The Human Genome 300000 bases The raw data
The Human Genome 300000 bases

The raw data NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN GATCTGATAAGTCCCAGGACTTCAGAAGagctgtgagaccttggccaagt cacttcctccttcag. GAACATTGCAGTGGGCCTAAGTGCCTCCTCTCGGG ACTGGTATGGGGACGGTCATGCAATCTGGACAACATTCACCTTTAAAAGT TTATTGATCTTTTGTGACATGCACGTGGGTTCCCAGTAGCAAGAAACTAA AGGGTCGCAGGCCGGTTTCTGCTAATTTCTTTAATTCCAAGACAGTCTCA AATATTTTCTTATTAACTTCCTGGAGGCTTATCATTCTCTCTTTTG GATGATTCTAAGTACCAGCTAAAATACAGCTATCATTTTCCTTGAT TTGGGAGCCTAATTTCTTTAATTTAGTATGCAAGAAAACCAATTTGGAAA TATCAACTGTTTTGGAAACCTTAGACCTAGGTCATCCTTAGTAAGATctt cccatttatataaatacttgcaagtagtagtgccataattaccaaacata aagccaactgagatgcccaaagggggccactctccttgcttttcctcctt tttagaggatttcccatttttcttaaaaaggaagaacaaactgtgc cctagggtttactgtgtcagaacagagtgtgccgattgtggtcaggactc catagcatttcaccattgagttatttccgcccccttacgtgtctctcttc agcggtctattatctccaagagggcataaaacactgagtaaacagctctt ttatatgtgtttcctggatgagccttcttttaattttgttaaggga tttcctctagggccactgcacgtcatggggagtcacccccagacactccc aattggccccttgtcacccaggggcacatttcagct. Atttgtaaaacctg aaatcactagaaaggaatgtctagtgacttgtgggggccaaggcccttgt tatggggatgaaggctcttaggtggtagccctccaagagaatagatggtg Aatgtctcttttcagacattaaaggtgtcagactctcagttaatctctcc tagatccaggaaaggcctagaaaaggcctgactgcattaatggaga ttctctccatgtgcaaaatttcctccacaaaagaaatccttgcagggcca ttttaatgtgttggccctgtgacagccatttcaaaatatgtcaaaaaata tattttggagtaaaatactttcattttccttcagagtctgctgtcgtatg atgccataccagagtcaggttggaaagtaagccacattatacagcgttaa cctaaaaaaactgtctaacaagattttatggtttatagagcat gattccccggacacattagaaatctgggcaagagaagaaaaaaagg tcagagtttaatcctca. TTCCTAAGTTAtgtaaaccaaaaattct gaagatgtcctgatcatctgaatggacccttcctctggaccagggcattc caaagttaacctgaaaattggtttgggccatgatgggaagggaggtttgg atatgcctcattatgccctcttccctttcagaattcaggaaaagccaacc agcattaacatcaacacagattttcagatcttaggtttccgatcta ttctctctgaaccctgctacctggaggcttcatctgcataataaaacttt agtctccacaaccccttatcttaccccagacattcctttctattgataat aactctttcaaccaattgccaatcagggtatgtttaaatctacctatgac ctggaagcccccactttgcaccctgagatcaaaccagtgcaaatcttata