The General Linear Model GLM the marriage between
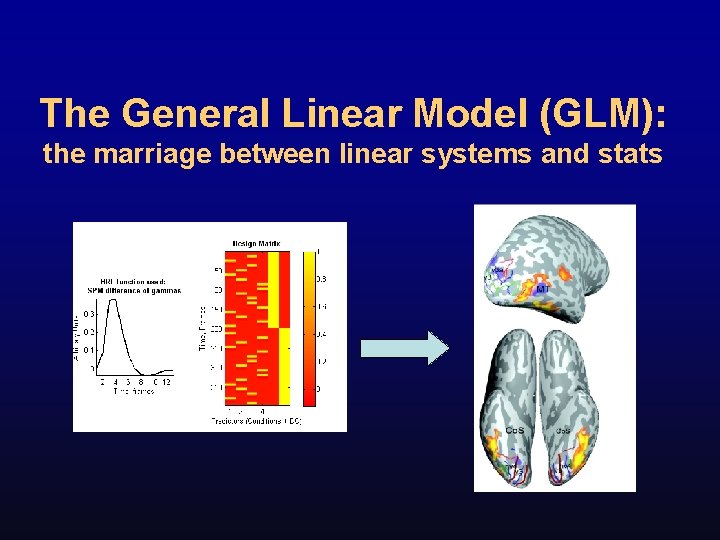
The General Linear Model (GLM): the marriage between linear systems and stats FFA

f. MRI Data Processing Stream raw scanner data preprocessing identify task related activity

The General Linear Model (GLM) • Jargon/Terms to remember – – – General Linear Model (GLM) HRF (HIRF): hemodynamic (impulse) response function Design matrix Predictors Betas ( ) Residual error • Deconvolution
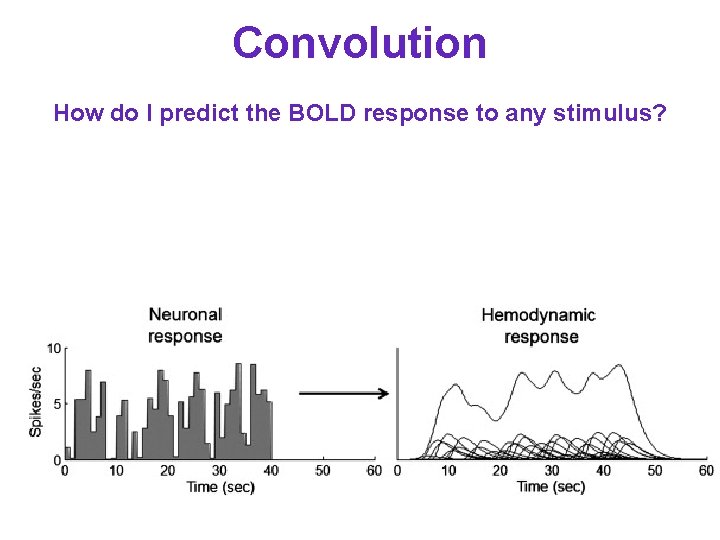
Convolution How do I predict the BOLD response to any stimulus?
