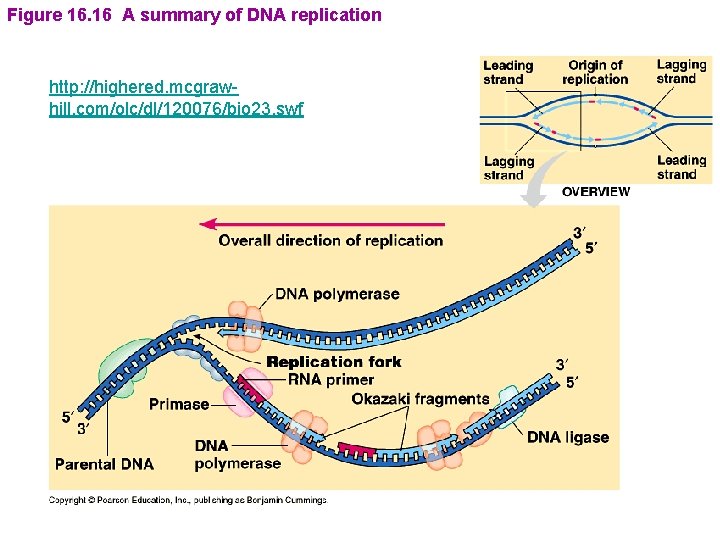
Figure 16. 16 A summary of DNA replication http: //highered. mcgrawhill. com/olc/dl/120076/bio 23. swf

Practice Question • Which enzymes catalyze the elongation of a DNA strand in the 5’ to 3’ direction? – Primase – DNA ligase – DNA polymerases – Topoisomerase – Helicase • Answer: DNA polymerases

Practice Question • What is the function of the following enzymes in DNA replication? – Helicase – SSB’s – Nuclease – Ligase – DNA polymerase – primase

Practice Question • In DNA, the designations 3’ and 5’ refer to what? • Answer: carbon atoms of deoxyribose to which phosphate groups may bond

Practice Question • Which of the following is LEAST related to the others on the list? – Okazaki fragment – Primer – Telomere – Leading strand – Lagging strand • Answer: telomere
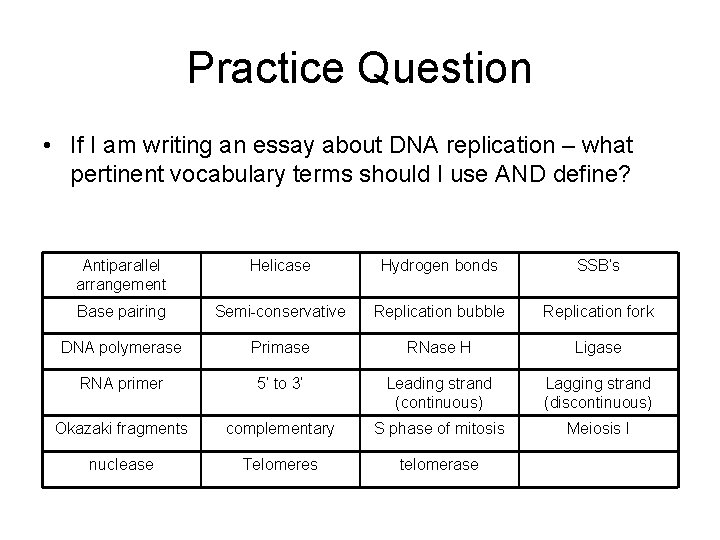
Practice Question • If I am writing an essay about DNA replication – what pertinent vocabulary terms should I use AND define? Antiparallel arrangement Helicase Hydrogen bonds SSB’s Base pairing Semi-conservative Replication bubble Replication fork DNA polymerase Primase RNase H Ligase RNA primer 5’ to 3’ Leading strand (continuous) Lagging strand (discontinuous) Okazaki fragments complementary S phase of mitosis Meiosis I nuclease Telomeres telomerase