tgtattgatttgtc. AATGAAAACAGTCAAAGCCagtcaggcacagtggct catgcctgtaatcccagcactttgggaggctgaggcgggtagatcacctg aggtcaggagttcgacaccagcctggccaacatggtgaaaccccgtccct actaaaatacaaaaattagcccagcttggtggtgggcacctgtaatctta gctactgcagagactgaggcaggagaatcgcttgaacccaggaggtggag gttgcagtgacctgagattttgccattgcactccagcctgggcaacagag caagactctatctcaaaaaacaaacaaacaaac. T gtcaaaatctgtacagtatgtgaagagatttgttctgaaccaaatatgaa tgaccatggtccatgacacagccctcagaagaccctgagaacatgtgccc aaggtggtcacagtgcatcttagttttgtacattttagggagatatgaga cttcagtcaaatacatttttaaaaaatacattggttttgtccagaaagcc agaaccactcaaagcaggggtttccaggttataagtagatttaaaatttt tctgattgacaattggttgaaagagttgtcaatagaaaggaatgtctgca ttgtgacaagaggttgtggagaccaagtttctgtcatgcagatgaagcct tcaggtagcaggcttccaagataacaggttgtaaatagttcttatcagac ttaa. GTTCTGTGGAGACGTAAAATGAGGCATATCTGACCTCCACTTccaa aaacatctgagacaggtctcagttaagaaagtttgttctgcctagt ttaaggacatgcccatgacactgcctcaggaggtcctgacagcatgtgcc caaggtggtcaggatacagcttctatatattttagggagaaaatac atca. GCCtgtaaacaaaaaattctaaggtccctgaaccatctgaa tgggctttcttctaggccagggcactctaaaattgaagaacctgaacatt cctttctattgataatactttcagccagttgagcccattcaga. CCACAGC AAGGTGCCAGGCAAGGGCTGACTTGAGATACCTGCCAGATGAGTC ACTGGCAAAAGGTGCTGCTCCCTGGTGAGGGAGAAACACCAGGGGCTGGG AGAGGCCCAGAAGGCTCTGAAGGAGTTTTGGCTGGCCATGTGTGC AATTAGCGTGATGAGCTCTGACATGGCCTTGCATGGACGGATTGGGCAGG A’s T’s C’s and G’s and N’s
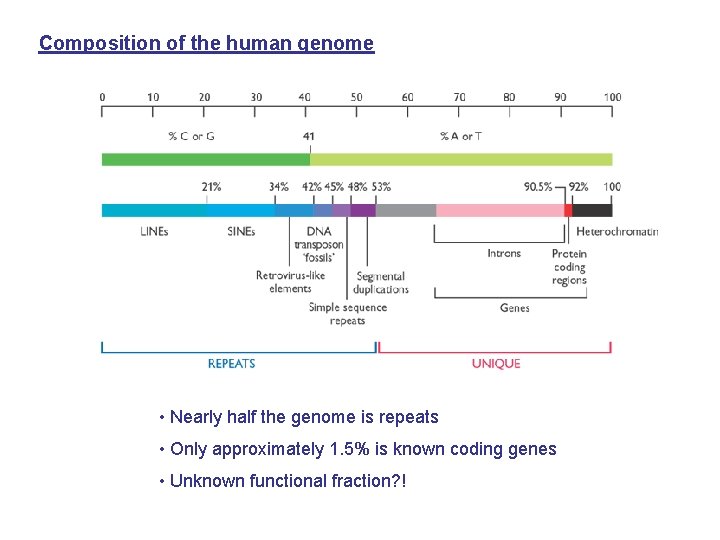
Composition of the human genome • Nearly half the genome is repeats • Only approximately 1. 5% is known coding genes • Unknown functional fraction? !
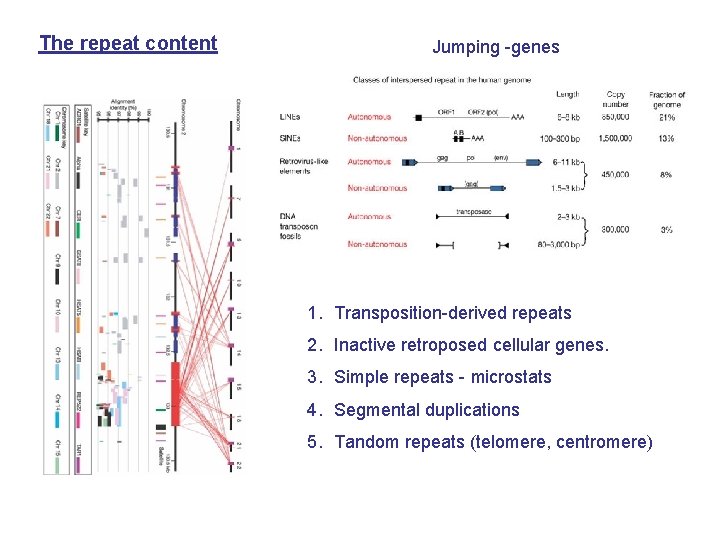
The repeat content Jumping -genes 1. Transposition-derived repeats 2. Inactive retroposed cellular genes. 3. Simple repeats - microstats 4. Segmental duplications 5. Tandom repeats (telomere, centromere)
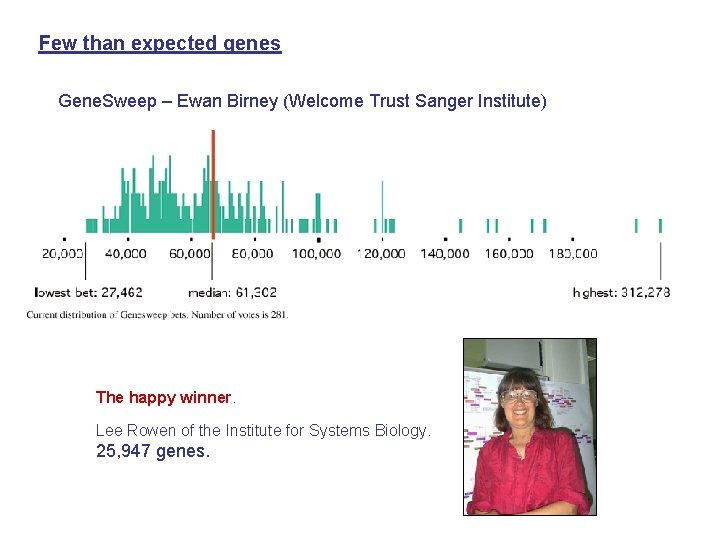
Few than expected genes Gene. Sweep – Ewan Birney (Welcome Trust Sanger Institute) The happy winner. Lee Rowen of the Institute for Systems Biology. 25, 947 genes.