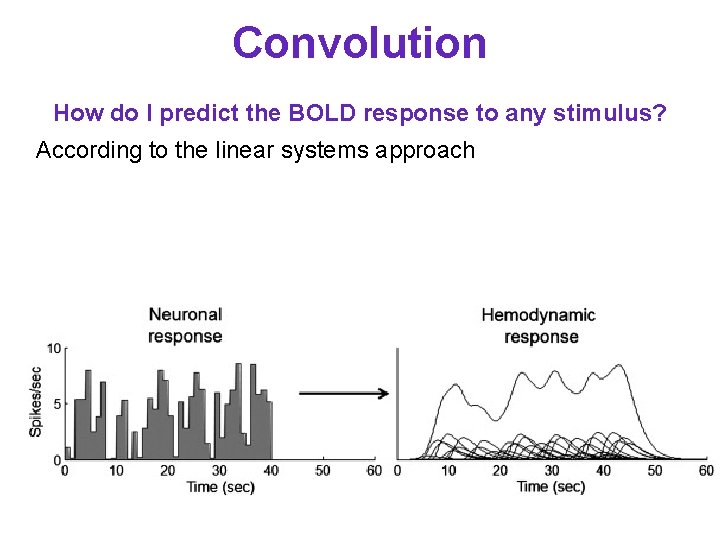
Convolution How do I predict the BOLD response to any stimulus? According to the linear systems approach - Need to know the impulse response function - Predicted response is the convolution between the input and the impulse response function
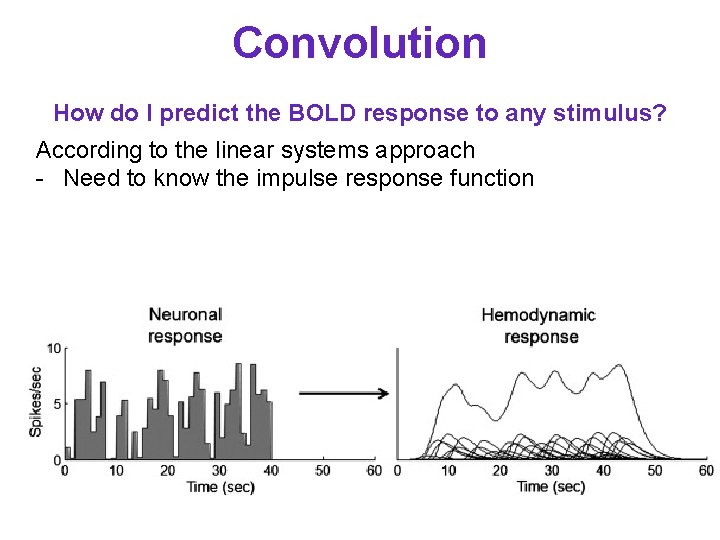
Convolution How do I predict the BOLD response to any stimulus? According to the linear systems approach - Need to know the impulse response function - Predicted response is the convolution between the input and the impulse response function
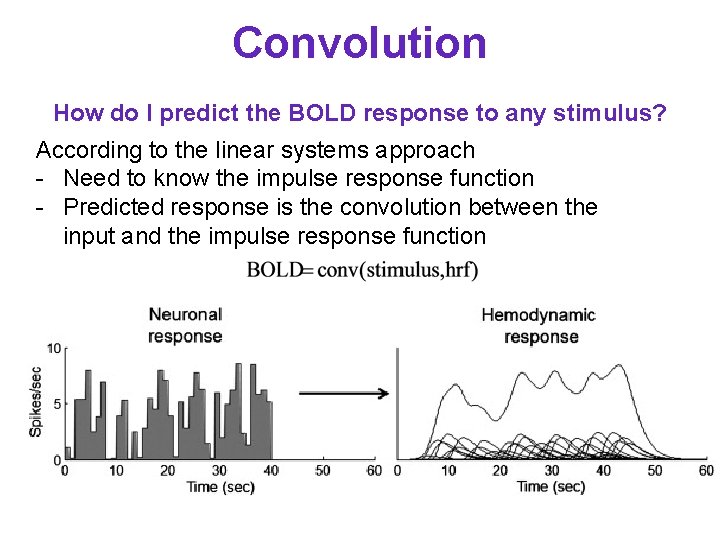
Convolution How do I predict the BOLD response to any stimulus? According to the linear systems approach - Need to know the impulse response function - Predicted response is the convolution between the input and the impulse response function
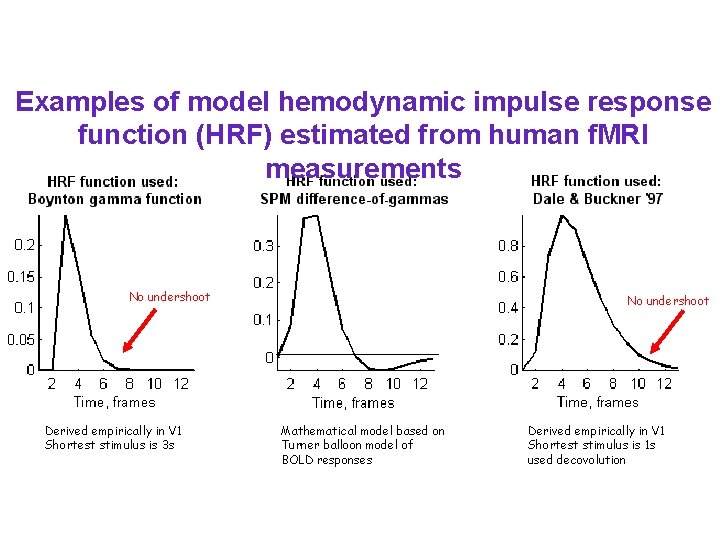
