The Genes the Whole Genes and Nothing But

The Genes, the Whole Genes, and Nothing But the Genes Jim Kent University of California Santa Cruz

Ben Franklin - Childhood Hero Hi Voltage Experiments

A Man of High Values

Early to bed Early to rise

Rock Collection

Shell Collection

Bottlecap Collection

Bug Collection

Jim Kent Genome Scientist (not to be confused with Richard Stallman)

Modern Bug Collection if (a = b) if (string == “something”) for (x=0; x<count; ++x); process(x); for (x=0; x<width; ++x) for (y=0; y<height; ++x) plot(x, y, data[x][y]);

Naive Biological Questions • • Is an ant an individual or is it the hive? Do dolphins talk with each other? How do amphibians and worms regenerate? How does an animal develop out of an egg?
From Egg to Adult in 3 x 109 Bases • A single cell, the fertilized egg, eventually differentiates into the ~300 different types of cells that make up an adult body. • With a few exceptions all of these cells contain the full human genome, but express only a subset of the genes. • Gene expression patterns are determined largely by the cell type, and vice versa.
From Totempotency to Senility • Human cells become more and more specialized during development • An egg can become anything. (Initially most of it will become placenta & amnion). • Liver cells only become liver cells. • Neurons can’t even reproduce.

Cell Type Determinants During Development • Cell type of parent cell. • Interactions with other cells. • Interactions with the extracellular environment.
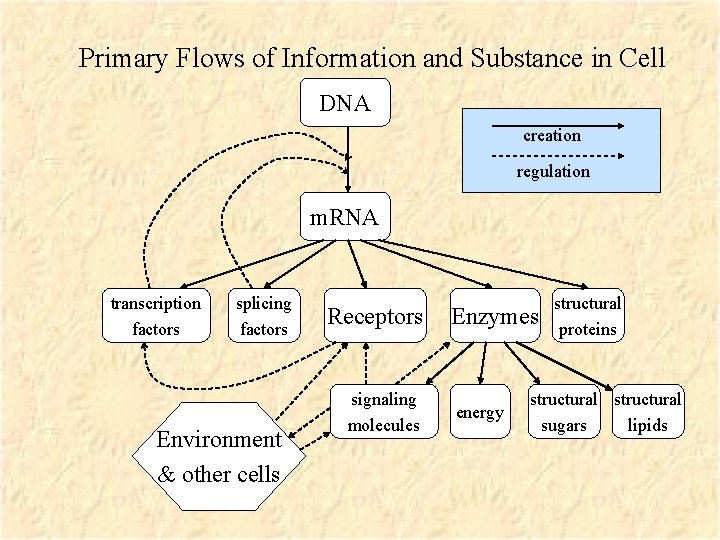
Primary Flows of Information and Substance in Cell DNA creation regulation m. RNA transcription factors splicing factors Environment & other cells Receptors signaling molecules Enzymes energy structural proteins structural sugars lipids

An Extreme Case of Dedifferentiation • The cloning of Dolly the sheep showed that a differentiated genome could be reset. • An egg is huge compared to a normal cell. • Putting a normal cell into an egg as Wilmut et al did, swamps out the normal cell transcription factor and receptors with egg transcription factors and receptors. • Cloning success rate sometimes improved by passing a nucleus through multiple eggs.

Regeneration by Nature • Among vertebrates only amphibians can regenerate limbs. • The process involves dedifferentiation, repatterning, and growth. • Not likely we’ll be able to engineer this soon. • Simpler regenerations though may be tractable and medically quite important.
Human Diseases Involving a Small Population of Cells • Parkinson’s - from the death of dopamine producing neurons in the substantia negra. • Macular degeneration - a leading cause of blindness in the elderly. • Type I Diabetes - from the death of insulinproducing cells in the pancreas.
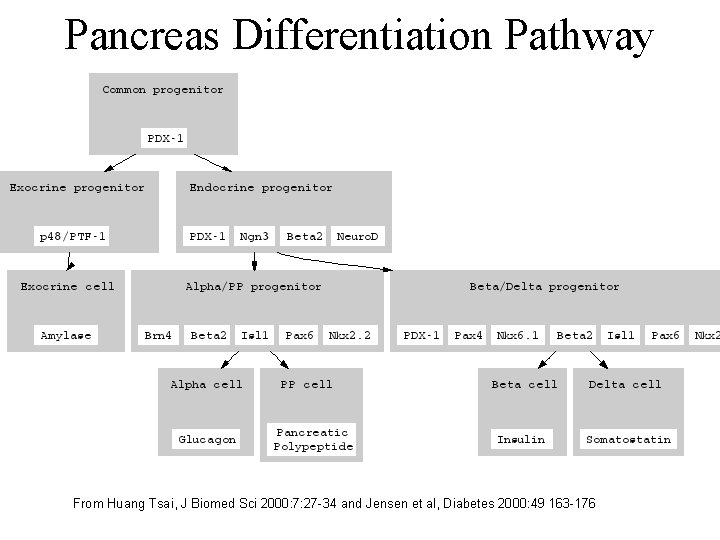
Pancreas Differentiation Pathway From Huang Tsai, J Biomed Sci 2000: 7: 27 -34 and Jensen et al, Diabetes 2000: 49 163 -176