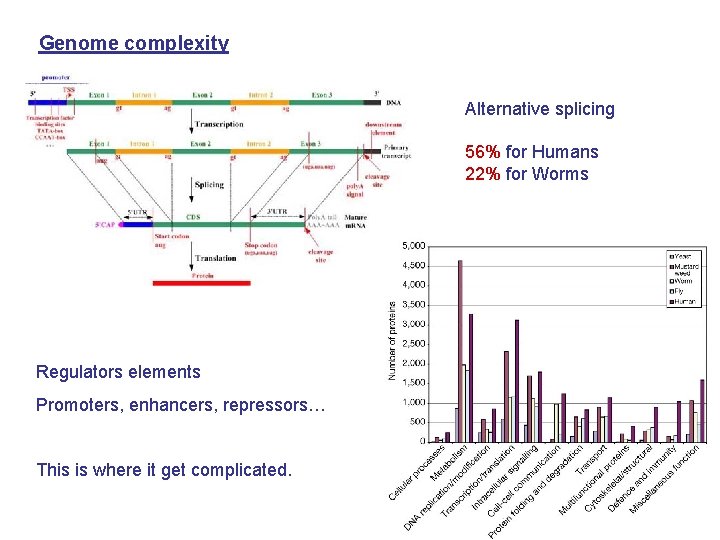
Genome complexity Alternative splicing 56% for Humans 22% for Worms Regulators elements Promoters, enhancers, repressors… This is where it get complicated.
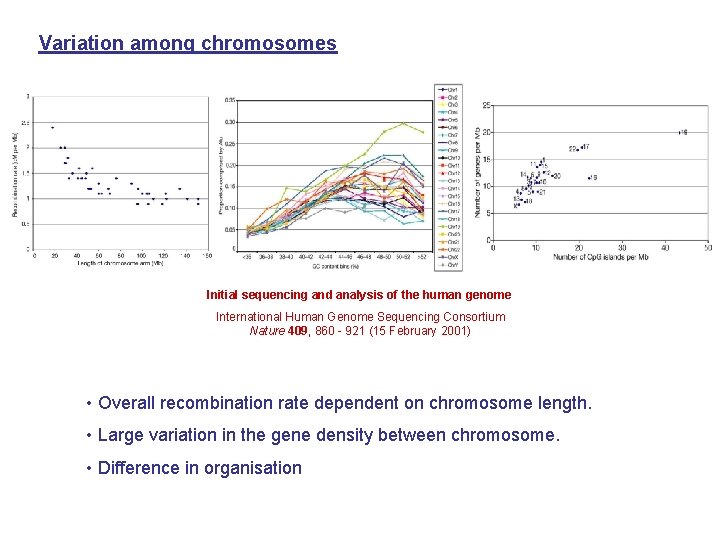
Variation among chromosomes Initial sequencing and analysis of the human genome International Human Genome Sequencing Consortium Nature 409, 860 - 921 (15 February 2001) • Overall recombination rate dependent on chromosome length. • Large variation in the gene density between chromosome. • Difference in organisation
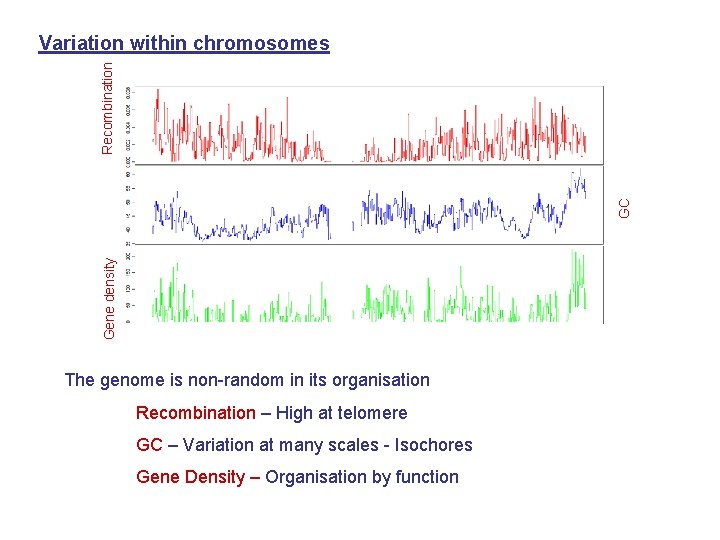
Gene density GC Recombination Variation within chromosomes The genome is non-random in its organisation Recombination – High at telomere GC – Variation at many scales - Isochores Gene Density – Organisation by function

New observations 2001 • Variation at multiple scales within and between chromosomes • Only twice as many genes as flies and worms – but more proteins • Genes have arrived from bacteria and transposable elements • Transposons inactive and LTR probably also (Alu’s in GC rich regions) • Most mutations occur in males (higher mutation rate) • GC poor regions correspond to dark bands. • Recombination rates are higher at telomeres • Lots of between individual variation

Completing the Human Genome Humans Genome Project starts 1990 Draft Human Genome completed 2001 Fewer gaps More continuity 147, 821 81 kb 341 38, 500 kb Gene rich regions completed 2003 • Error rate of ~1 in per 100, 000 bases • 2. 85 billion bases • Covers ~99% of the euchromatic genome. Each chromosome compiled annotated. Go home? 2006!
