The Interdisciplinary Center for Biotechnology Research An Overview

The Interdisciplinary Center for Biotechnology Research – An Overview 2018 Pathology Research Infrastructure Talk April 11, 2018 Steven J. Madore, Ph. D. ICBR Associate Director for Science UFHCC Associate Director for Core Technologies

Talk Outline § The ICBR – an introduction and historical overview § Administrative Functions & Accessing ICBR Services § Technologies & Expertise – the ICBR Cores

Introduction § Founded in 1987 by the Florida Legislature as a research support organization to develop biotechnology resources to support the UF research community. § State of Florida and UF provide funding to support administration, scientific directors and technical personnel. § Main facility located in Cancer & Genetics Research Complex with satellite locations in Mac. Knight Brain Institute and Microbiology and Cell Sciences. § Mission is to enable, strengthen and energize all aspects of molecular life science research by providing scientific and technical instrumentation and expertise. § Long term aim is to jump start research for technology transfer and accelerate molecular biology research success from concept through to data that advances science. § Organized under the Vice President for Research, Dr. David Norton and led by Dr. Robert Ferl.

ü Ease of use ü Cost effective ü Impactful Your pursuit is what drives us.
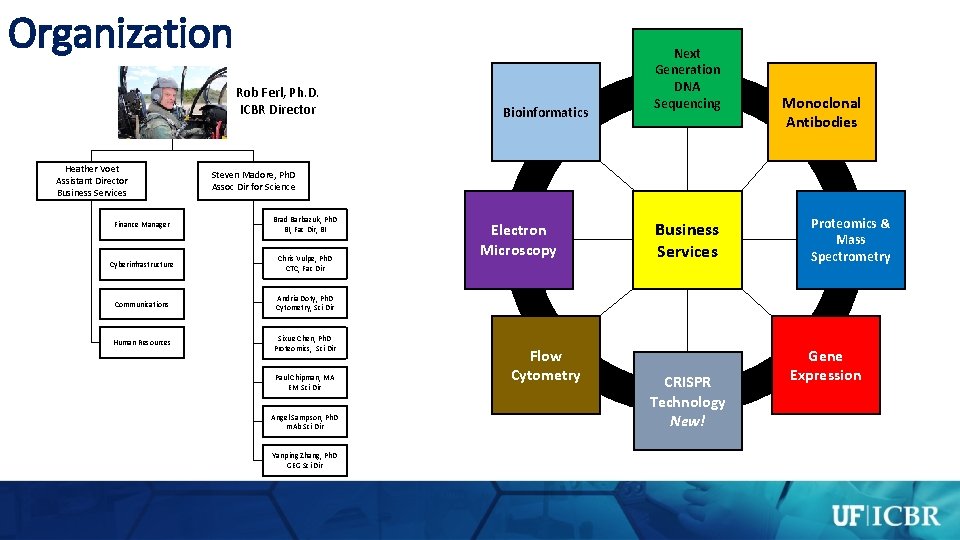
Organization Rob Ferl, Ph. D. ICBR Director Heather Voet Assistant Director Business Services Bioinformatics Next Generation DNA Sequencing Monoclonal Antibodies Steven Madore, Ph. D Assoc Dir for Science Finance Manager Brad Barbazuk, Ph. D BI, Fac Dir, BI Cyberinfrastructure Chris Vulpe, Ph. D CTC, Fac Dir Communications Andria Doty, Ph. D Cytometry, Sci Dir Human Resources Sixue Chen, Ph. D Proteomics, Sci Dir Paul Chipman, MA EM Sci Dir Angel Sampson, Ph. D m. Ab Sci Dir Yanping Zhang, Ph. D GEG Sci Dir Electron Microscopy Flow Cytometry Business Services CRISPR Technology New! Proteomics & Mass Spectrometry Gene Expression
Easy access Cost effective