Flexibility of Stem Cells • In many cases stem cells are flexible enough that putting them into a particular tissue will cause them to differentiate into the type of cells that make up that tissue. • At low levels bone transplanted bone marrow (blood stem cells) develops into neurons in stroke victims! • Making this happen at high enough levels to be useful will likely require some engineering.

To Understand the Body Need • • The genome A comprehensive list of genes Gene expression data Protein localization in cell Protein/protein and protein/DNA interaction information. Ways to store, display and query masses of data so human investigators can focus on relevant bits. Many human investigators.

Where are we now? • • • The genome >95% complete. 98% complete in April. A comprehensive list of genes - ~75% of coding regions. <50% of transcription start sites. Gene expression data - publically available on ~1/3 of genes. Protein localization in cell - very spotty. Computer predictions are about 75% accurate. Protein/protein and protein/DNA interaction information - just getting started.

The Genes • Identifying genes is a prerequisite for a great deal of other research. – Expression microarrays – In situ m. RNA hybridization – Producing proteins for cellular localization experiments – Etc.

The Whole Genes • The full gene including the 5’ and 3’ UTRs are critical for – Avoiding misleading fragmentation/fusion artifacts. – Understanding m. RNA targeting and stability – Finding transcription factor binding sites – Understanding the regulatory networks that drive and maintain cell differentiation.

Nothing But the Genes • Experimental analysis is expensive. • Unreal genes can mislead: – Analysis of multiple alignments to look for active sites etc. – Protein classification systems and phylogenies • One bogus gene can lead to another as much annotation is done via homology.

Methods of Identifying Genes • • • m. RNA/c. DNA sequencing Microarrays covering entire genome Genetics in model organisms Cross species protein homology Cross species genomic homology HMMs and other computational genefinding.

c. DNA Sequencing • Extract RNA from cells. • Use reverse transcriptase and a poly-U primer to convert to c. DNA starting at poly-A tail. • Insert c. DNA into vectors that grow in E. coli • Sequence a read from one or both sides of insert using primers on vector • If EST looks to be new sequence full c. DNA. • Artifacts and limitations are possible at each stage!

Common c. DNA Problems & Solutions • For rarely expressed genes little RNA is available. – Normalize libraries. Use embryonic and exotic tissues as m. RNA source. • Splicing is not instantanious, can get retained introns. – Spin out nuclei and just use cytoplasmic m. RNA – Align to genome and look for splicing • Reverse transcriptase falls off before it’s finished – Preferentially taking larger c. DNAs. – G-cap selected libraries (Sugano) – Normalizing only on 5’ ends (Soares)

More c. DNA Problems & Solutions • Reverse transcriptase has a high error rate and is prone to small deletions. – Compare c. DNA to genomic DNA – Sequence multiple c. DNA clones • At a low level cell seems to tolerate a certain degree of nonsense transcription and splicing. Normalizing increases concentration of these as well as of rare genes. – Ignore everything that’s not coding (ouch) – ? ? ?

c. DNA Status & Summary • ~10, 000 c. DNA sequence have been accumulated over years by various labs working on gene families and pathways. • Riken project has ~33, 000 unique c. DNAs in mouse. ~11, 000 of these seem to have retained introns. ~3, 000 are noncoding antisense. ~70% include initial ATG • Mammalian Gene Collection (MGC) has ~15, 000 human c. DNAs with initial ATGs. Having to resort to exotic libraries and RT-PCR to get more. • Human ref. Seq has ~18, 000 human c. DNAs.