Examples of model hemodynamic impulse response function (HRF) estimated from human f. MRI measurements No undershoot Derived empirically in V 1 Shortest stimulus is 3 s No undershoot Mathematical model based on Turner balloon model of BOLD responses Derived empirically in V 1 Shortest stimulus is 1 s used decovolution
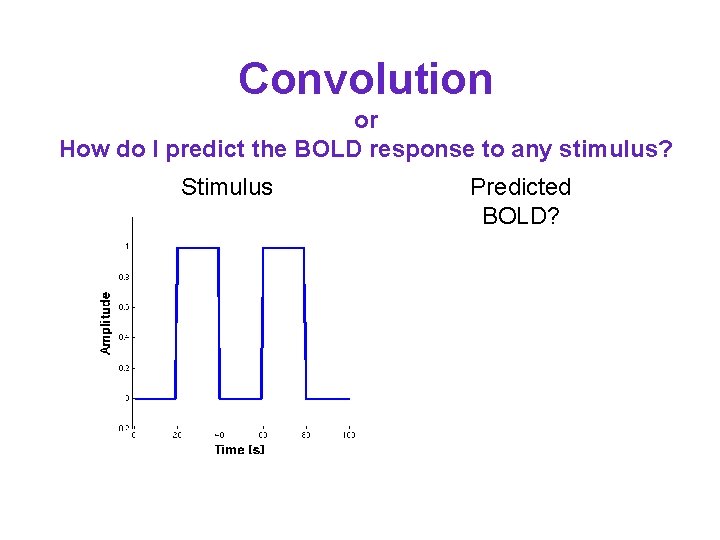
Convolution or How do I predict the BOLD response to any stimulus? Stimulus Predicted BOLD?

Convolution or How do I predict the BOLD response to any stimulus? Stimulus Predicted BOLD? • Turn the stimulus into a series of impulses • Sum up the time shifted impulse response • In other words convolve your stimulus with the hrf
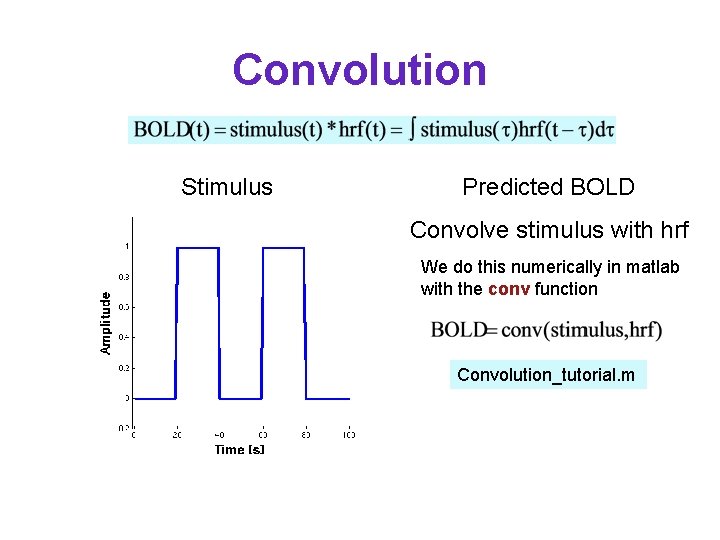
Convolution Stimulus Predicted BOLD Convolve stimulus with hrf We do this numerically in matlab with the conv function Convolution_tutorial. m
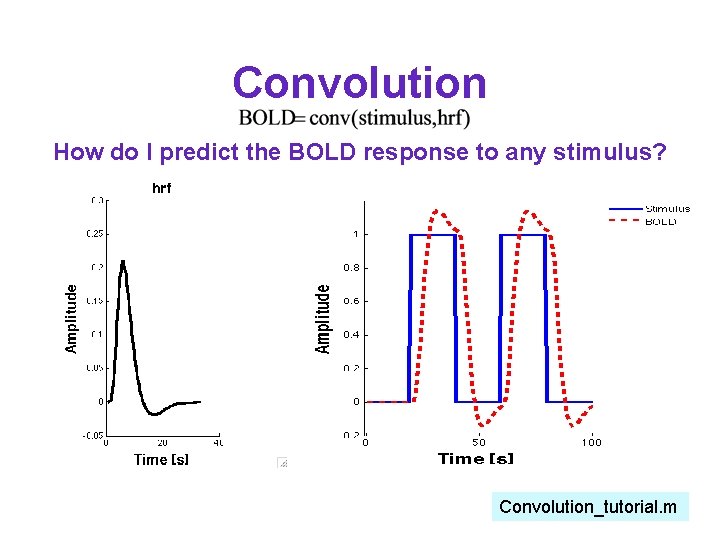
