The International Tomato Sequencing Project The first Cornerstone

















- Slides: 26

The International Tomato Sequencing Project: The first Cornerstone of the SOL Project Lukas Mueller on behalf of International SOL Tomato Sequencing Project

Overview • • • Aims Why sequence the tomato genome? How to sequence the tomato genome? Who is sequencing the tomato genome? Resources for Sequencing the Tomato Genome – – – Genetic Map BAC libraries Overgo mapping BAC End Sequences Minimal Tiling Path Bioinformatics • Summary

Mingsheng Chen Zhukuan Cheng Chuanyou Li Hongqing Ling Yongbiao Xue Ramesh Sharma Jiten Khurana Akhilesh Tyagi Doil Choi Byung Dong Kim Antonio Granell Miguel A. Botella Giovanni Giuliano Luigi Fruciante Steven D. Tanksley Jim J. Giovannoni Stephen Stack, Joyce van Eck Graham Seymour Gerard Bishop Willem Stiekema P. Lindhout Taco Jesse Rene Klein Lankhorst Daisuke Shibata Satoshi Tabata Mondher Bouzayen Mathilde Causse

Aims • Provide a high quality reference sequence for the Solanaceae genomes • Using mapping of other Solanaceae sequences onto the tomato sequence, and comparative genetic maps to derive “virtual” genomes for other Solanaceae • Prerequisite for studying natural diversity and linking genotype to phenotype • Build a Solanaceae bioinformatics platform to integrate, analyze and distribute the information

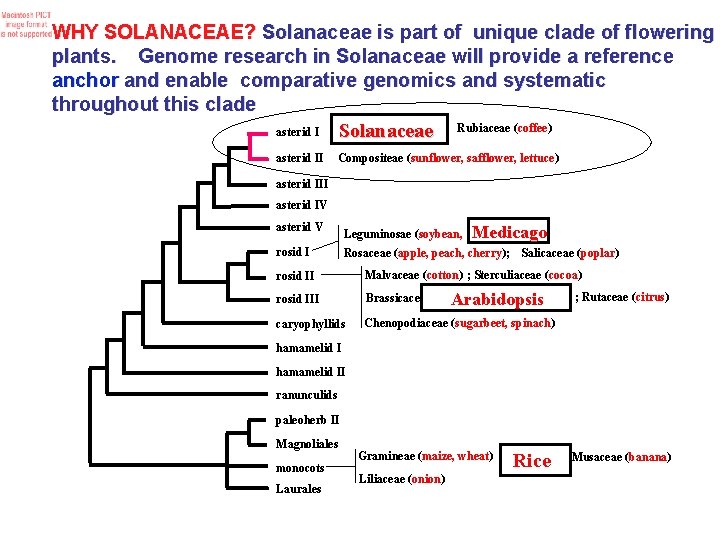
WHY SOLANACEAE? Solanaceae is part of unique clade of flowering plants. Genome research in Solanaceae will provide a reference anchor and enable comparative genomics and systematic throughout this clade asterid I Solanaceae Rubiaceae (coffee) asterid II Compositeae (sunflower, safflower, lettuce) asterid III asterid IV asterid V rosid I Leguminosae (soybean, Medicago Rosaceae (apple, peach, cherry); Salicaceae (poplar) rosid II Malvaceae (cotton) ; Sterculiaceae (cocoa) rosid III Brassicaceae caryophyllids Chenopodiaceae (sugarbeet, spinach) Arabidopsis ; Rutaceae (citrus) hamamelid II ranunculids paleoherb II Magnoliales monocots Laurales Gramineae (maize, wheat) Liliaceae (onion) Rice ; Musaceae (banana)

Why sequence tomato? • Tomato is the most intensively researched Solanaceae genome encoding approx. 35, 000 genes euchromatic regions corresponding to less than a 25% of the total DNA in the tomato nucleus (220~250 Mb). • Tomato provides the smallest diploid genome for which homozygous inbreds are available. • Its sequence will facilitate positional cloning in tomato and other Solanaceae genomes (via synteny maps).

How to sequence the tomato genome? • Whole Genome Shotgun – Advantages: Fast, cheaper, ok with reference genome – Disadvantages: Unordered contigs • Methylation Filtering (Tobacco) – Advantages: Selects for expressed genome, cheaper – Disadvantages: unordered contigs • Tiling Path (Arabidopsis, Drosophila, Rice) – Advantages: Sequence and gene order; select gene rich regions; easy to divide work – Disadvantages: Relatively expensive, time consuming • ORDER IMPORTANT FOR COMPARING GENOMES