Overall ICBR Customer Satisfaction 1 (“very dissatisfied”) 2 3 4 5 (“very satisfied”) Not sure Frequency 1 4 9 25 35 1 % (N=75) 1. 3% 5. 3% 12. 0% 33. 3% 46. 7% 1. 3% What did the ICBR do really well? Angel Sampson was great about getting information to us at the start Came up with good suggestions and helped interpret the results Communicate Communication (especially David Moraga) - Accommodate tight schedule. Communication when working through a project. Customer service, quality of training and data Easy requesting, timing EM core Expert advice on SEM use Fast service, constant updates, quality What could the ICBR do better? Better maintenance of instruments Bring back Sanger Sequencing Communication regarding technical issues Data analysis education Easier access/location to the college of medicine Fewer password requirements to access scheduling calendars

Talk Outline § The ICBR – an introduction and historical overview § Administrative Functions & Accessing ICBR Services § Technologies & Expertise – the ICBR Cores

Core Technologies – Electron Microscopy § Support researchers visualizing structures of microscopic samples –full service imaging projects –user training to facilitate solving research questions § Major service categories include: –cryo. EM, transmission and scanning electron microscopy –confocal laser scanning and epifluorescence microscopy –preparation of samples for microscopic analyses –training users to operate microscopes and ancillary equipment
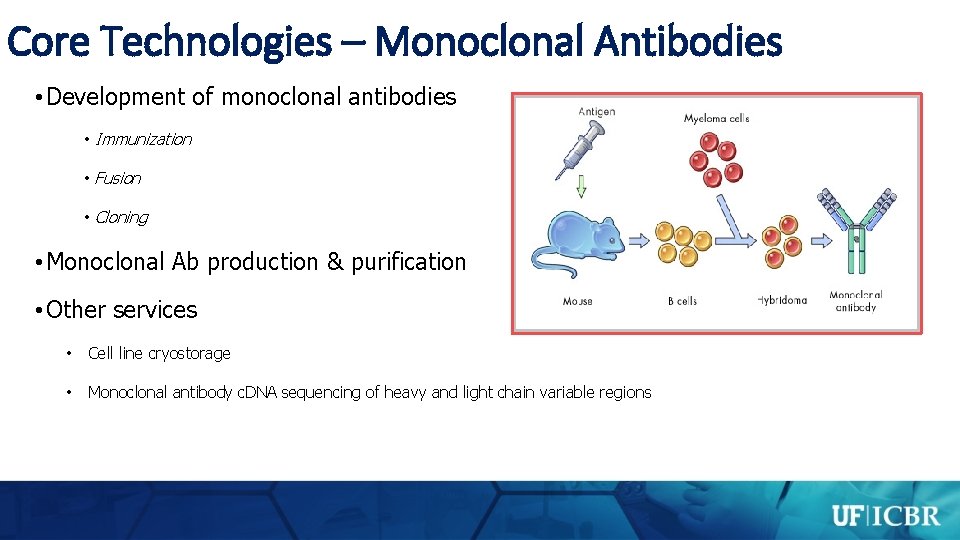
Core Technologies – Monoclonal Antibodies • Development of monoclonal antibodies • Immunization • Fusion • Cloning • Monoclonal Ab production & purification • Other services • Cell line cryostorage • Monoclonal antibody c. DNA sequencing of heavy and light chain variable regions
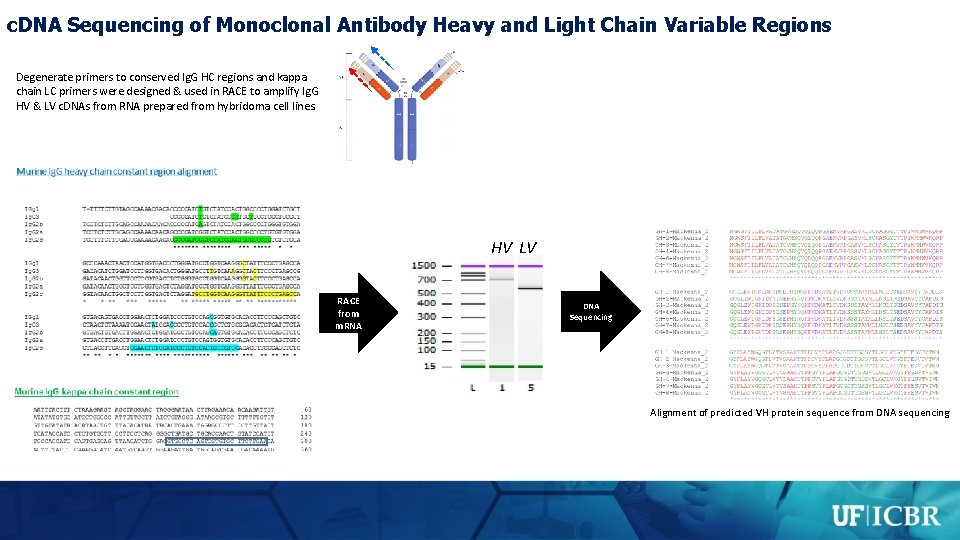
c. DNA Sequencing of Monoclonal Antibody Heavy and Light Chain Variable Regions Degenerate primers to conserved Ig. G HC regions and kappa chain LC primers were designed & used in RACE to amplify Ig. G HV & LV c. DNAs from RNA prepared from hybridoma cell lines. HV LV RACE from m. RNA DNA Sequencing Alignment of predicted VH protein sequence from DNA sequencing