Whole Genome Microarrays • Perlegen and Affymetrix are making microarrays that cover entire non-Repeat. Masker masked genome. Results on chromosome 21 and 22 published. • Based on 25 -mers. • Rarely expressed genes may not stand out above background. • Have to cope with cross-hybridization issues, GC content, etc. • Advantages - no homology required, can sense lower concentrations of m. RNA than random EST sequencing.
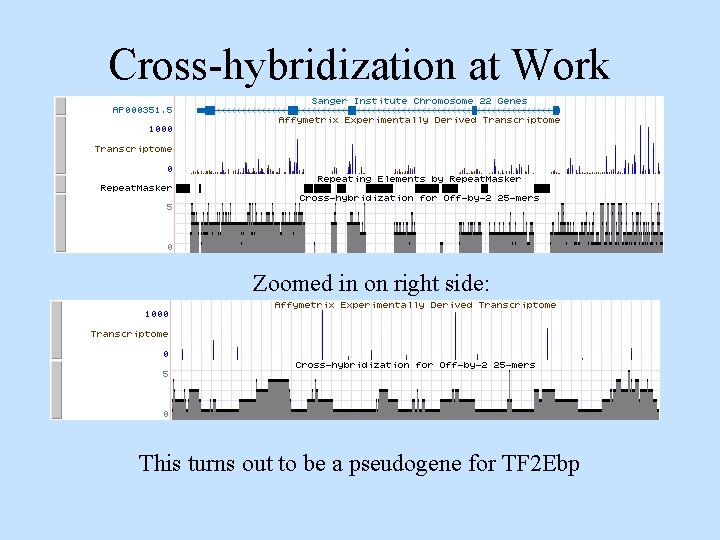
Cross-hybridization at Work Zoomed in on right side: This turns out to be a pseudogene for TF 2 Ebp

Model Organism Genetics • Zap hapless yeast, worms, flies, and mice • Inbreed offspring and look for twisted ones. • Advantages: – Works at DNA level, so expression level doesn’t matter – You get hints of function right away. – Can look for gene interactions simply by breeding mutants. • Disadvantages: – Finding which DNA is mutated can take a long time. – Essential genes can be hard to find - all you see is reduced fertility in the inbreeding stage. – Genes only needed in certain environments and duplicated genes may be missed in screens.

Comparative Genomics

Cross Species Genome Comparisons • Mutations occur more or less randomly across genome but • Mutations in functional areas tend to be weeded out by selection • In comparing DNA across species, the functional areas are more conserved than the nonfunctional areas in general • Caveat: ~5% of human genes don’t have clear mouse orthologs (though most do have paralogs).
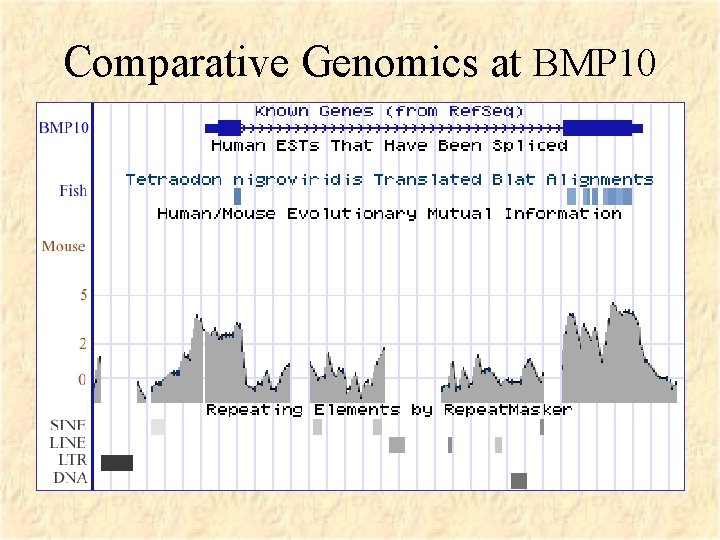
Comparative Genomics at BMP 10

Conservation of Gene Features Intron Mid CDS Intron Up 200 5’ UTR 1 st CDS last CDS 3’ UTR Down 200 Conservation pattern across 3165 mappings of human Ref. Seq m. RNAs to the genome. A program sampled 200 evenly spaced bases in introns, coding exons, and UTRs, and 200 bases upstream of transcription, and downstream of poly-adenlyation. There are peaks of conservation at the transition from one region to another.

Detail Near Translation Start Note the relatively conserved base 3 before translation Start (constrained to be a G or an A by the Kozak Consensus sequence, and the first three translated bases (ATG).
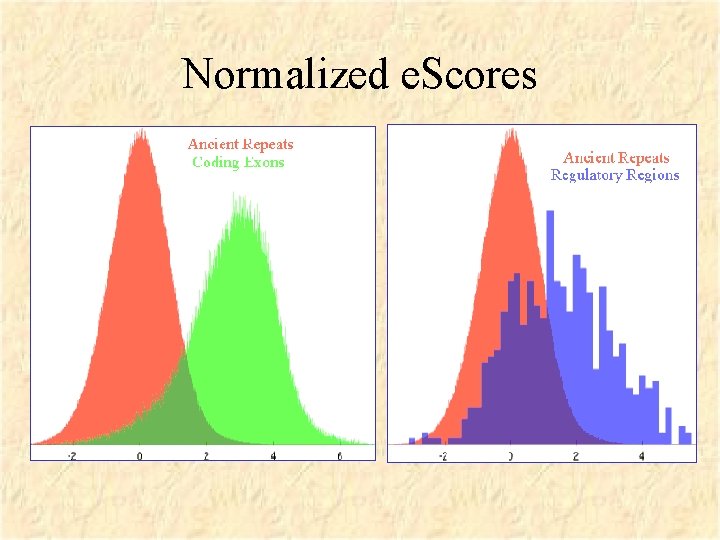
Normalized e. Scores

Computational Gene Finding 10 Different Gene Finders over a region containing two genes. One gene has a ref. Seq m. RNA and is ‘Known. ’ There is also a full length m. RNA for an ortholog of the gene at the right.