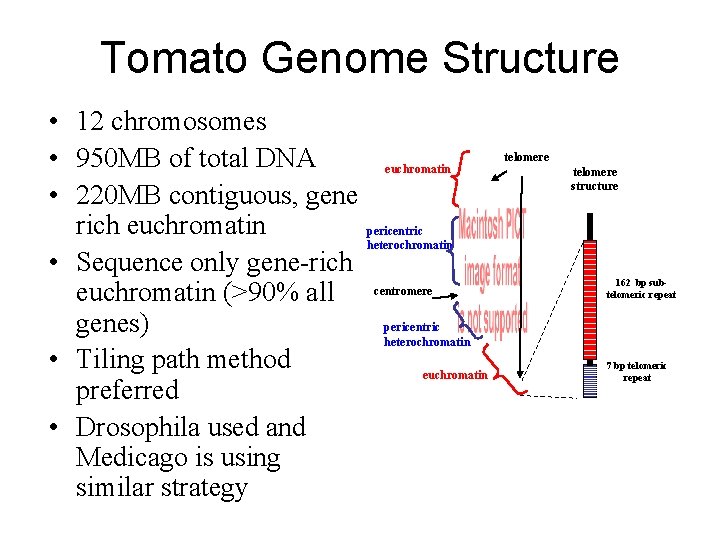
Tomato Genome Structure • 12 chromosomes • 950 MB of total DNA euchromatin • 220 MB contiguous, gene rich euchromatin pericentric heterochromatin • Sequence only gene-rich euchromatin (>90% all centromere pericentric genes) heterochromatin • Tiling path method euchromatin preferred • Drosophila used and Medicago is using similar strategy telomere structure 162 bp subtelomeric repeat 7 bp telomeric repeat

BAC libraries • All libraries derived from Solanum lycopersicum Heinz 1706. • Hind. III library (Rod Wing, Clemson U) – – – ~120, 000 clones, 120 k. B average size ~15 x coverage FPC contigged Overgo analysis 75, 000 clones BAC end sequenced • Mbo. I library – 50, 000 clones, 140 kb average size – Will be BAC end sequenced • Eco. RI library (being prepared) – Will be BAC end sequenced

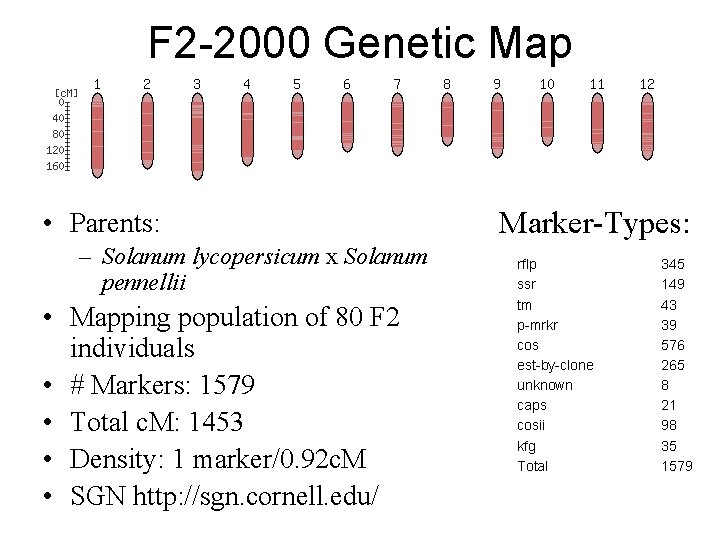
F 2 -2000 Genetic Map • Parents: – Solanum lycopersicum x Solanum pennellii • Mapping population of 80 F 2 individuals • # Markers: 1579 • Total c. M: 1453 • Density: 1 marker/0. 92 c. M • SGN http: //sgn. cornell. edu/ Marker-Types: rflp ssr tm p-mrkr cos est-by-clone unknown caps cosii kfg Total 345 149 43 39 576 265 8 21 98 35 1579

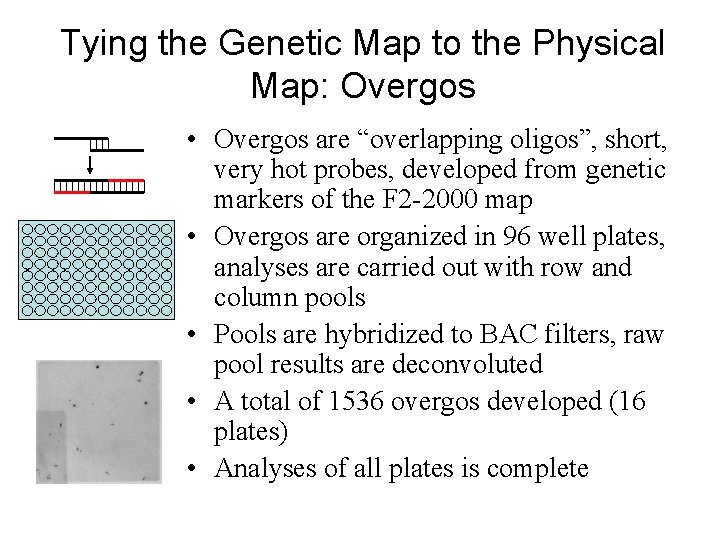
Tying the Genetic Map to the Physical Map: Overgos • Overgos are “overlapping oligos”, short, very hot probes, developed from genetic markers of the F 2 -2000 map • Overgos are organized in 96 well plates, analyses are carried out with row and column pools • Pools are hybridized to BAC filters, raw pool results are deconvoluted • A total of 1536 overgos developed (16 plates) • Analyses of all plates is complete