c. DNA Sequencing of Monoclonal Antibody Heavy and Light Chain Variable Regions Derived from Cryo. EM, ICBR Mavis Agbandje-Mc. Kenna, Ph. D. Director, UF Center for Structural Biology

Core Technologies – Proteomics & Mass Spectrometry 1. Services and Technologies Protein fractionation Qualitative and quantitative mass spectrometry analysis de novo protein sequencing Post-translational modification analysis and accurate molecular weight determination o 2 -D gel electrophoresis and Western blotting o Protein enrichment and abundant protein depletion from biological fluids o Consultation, data processing and grant application support o o 2. Key equipment o Agilent 6495 Triple Quadrupole Mass Spectrometer coupled with the new Agilent 1290 UPLC

Core Technologies – Proteomics & Mass Spectrometry § Protein mixture is enzymatically digested into peptides § Peptide mix fractionated via HPLC § Analyzed by MS to generate spectrum § Spectrum compared to databases of known peptides to identify proteins in original mixture M stands for mass and Z stands for charge number of ions.

Isobaric Tags for Relative and Absolute Quantification (i. TRAQ) labeling • Labeling protein digests with isobaric tags allows many different biological samples to be mixed together in one analysis. • Enables simultaneous identification of proteins including relative and absolute quantitation. • Low molecular weight reporter ions from the mass tags are generated by fragmentation in the mass spectrometer and their intensity is measured, the ratios of which determine the relative or absolute amount of protein in each sample