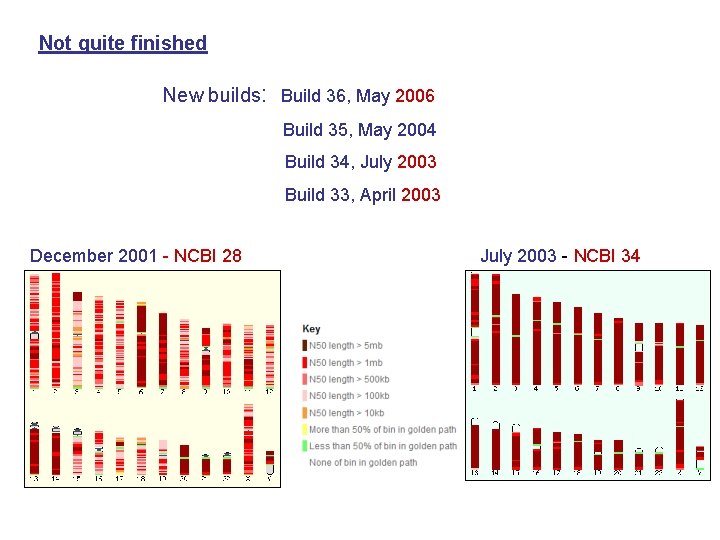
Not quite finished New builds: Build 36, May 2006 Build 35, May 2004 Build 34, July 2003 Build 33, April 2003 December 2001 - NCBI 28 July 2003 - NCBI 34
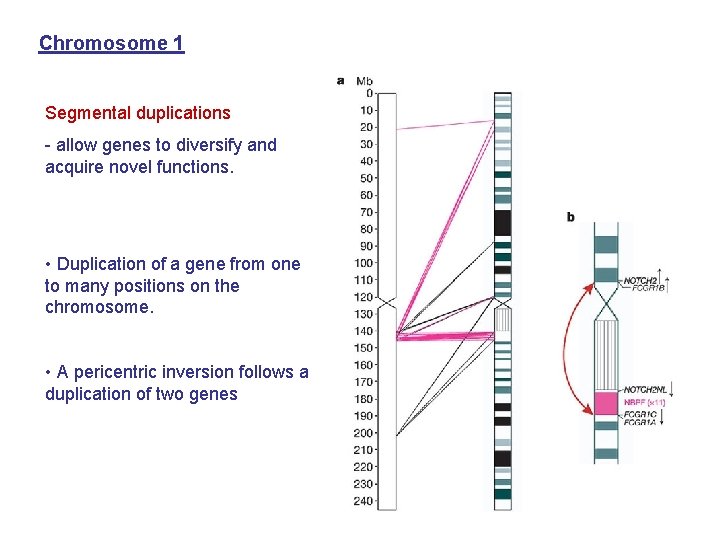
Chromosome 1 Segmental duplications - allow genes to diversify and acquire novel functions. • Duplication of a gene from one to many positions on the chromosome. • A pericentric inversion follows a duplication of two genes
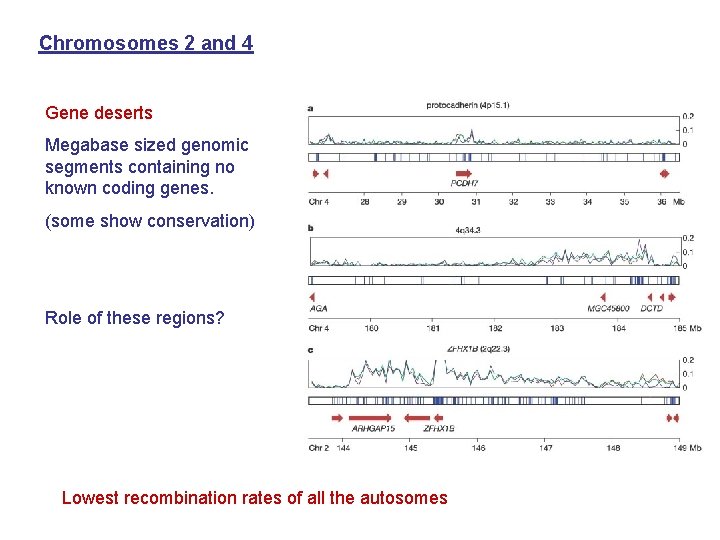
Chromosomes 2 and 4 Gene deserts Megabase sized genomic segments containing no known coding genes. (some show conservation) Role of these regions? Lowest recombination rates of all the autosomes
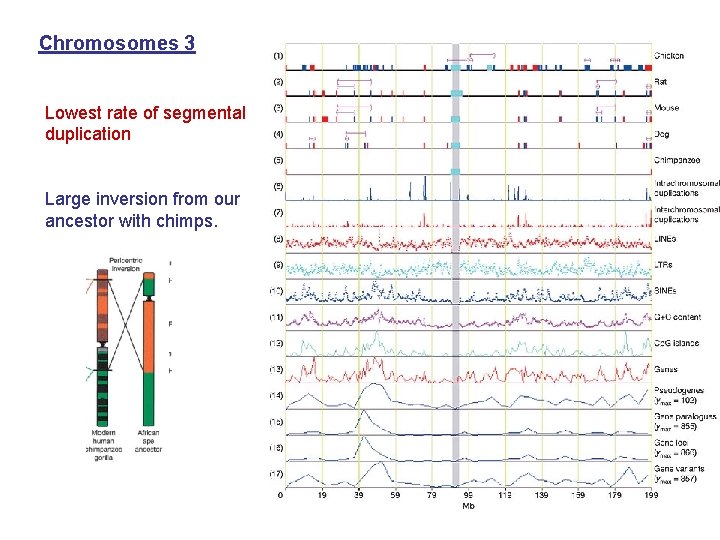
Chromosomes 3 Lowest rate of segmental duplication Large inversion from our ancestor with chimps.