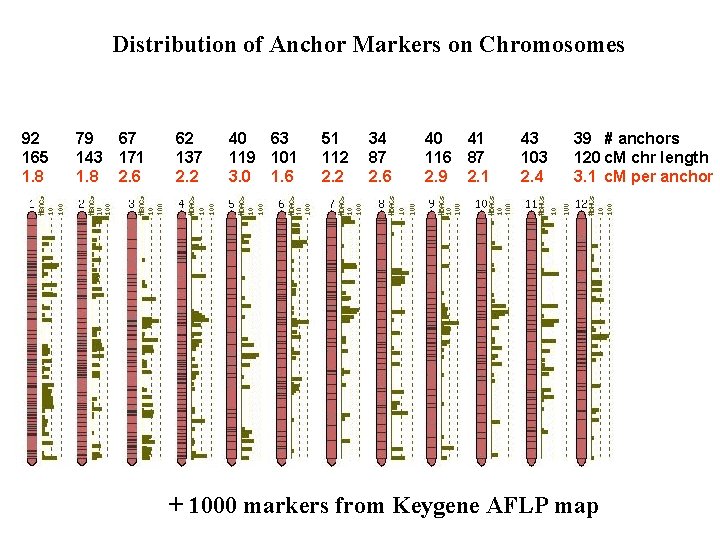
Overgo Anchoring Results Anchors: • 652 anchor markers are involved in plausible non-conflicted associations with BACs. • 4857 good marker--BAC associations FPC contigs: • 1880 BACs in 705 plausible contigs • 2166 BAC singletons • 652 seed BACs ==> 1/3 of euchromatic genome sequence

Distribution of Anchor Markers on Chromosomes 92 165 1. 8 79 67 143 171 1. 8 2. 6 62 137 2. 2 40 63 119 101 3. 0 1. 6 51 112 2. 2 34 87 2. 6 40 41 116 87 2. 9 2. 1 43 103 2. 4 39 # anchors 120 c. M chr length 3. 1 c. M per anchor + 1000 markers from Keygene AFLP map

Verification of overgo mappings • Fluorescence In-Situe Hybridization (FISH) – BAC probe on pachytene chromosomes • IL lines (Zamir lab) – Map BACs to IL lines – CAPS assays

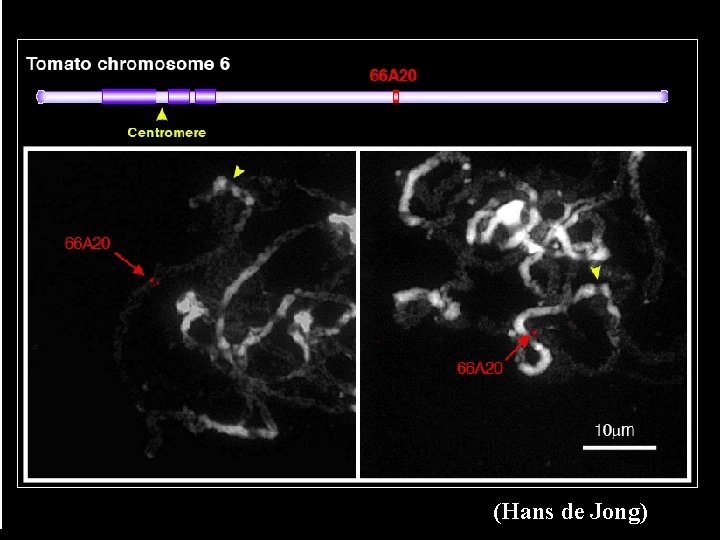
(Hans de Jong)

Summary of FISH verification • Song-Bin Cheng, Hans de Jong (Holland, chromosome 6): – 9 BACs analyzed – 8 mapped to chromsome 8 in right order – 1 BAC gave signals on centromere of chromosome 1 • Sangheob Lee, Doil Choi (Korea, chromosome 2): – 27 BACs analyzed with FISH – 25 confirmed to specific location, same order as F 2 -2000 map – 2 match to other chromosomes • Chuanyou Li (China, chromosome 3) – >30 BACs being analyzed • Steven Stack (USA): – Telomere and heterochromatic boundary determination – FISH service for countries without FISH capability

BAC end sequences • Total of 400, 000 reads (200, 000 BACs from both ends) selected from the 3 BAC libraries • Batch of 75, 000 BACs in process (Hind. III library) • ~45, 000 BAC end sequences already obtained (ftp: //ftp. sgn. cornell. edu/tomato_genome/) • Average read length 655 bp • Annotation in progress • Seq. Wright Inc, Houston, TX • Seq. Wright is sponsoring a happy hour after this session.