Convolution How do I predict the BOLD response to any stimulus? Convolution_tutorial. m

Hypothesis: there are neurons in the brain that respond more (increase firing) to moving than still visual stimuli
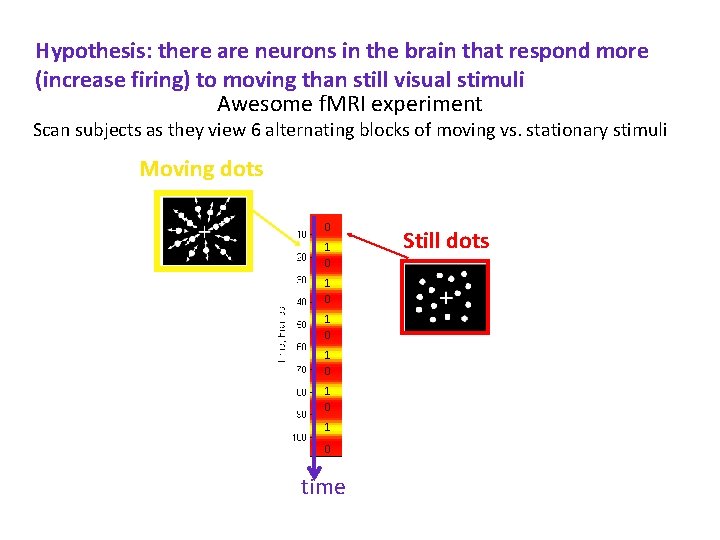
Hypothesis: there are neurons in the brain that respond more (increase firing) to moving than still visual stimuli Awesome f. MRI experiment Scan subjects as they view 6 alternating blocks of moving vs. stationary stimuli Moving dots 0 1 0 1 0 1 0 time Still dots

Hypothesis: there are neurons in the brain that respond more (increase firing) to moving than still visual stimuli Awesome f. MRI experiment Scan subjects as they view 6 alternating blocks of moving vs. stationary stimuli Moving dots 0 1 0 Prediction: BOLD response in regions containing motion sensitive neurons will be stronger during blocks of moving dots than blocks of still dots 1 0 1 0 time Still dots