Chromosomes 7 Complex repeat patterns and fragile locations Williams-Beuren syndrome associated with a large deletion (1. 6 Mb). Lots of repetitive and duplicated DNA. What is the true sequences? “It is characterized by a distinctive, "elfish" facial appearance, along with a low nasal bridge; an unusually cheerful demeanor and ease with strangers, coupled with unpredictably occurring negative outbursts; mental retardation coupled with an unusual facility with language; a love for music; and cardiovascular problems, such as supravalvular aortic stenosis and transient hypercalcemia. ”

Chromosomes 10 Multi-species alignment – gene involved in cancer Conservation indicates the location of functional elements. Some are known genes. Others aren’t – higher levels of conservation!
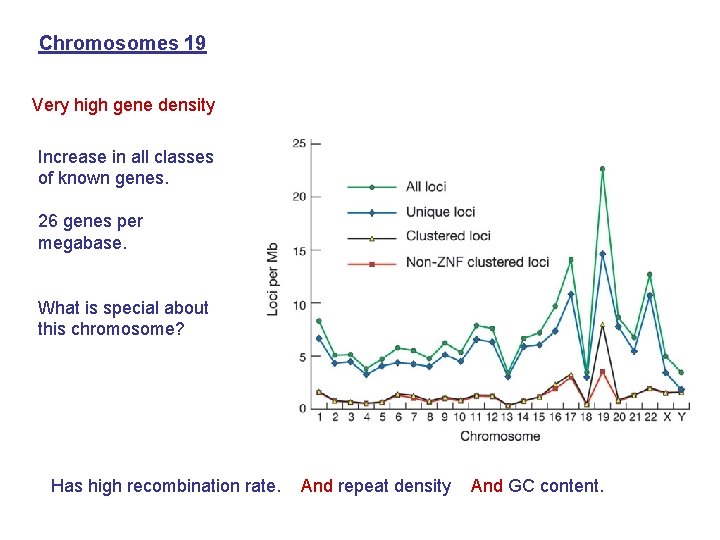
Chromosomes 19 Very high gene density Increase in all classes of known genes. 26 genes per megabase. What is special about this chromosome? Has high recombination rate. And repeat density And GC content.