Core Technologies – Flow Cytometry 1. Services and Technologies • • Multi-parameter flow cytometry and live cell sorting Training and Consultation; Courses and Workshops Confocal microscopy with live cell imaging Microparticle analysis using light scattering technology 2. Space and equipment • • 1400 sq. ft. - Cancer and Genetics Research Center, 682 sq. ft. -Mc. Knight Brain Institute Key equipment: o o Becton Dickenson LSR FORTESSA Flow Cytometer with 16 color 4 laser, 18 parameter analysis BD LSR II, CANTO II, Two - ACURI C 6 flow cytometers, three - ARIA II/III SONY SH 800 cell sorters 5 -laser scanning Leica ST 5 Confocal microscope with a live cell imaging incubation chamber Nano. Sight 300 microparticle analyzer Andria Doty, Ph. D. Scientific Director
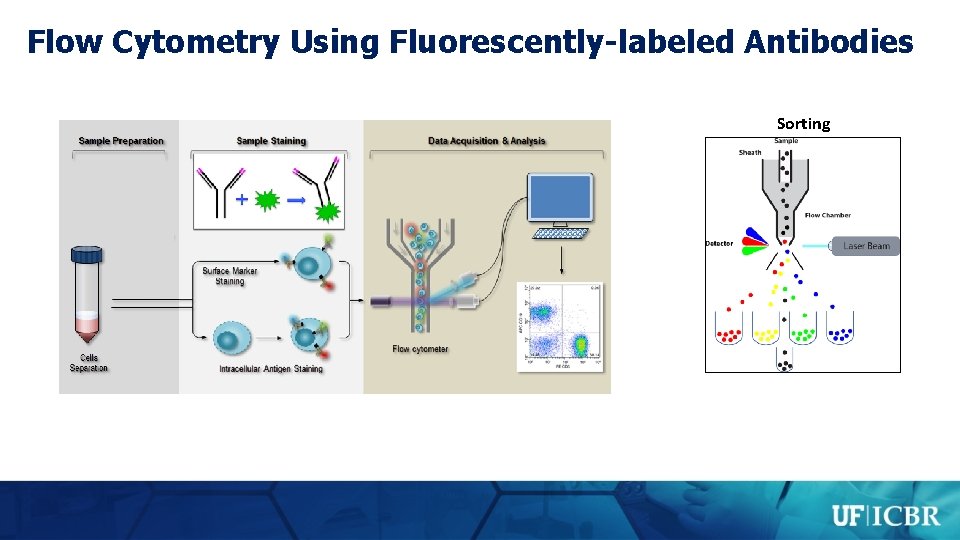
Flow Cytometry Using Fluorescently-labeled Antibodies Sorting
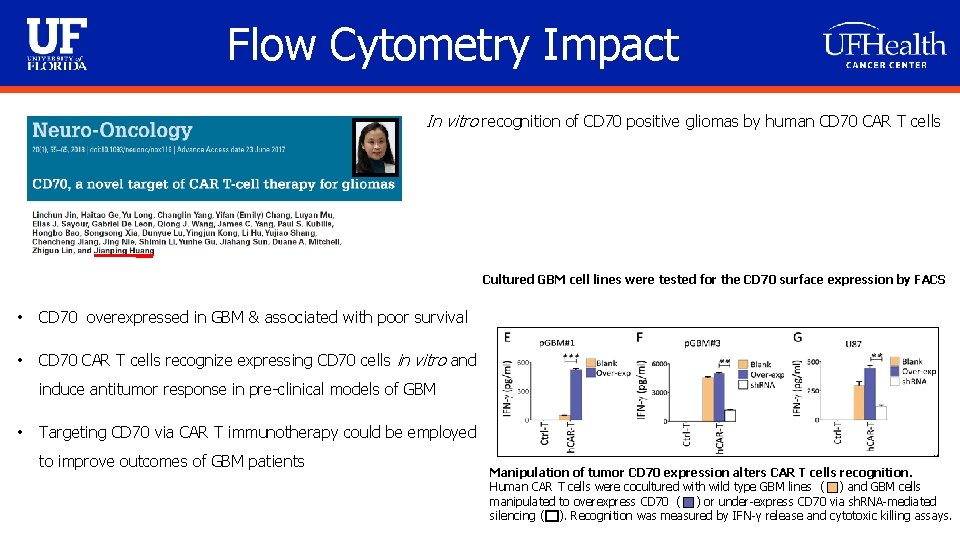
Flow Cytometry Impact In vitro recognition of CD 70 positive gliomas by human CD 70 CAR T cells Cultured GBM cell lines were tested for the CD 70 surface expression by FACS • CD 70 overexpressed in GBM & associated with poor survival • CD 70 CAR T cells recognize expressing CD 70 cells in vitro and induce antitumor response in pre-clinical models of GBM • Targeting CD 70 via CAR T immunotherapy could be employed to improve outcomes of GBM patients Manipulation of tumor CD 70 expression alters CAR T cells recognition. Human CAR T cells were cocultured with wild type GBM lines ( ) and GBM cells manipulated to overexpress CD 70 ( ) or under-express CD 70 via sh. RNA-mediated silencing ( ). Recognition was measured by IFN-γ release and cytotoxic killing assays.