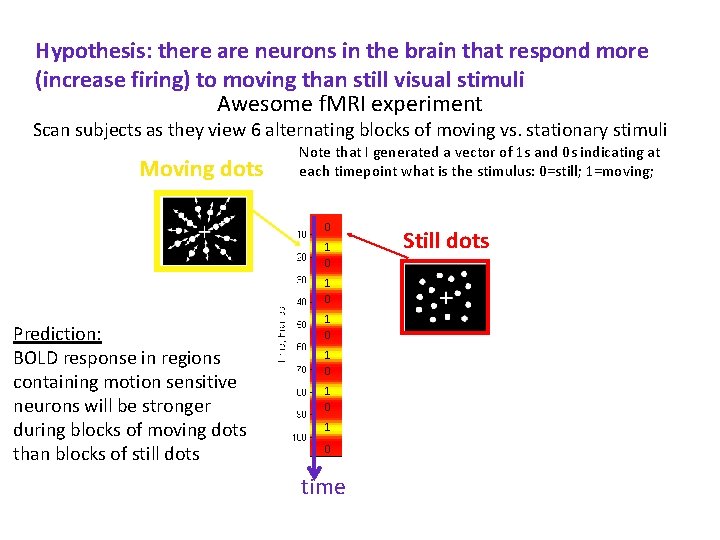
Hypothesis: there are neurons in the brain that respond more (increase firing) to moving than still visual stimuli Awesome f. MRI experiment Scan subjects as they view 6 alternating blocks of moving vs. stationary stimuli Moving dots Note that I generated a vector of 1 s and 0 s indicating at each timepoint what is the stimulus: 0=still; 1=moving; 0 1 0 Prediction: BOLD response in regions containing motion sensitive neurons will be stronger during blocks of moving dots than blocks of still dots 1 0 1 0 time Still dots
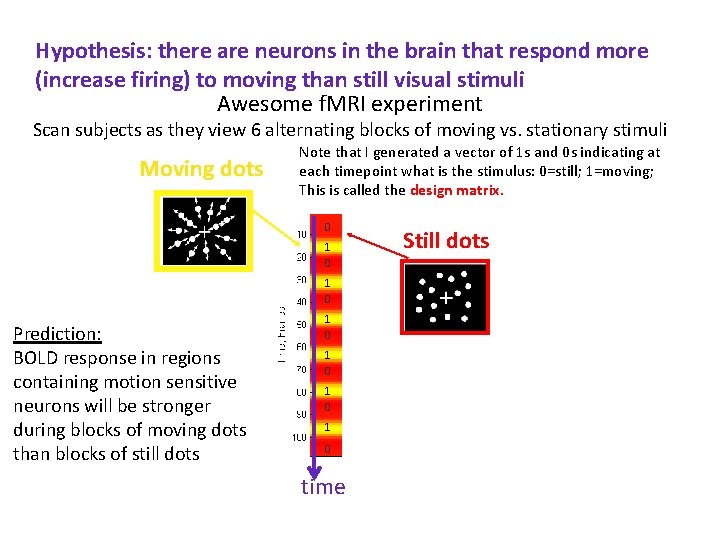
Hypothesis: there are neurons in the brain that respond more (increase firing) to moving than still visual stimuli Awesome f. MRI experiment Scan subjects as they view 6 alternating blocks of moving vs. stationary stimuli Moving dots Note