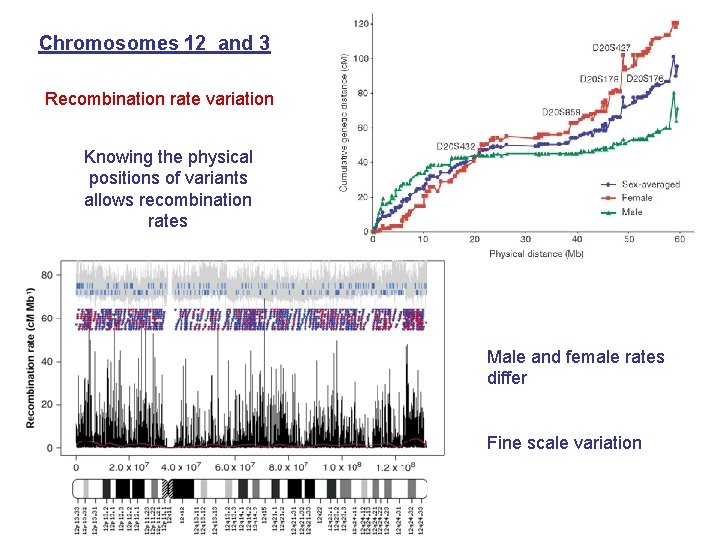
Chromosomes 12 and 3 Recombination rate variation Knowing the physical positions of variants allows recombination rates Male and female rates differ Fine scale variation

Where is the data available N. C. B. I. www. ncbi. nlm. nih. gov/genome/guide/human/ • Part of the National Institute of Health. • Has a number of important associated projects. • Mr NCBI – David Lipman. Ensembl www. ensembl. org/Homo_sapiens/ • A joint project between EMBL and the Sanger Institute. • Primarily funded by the Welcome Trust. • Mr Ensembl – Ewan Birney UCSC genome. ucsc. edu/cgi-bin/hg. Gateway • Based at the University of California Santa Cruz. • Largely funded by the NHGRI. • Mr UCSC – David Hassler

What data available • Compositional Base composition Insertion deletions Segmental duplications Repeats Transposable elements • Functional Genes Regulatory elements Gene expression • Evolutionary Species comparison Variation data Population genetic analysis Use drop down controls below and press refresh to alter tracks displayed. Tracks with lots of items will automatically be displayed in more compact modes. Mapping and Sequencing Tracks Base Chromoso STS FISH Recomb Positio me Band Markers Clones Rate n Map Contigs Assembly Gap Coverage BAC End Pairs Fosmid End Pairs GC Percent WSSD Duplicatio n Short Match Restr Enzymes Phenotype and Disease Associations RGD Human QTL Mutation Genes and Gene Prediction Tracks Known Genes CCDS Ref. Seq Genes Other Ref. Seq MGC Genes Vega Pseudogen es Ensembl Genes Ace. View Genes ECgene Genes NSCAN SGP Genes Geneid Genes Genscan Genes Exoniphy August us Genes Retropose d Genes Superfami ly Yale Pseudo Evo. Fold sno/mi RNA Exon. Walk m. RNA and EST Tracks Human m. RNAs Spliced ESTs Human ESTs Other m. RNAs Other ESTs H-Inv TIGR Gene Index Uni. Gene Bounds Alt-Splicing Affy Hu. Ex 1. 0 Affy U 133 Expression and Regulation Allen GNF Atlas GNF Ratio Brain 2

Orientation • Human chromosomes are numbered • Arms are labelled p and q • Regions labelled ascending from centromere. • Bases numbered from beginning of small arm to end of long arm.

Annotation - Repeats Transposable elements • Make up a large proportion of the genome Microsatellites and repeats • Important in many common diseases • Some of the most polymorphic loci

Annotation - genes • Different levels of evidence for genes m. RNA evidence • Based on homology • Based on expression • Based on prediction Protein evidence Gene prediction EST evidence Predicted transcripts - Known Novel Manually annotated genes
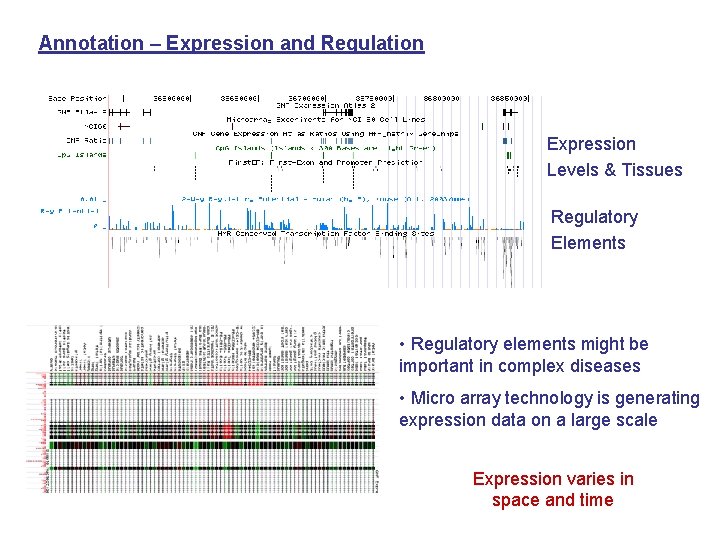
Annotation – Expression and Regulation Expression Levels & Tissues Regulatory Elements • Regulatory elements might be important in complex diseases • Micro array technology is generating expression data on a large scale Expression varies in space and time
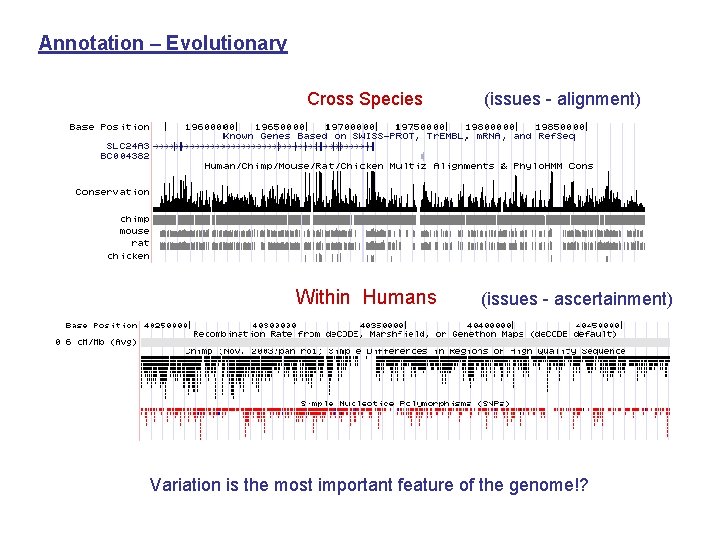
Annotation – Evolutionary Cross Species Within Humans (issues - alignment) (issues - ascertainment) Variation is the most important feature of the genome!?