Core Technologies – Gene Expression & Genotyping § RNA analysis § RNA qualitative/quantitative analysis § q. PCR and Droplet digital PCR § m. RNA splice variant analysis § mi. RNA and non-coding RNA analysis § RNASeq libraries § Whole transcriptome § Sequence capture libraries (amplicon or exome) § DNA genotyping § Mouse tail DNA genotyping by PCR § AFFYMETRIX AXIOM SNP genotyping § Microsatellite analysis § Human cell line authentication
Core Technologies - Bioinformatics § Transcriptome analysis § Metagenomics § Genomic analysis of complex mixture of microorganisms § Environmental samples, human gut microbiota § Taxonomical resolution using whole genome vs 16 S r. RNA sequencing § Next-generation sequence analysis § de novo genome assembly § SVs, CNVs § Statistical and functional analysis of microarray-based data § Proteomics data analysis

Core Technologies – Next Generation DNA Sequencing 1. Services and Technologies Experimental design consultation Illumina “short read” DNA sequencing technology Pac. Bio “long read” DNA sequencing technology NGS library construction (whole genome, exome) RNASeq Epigenomics 2. Space and Equipment • 3600 sq. ft. within the ICBR space in the Cancer and Genetics Research Complex (CGRC) • Key equipment includes: o o Illumina Mi. Seq (2), Next. Seq, and Hi. Seq 3000 DNA sequencers Pac. Bio SEQUEL DNA sequencer Dr. David Moraga, Scientific Director ICBR NGS Core
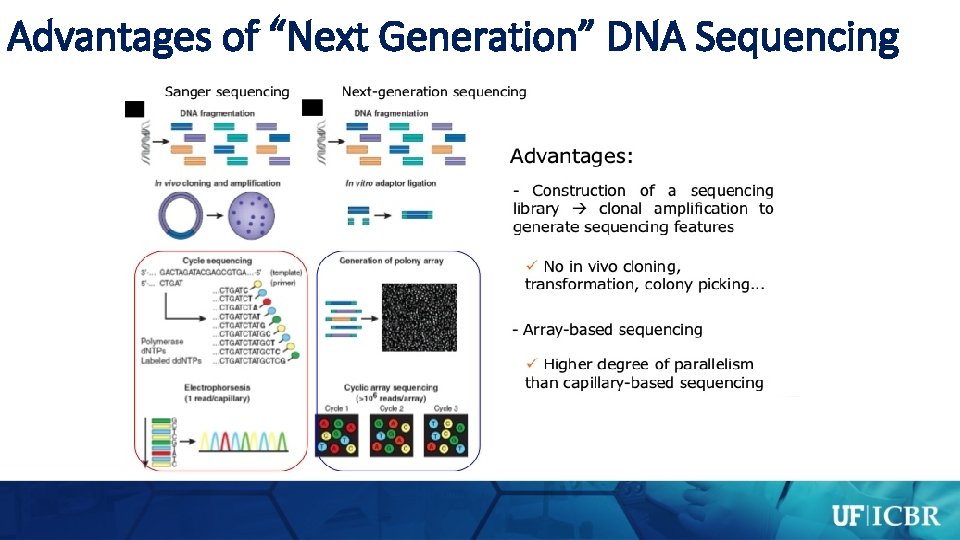
Advantages of “Next Generation” DNA Sequencing
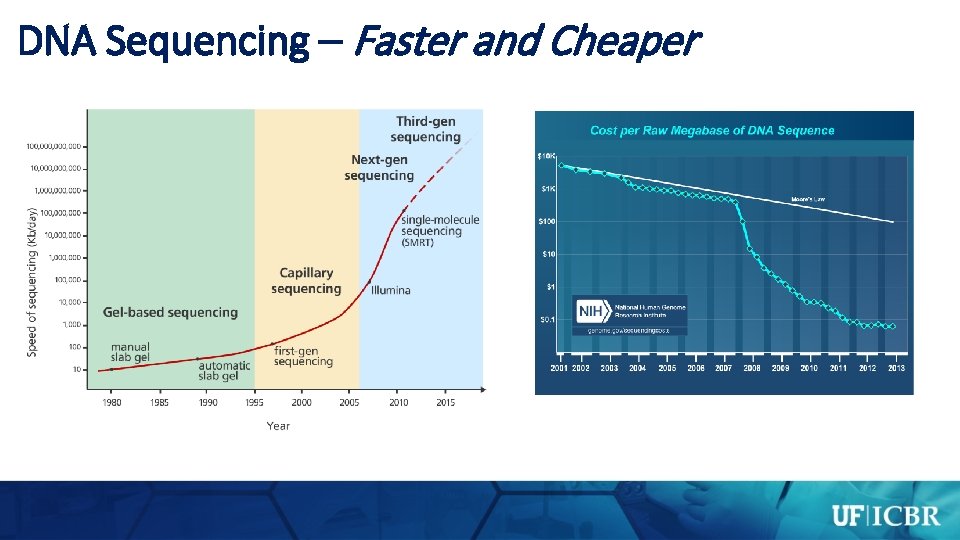
DNA Sequencing – Faster and Cheaper
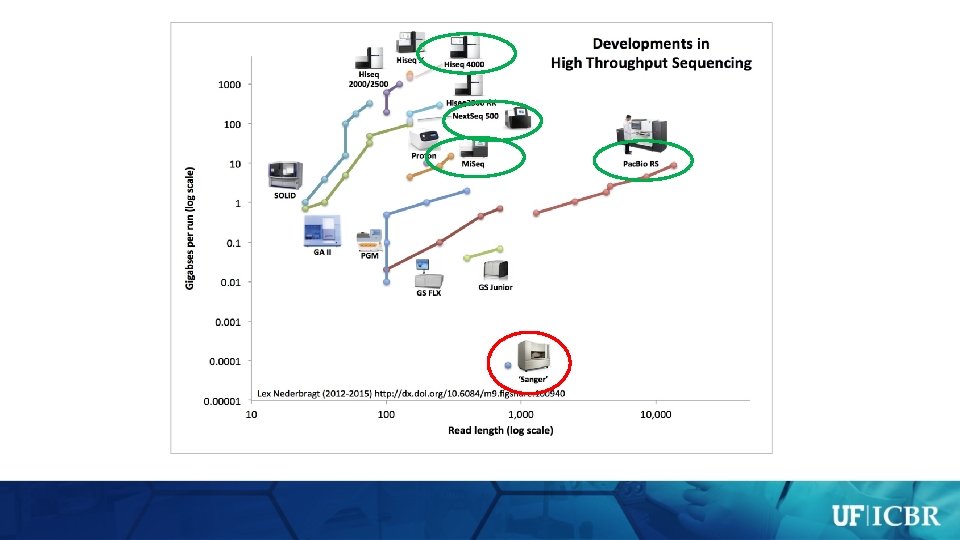
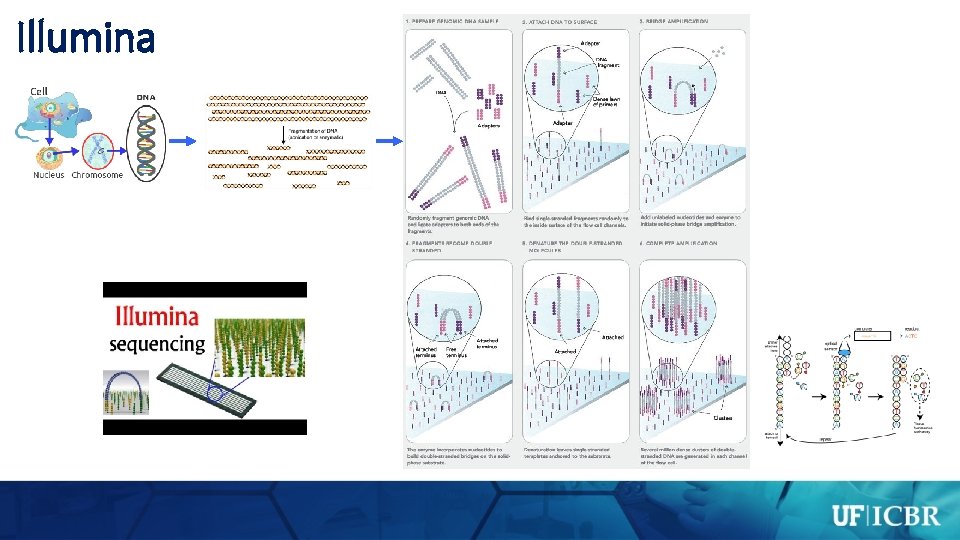
Illumina
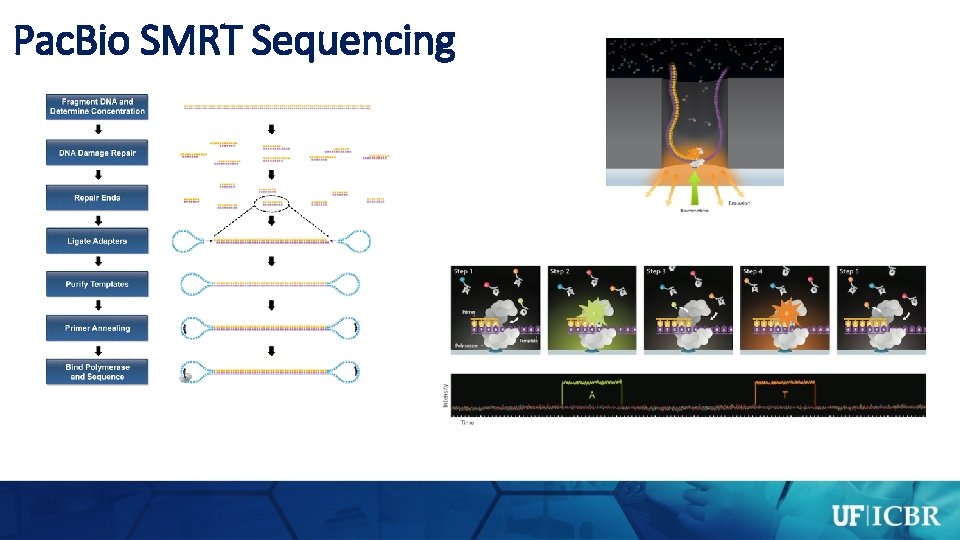
Pac. Bio SMRT Sequencing