that I generated a vector of 1 s and 0 s indicating at each timepoint what is the stimulus: 0=still; 1=moving; This is called the design matrix. 0 1 0 Prediction: BOLD response in regions containing motion sensitive neurons will be stronger during blocks of moving dots than blocks of still dots 1 0 1 0 time Still dots
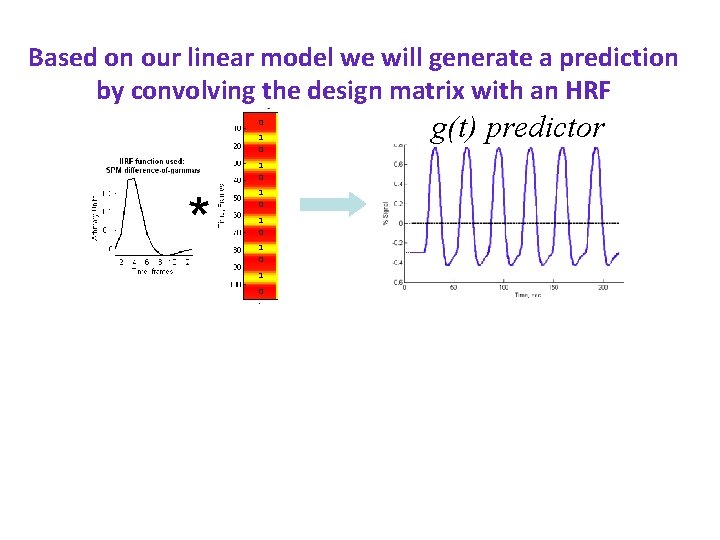
Based on our linear model we will generate a prediction by convolving the design matrix with an HRF 0 HRF 1 0 * 1 0 1 0 g(t) predictor
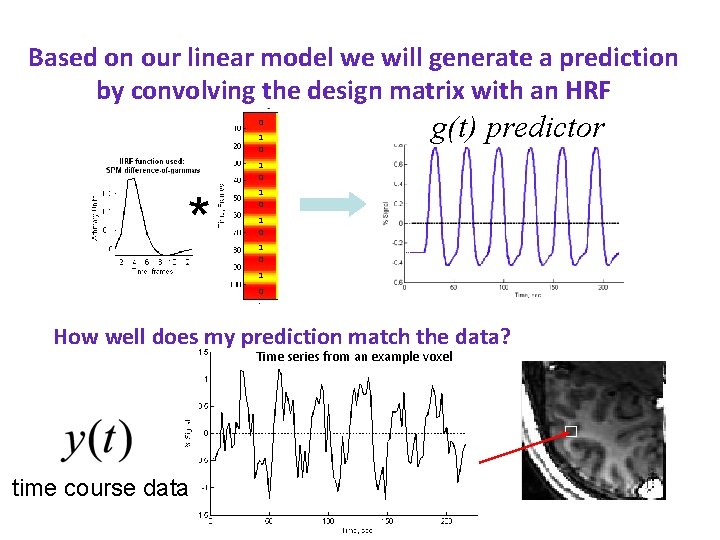
Based on our linear model we will generate a prediction by convolving the design matrix with an HRF 0 HRF 1 0 g(t) predictor 1 0 * 1 0 1 0 How well does my prediction match the data? Time series from an example voxel time course data
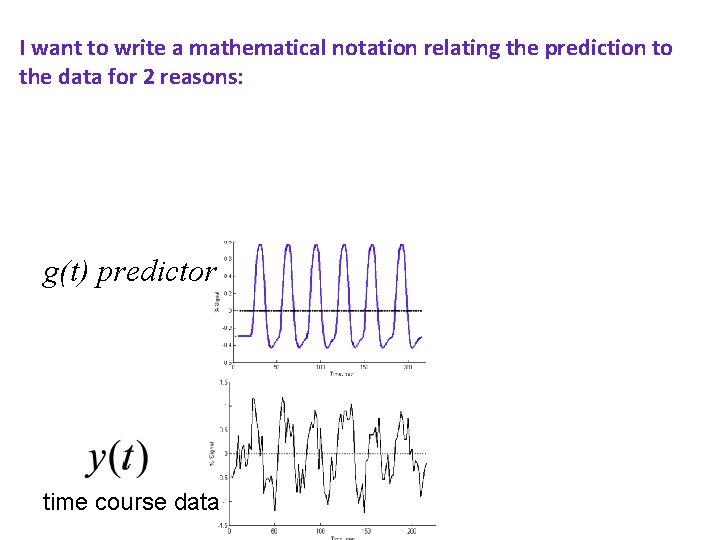
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is g(t) predictor time course data