Basic Techniques • Bacteria - look for open reading frames - long stretches between start and stop codons. • Eukaryotes - introns are challenging – Look for coding exons (bounded by AG / GT) – HMMs can model coding regions and splice sites simultaniously – Generalized HMMs (genscan) can string together probable exons – Homology based ones (Gene. Wise) can map proteins to genome allowing for some evolutionary divergence and sequencing error.

Limitations of Basic Approach • Introns are vast, GT/AG splice signals are small. • Coding signal is stronger than start/stop signal. As a result gene exons are often correct but genes are fused and split. • Pseudo genes, processed and otherwise, mimic coding regions. • Pure HMM approaches tend to overpredict • Pure homology approaches only can tell us about what we already know.

Composite Approaches • Use protein homology info on top of HMMs (fgenesh++, Genome. Scan, Ensembl) • Use EST info to constrain HMMs (Genie) • Use cross species genomic alignments on top of HMMs (twinscan, fgenesh 2, SLAM, SGP)

Computational Gene Finding Ref. Seq ESTs protein X genome protein Ab initio

Computational + Wet • Even the best computational genefinding is not good enough. • Mouse/human genomic homology reduces false positive exons a fair amount typically for a given level of sensitivity. • Getting c. DNA data on a gene is much easier if you have a piece of it to start with. You can use the piece to probe c. DNA libraries rather than doing random sequencing. • On chromosome 22 Sanger probed ~60 libraries with ~every genscan prediction, spliced EST, and pufferfish homology.

Conclusions • Human genome sequencing essentially complete. • Currently have identified full coding region of ~18000 of ~25000 human protein coding genes. • The remaining genes are generally harder to identify than the ones we have identified already. • The remaining genes will require a careful mix of computational and wet work.

Acknowledgements Individuals Institutions David Haussler, Angie Hinrichs, Chuck Sugnet, Matt Schwartz, Robert Baertsch, Donna Karolchik, NHGRI, The Wellcome Trust, HHMI, NCI, Taxpayers in the US and worldwide. Francis Collins, Bob Waterston, Eric Lander, John Sulston, Richard Gibbs Roderic Guigo, Michael Brent, Chris Burges, Olivier Jaillon, David Kulp, Victor Solovyev, Ewan Birney, Greg Schuler, Deanna Church, Asif Chinwalla, the Gene Cats. Everyone else! Whitehead, Sanger, Wash U, Baylor, Stanford, DOE, and the international sequencing centers. UCSC, Mouse Sequencing Consortium, NCBI, Ensembl, Genoscope, MGC, Softberry, Affymetrix.

THE END
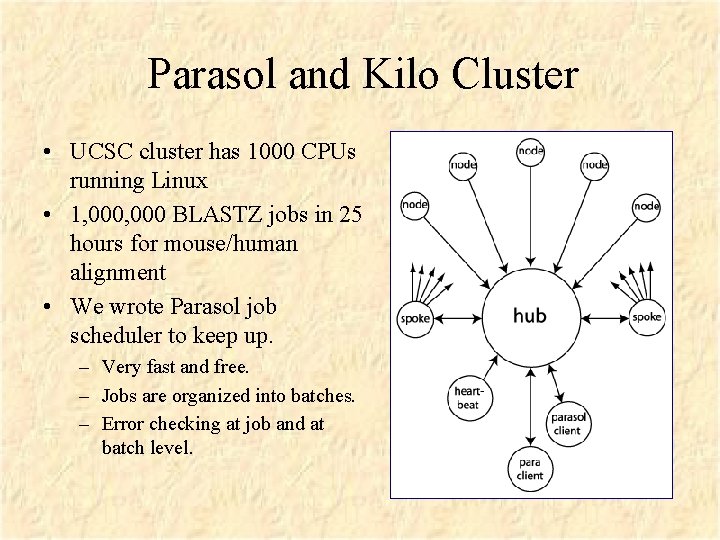
Parasol and Kilo Cluster • UCSC cluster has 1000 CPUs running Linux • 1, 000 BLASTZ jobs in 25 hours for mouse/human alignment • We wrote Parasol job scheduler to keep up. – Very fast and free. – Jobs are organized into batches. – Error checking at job and at batch level.
- Slides: 49