NGS Applications § Whole genome sequencing § De novo genome assembly § Metagenomics § Targeted sequencing § Exome sequencing § Amplicon sequencing § Variant & structural analysis § 16 S r. RNA sequencing § Transcriptomics (RNASeq) § m. RNA expression levels § m. RNA splice variant analysis § Non coding RNAs (mi. RNAs, lnc. RNAs) § Epigenetics § DNA methylation § Nucleic acid-Protein analysis § Chromatin Immunoprecipitation (Ch. IP) § RNA – protein footprinting § Nucleosome footprinting § Clinical diagnosis § Tumor profiling § Gene polymorphisms § Fetal chromosome abnormalities § Pharmacogenomics

Sequencing Requires Shredding Chromosomal DNA…
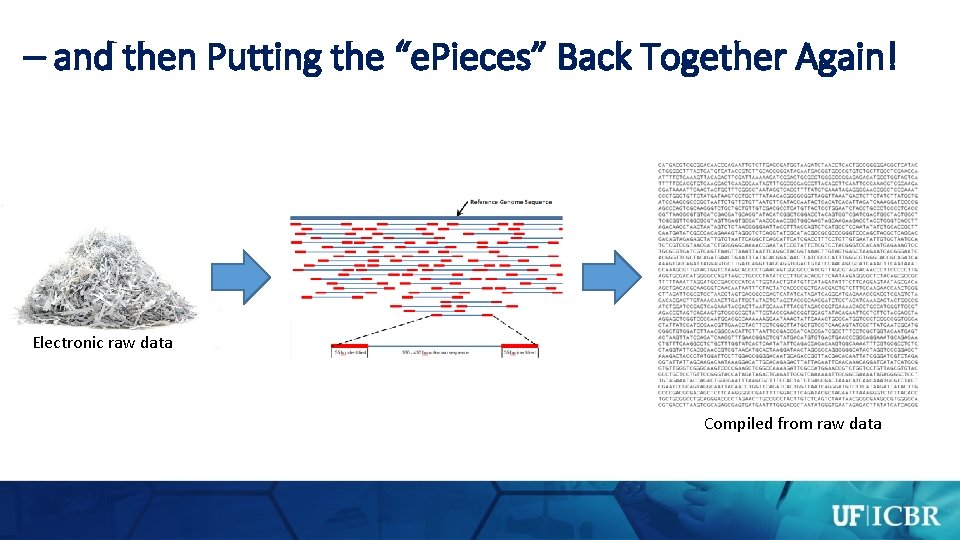
– and then Putting the “e. Pieces” Back Together Again! Electronic raw data Compiled from raw data
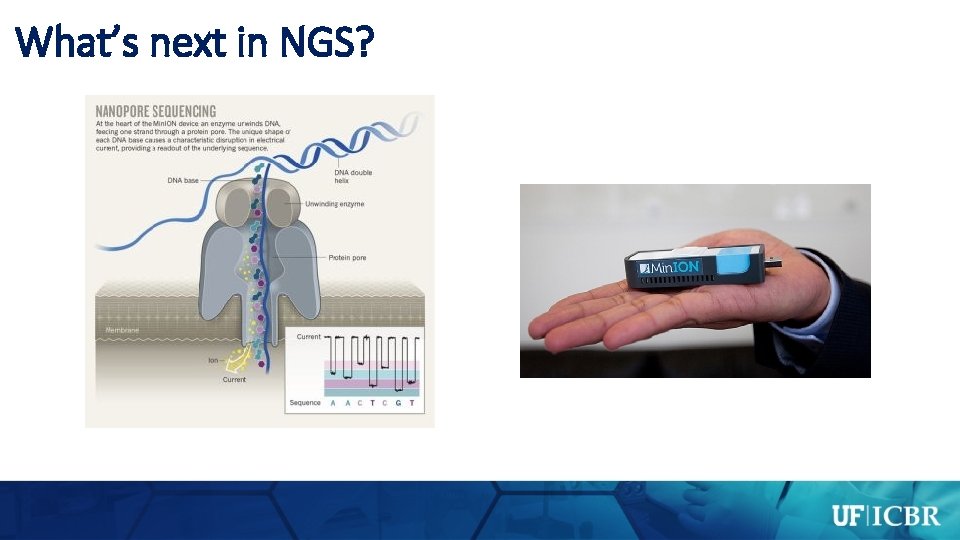
What’s next in NGS?

New ICBR Core – CRISPR Technology • Co-managed by UFHCC and ICBR • Led by Dr. Chris Vulpe, UFHCC Faculty Director • Clustered Regularly Interspaced Short Palindromic Repeats • Hallmark of a adaptive anti-phage immune system in bacteria first characterized in 1993 by Francisco Mojica, University of Alicante, Spain • Simple enzymatic process modified to easily work in mammalian cells • Gene editing in mammalian cells • Generation of cell pools or clonal cell lines with targeted allele editing • CRISPR-based whole genome and targeted screening

Gene Editing with CRISPR Cas 9 g. RNA Puromycin R

QUESTIONS? §Thank you!
- Slides: 33