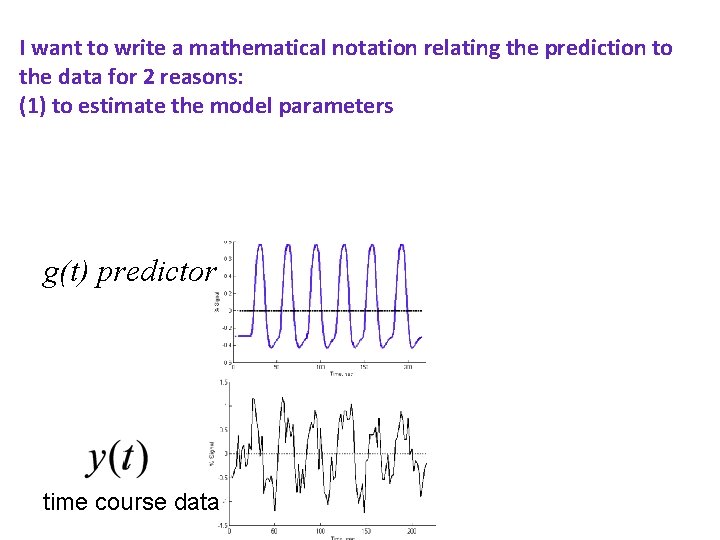
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is g(t) predictor time course data
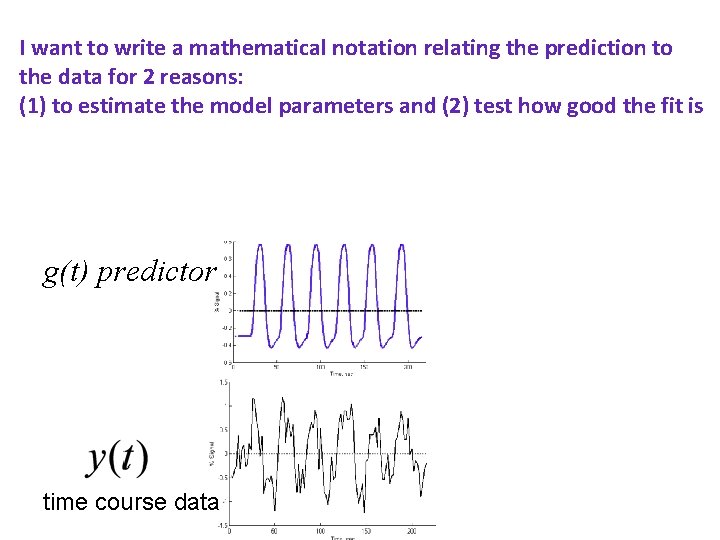
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is g(t) predictor time course data
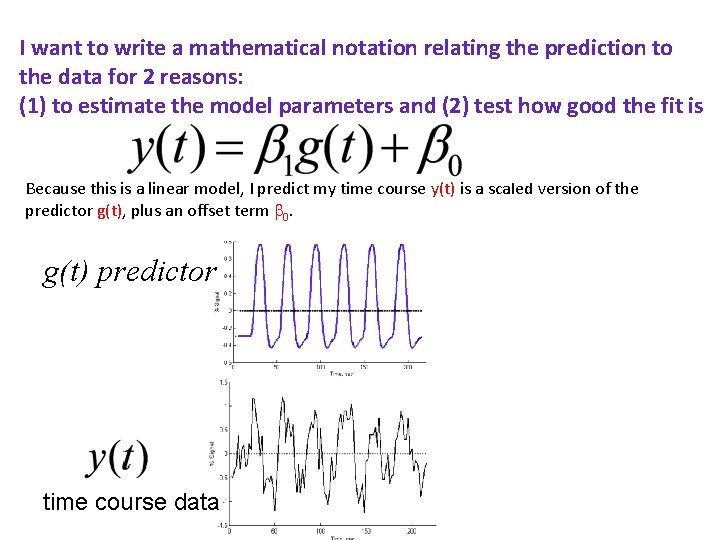
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is Because this is a linear model, I predict my time course y(t) is a scaled version of the predictor g(t), plus an offset term 0. g(t) predictor time course data
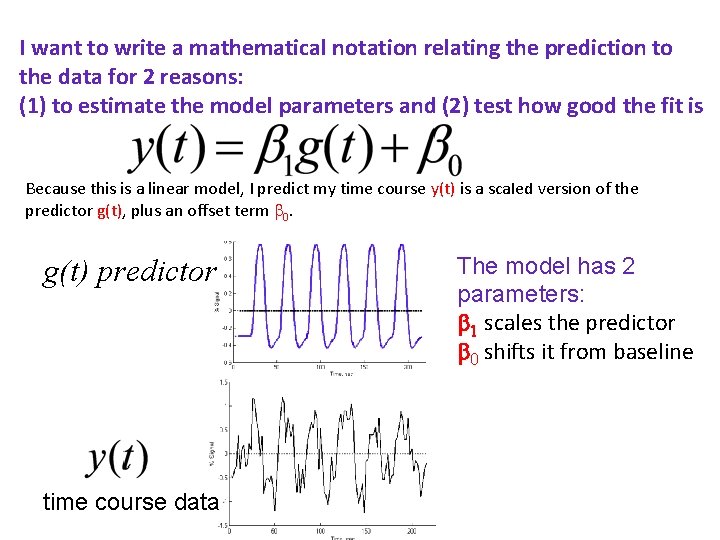
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is Because this is a linear model, I predict my time course y(t) is a scaled version of the predictor g(t), plus an offset term 0. g(t) predictor time course data The model has 2 parameters: 1 scales the predictor 0 shifts it from baseline and an error term e(t), residual error: what the linear model doesn’t explain in the data