PROOFREADING DNA FOR ERRORS n n Enzymes proofread DNA during its replication and repair damage in existing DNA See page 299 – ¨ errors in completed DNA molecule amount to only one in a billion nucleotides ¨ Initial pairing errors – error rate of 1 per 10000 base pairs ¨ exposure damage – reactive chemicals, radiation, Xrays, ultraviolet light (unpredictable, but common) n Ex. Skin cells and UV damage ¨ Mismatch repair – cells use special enzymes to fix incorrectly paired nucleotide ¨ Nucleotide excision repair – uses nuclease (DNA cutting enzyme that excises damaged area)
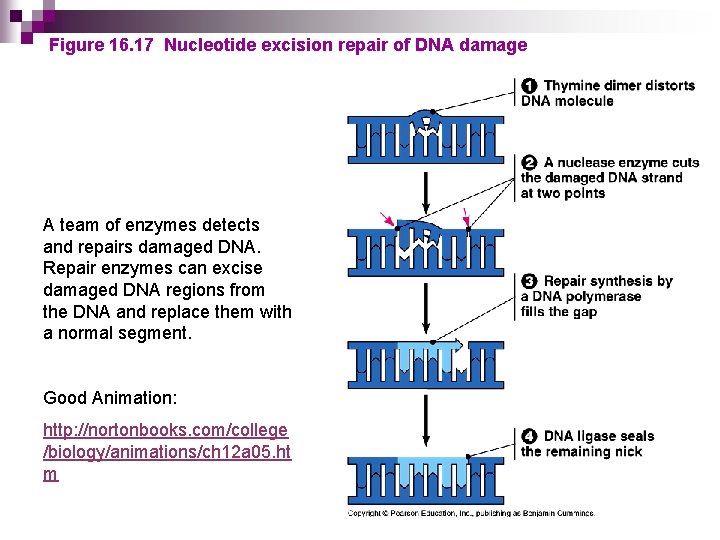
Figure 16. 17 Nucleotide excision repair of DNA damage A team of enzymes detects and repairs damaged DNA. Repair enzymes can excise damaged DNA regions from the DNA and replace them with a normal segment. Good Animation: http: //nortonbooks. com/college /biology/animations/ch 12 a 05. ht m

The Dilemma of DNA Ends n n For linear DNA, like that found in eukaryotes, DNA polymerase can only add nucleotides to the 3’ end of a preexisting polynucleotide – this is a potential problem! The normal replication machinery has no way to complete the 5’ ends of daughter DNA strands; as a result – repeated rounds of replication produce shorter and shorter DNA molecules. ¨ Result: n deletion of essential genes! NOTE: prokaryotes avoid this problem by having circular DNA!
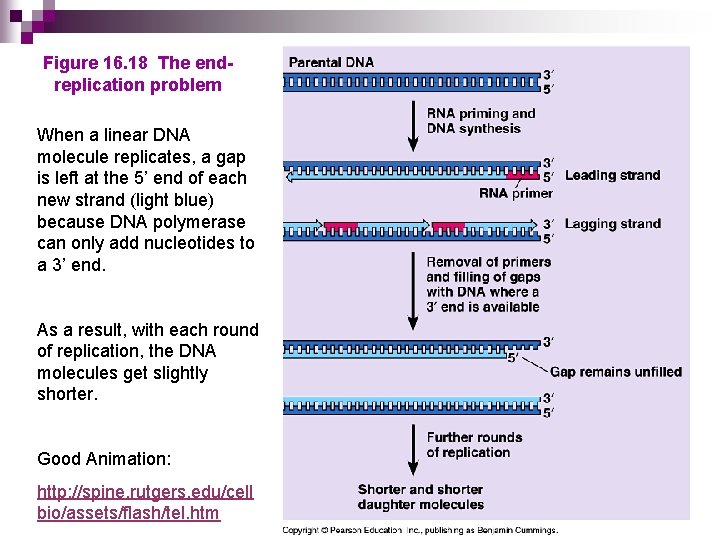
Figure 16. 18 The endreplication problem When a linear DNA molecule replicates, a gap is left at the 5’ end of each new strand (light blue) because DNA polymerase can only add nucleotides to a 3’ end. As a result, with each round of replication, the DNA molecules get slightly shorter. Good Animation: http: //spine. rutgers. edu/cell bio/assets/flash/tel. htm