Encylopedia of DNA Elements - Encode 1% of genome 14 manually chosen regions (Alpha & beta globin, HOXA, FOXP 2 and CFTR) Plus 26 random regions • Variation group – SNPs indels • Function group – Promoters, transcription and binding • Chromatin group – Chromatin modification, replication origins • Multiple sequence alignment – Conservation vs Constraint Aim: Understand everything possible about these regions.
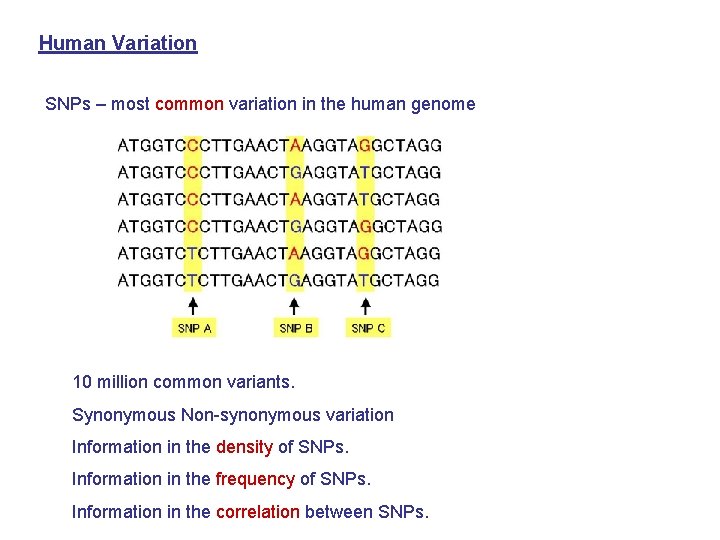
Human Variation SNPs – most common variation in the human genome 10 million common variants. Synonymous Non-synonymous variation Information in the density of SNPs. Information in the frequency of SNPs. Information in the correlation between SNPs.

Hap. Map Project 2002 Hap. Map phase I begins § Three populations § § (YRI) Yoruba in Ibadan, Nigeria § (CEU) Utah, USA § (CHB) Han Chinese in Beijing § (JPT) Japanese in Tokyo Approximately 1 million SNPs 2005 90 90 45 44 Phase I complete, phase II begins § Increase from 1 million to ~ 4. 6 million 2006 Phase II complete, “phase III” begins § Additional 6 populations § Kenya, African Americans, Mexican Americans, Italy, India