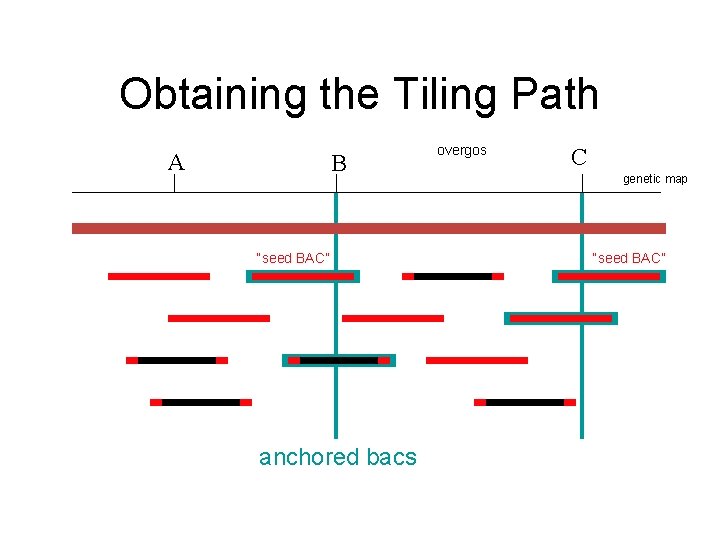
Obtaining the Tiling Path A B “seed BAC” anchored bacs overgos C genetic map “seed BAC”

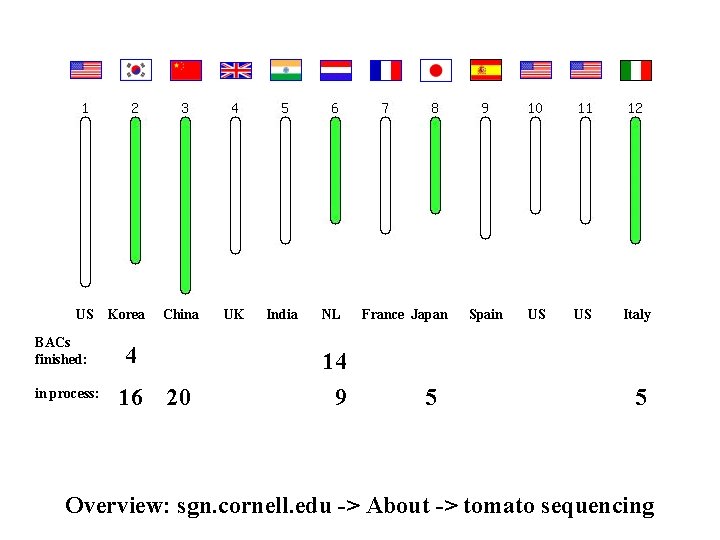
US BACs finished: in process: Korea China 4 16 20 UK India NL 14 9 France Japan 5 Spain US US Italy 5 Overview: sgn. cornell. edu -> About -> tomato sequencing

Building a Bioinformatics Platform for the Solanaceae • Project-wide standards for quality, gene naming, annotation (http: //sgn. cornell. edu/solanaceae-project/) • Create a unified web presence for the entire project • Develop distributed model for annotation, web presentation, involving different centers in SOL countries • All data and programs developed in the project are shared in an open source format • Integrate all data into the SOL bioinformatics platform, facilitating a systems approach to explore diversity and adaptation and the complex interactions that occur on all levels of biological organization

CAS Genome India SGN Agronanotech Kazusa VIB Ghent

Annotation Phases 1. First pass annotations of sequences and gene models on BAC basis, available immediately 2. BAC based, common, distributed platform, stable BAC-based identifiers 3. Chromosome based, stable identifiers

Summary • Sequencing of tomato is under way by a consortium of 10 countries • High quality, ordered sequence using BAC tiling path • BAC ends available, overgo results verified by FISH analyses • Sequence will be tied to other Solanaceae and closely related species (coffee and beyond) • Provide a foundation for shared biology for this economically important clade of plants

Acknowledgments SOL community Tomato Sequencing Project Funding National Science Foundation Other National Funding Sources Keygene NV Seqwright Inc. (Happy Hour) Colleagues Steven Tanksley, Jim Giovannoni, Joyce van Eck , Steven Stack SGN: Teri Solow, Beth Skwarecky, Nick Taylor, Robert Buels, John Binns, Chenwei Lin