Telomeres and Telomerase n Eukaryotic chromosomal DNA molecules have special nucleotide sequences called telomeres at their ends: ¨ n Typically, the repeated unit TTAGGG is seen in human telomeres. ¨ n Telomeres are regions of DNA that do not contain genes; rather, are multiple repetitions of one short nucleotide sequence. Telomeric DNA protects an organism’s genes from being eroded through successive rounds of DNA replication! These (along with special protein called telomerase associated with it) prevent the ends from activating the cell’s systems for monitoring DNA damage. ¨ Ex. The end of DNA strand seen as double strand “break” might trigger signal transduction pathways leading to cell cycle arrest or cell death)!
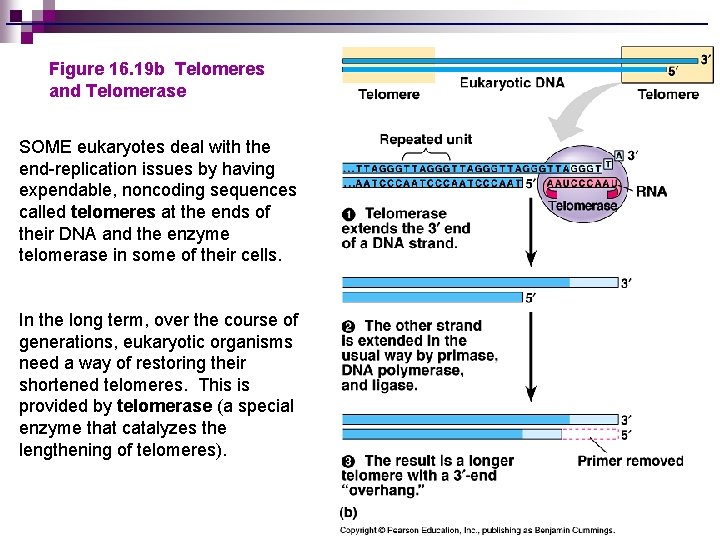
Figure 16. 19 b Telomeres and Telomerase SOME eukaryotes deal with the end-replication issues by having expendable, noncoding sequences called telomeres at the ends of their DNA and the enzyme telomerase in some of their cells. In the long term, over the course of generations, eukaryotic organisms need a way of restoring their shortened telomeres. This is provided by telomerase (a special enzyme that catalyzes the lengthening of telomeres).

Telomeres – Limiting Life? n Telomerase is NOT present in most cells of multicellular organisms like ourselves, and the DNA of dividing somatic cells does tend to be shorter in older individuals and in cultured cells that have divided many times. ¨ n Thus, it is possible that telomeres are a limiting factor in the life span of certain tissues and even organisms as a whole. Telomerase is however present in germ-line cells that give rise to gametes – and here the enzyme produces long telomeres in these cells and hence in the newborn. Intriguingly, telomerase is also found in somatic cells that are cancerous – these usually have unusually short telomeres, which one would expect for cells that have undergone many rounds of division. ¨ Progressive shortening would eventually lead to self-destruction of cancer unless telomerase became available to stabilize telomere length. ¨ This is exactly what seems to happen in cancer cells. IF this is an important factor – it may well provide a useful target for cancer diagnosis and chemotherapy! ¨

The Essay n Scientists seeking to determine which molecule is responsible for the transmission of characteristics from one generation to the next knew that the molecule must ¨ ¨ ¨ n n (1) copy itself precisely, (2) be stable but able to be changed, and (3) be complex enough to determine the organism's phenotype. Explain how DNA meets each of the three criteria stated above. Select one of the criteria stated above and describe experimental evidence used to determine that DNA is the hereditary material.
- Slides: 57