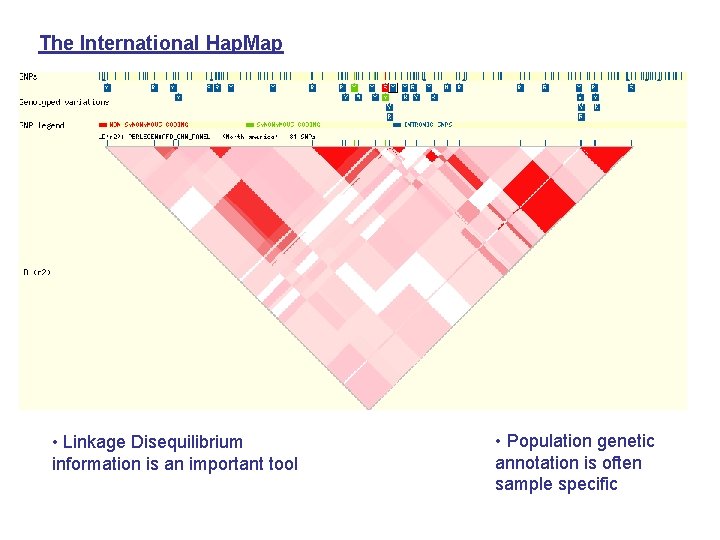
The International Hap. Map • Linkage Disequilibrium information is an important tool • Population genetic annotation is often sample specific
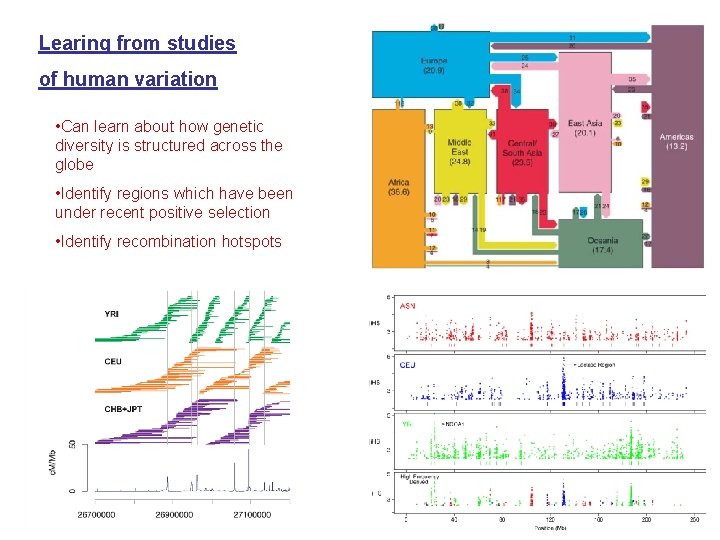
Learing from studies of human variation • Can learn about how genetic diversity is structured across the globe • Identify regions which have been under recent positive selection • Identify recombination hotspots
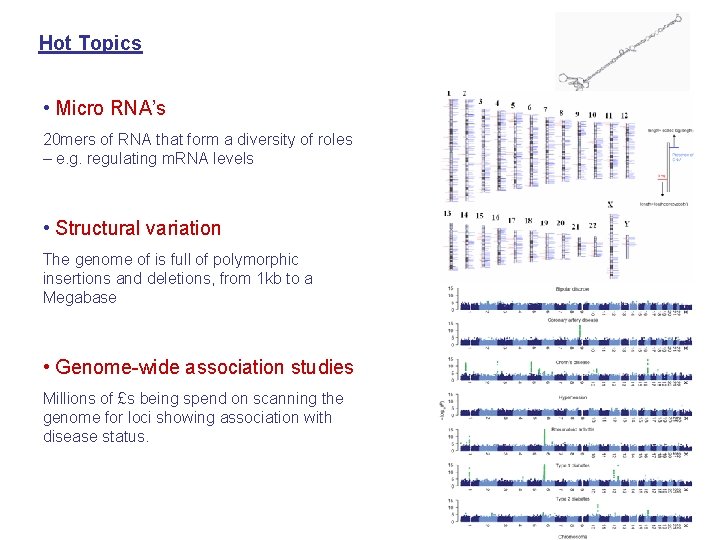
Hot Topics • Micro RNA’s 20 mers of RNA that form a diversity of roles – e. g. regulating m. RNA levels • Structural variation The genome of is full of polymorphic insertions and deletions, from 1 kb to a Megabase • Genome-wide association studies Millions of £s being spend on scanning the genome for loci showing association with disease status.
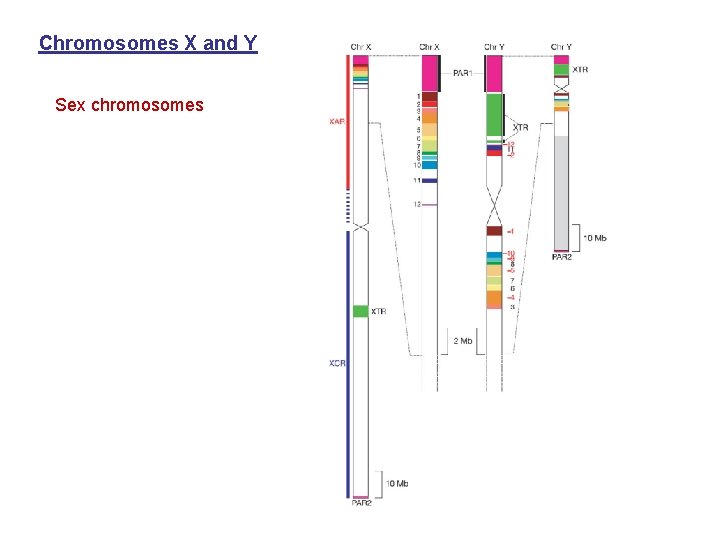
Chromosomes X and Y Sex chromosomes
- Slides: 32