I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is Because this is a linear model, I predict my time course y(t) is a scaled version of the predictor g(t), plus an offset term 0. g(t) predictor time course data The model has 2 parameters: 1 scales the predictor 0 shifts it from baseline and an error term e(t), residual error: what the linear model doesn’t explain in the data
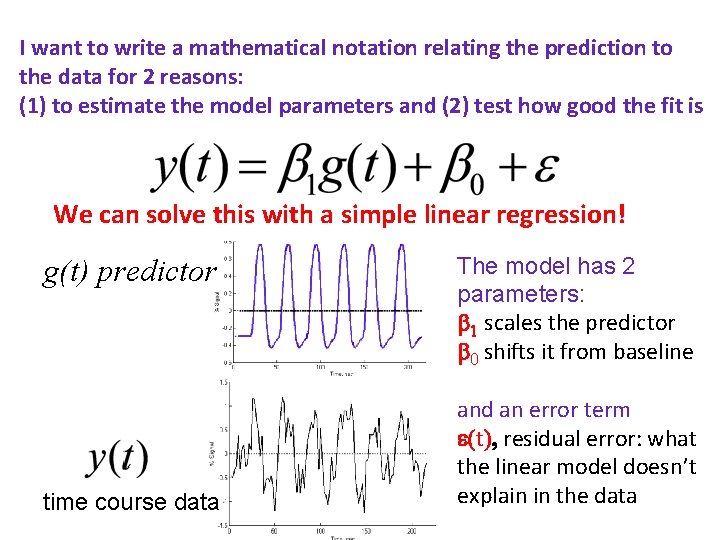
I want to write a mathematical notation relating the prediction to the data for 2 reasons: (1) to estimate the model parameters and (2) test how good the fit is We can solve this with a simple linear regression! g(t) predictor time course data The model has 2 parameters: 1 scales the predictor 0 shifts it from baseline and an error term e(t), residual error: what the linear model doesn’t explain in the data
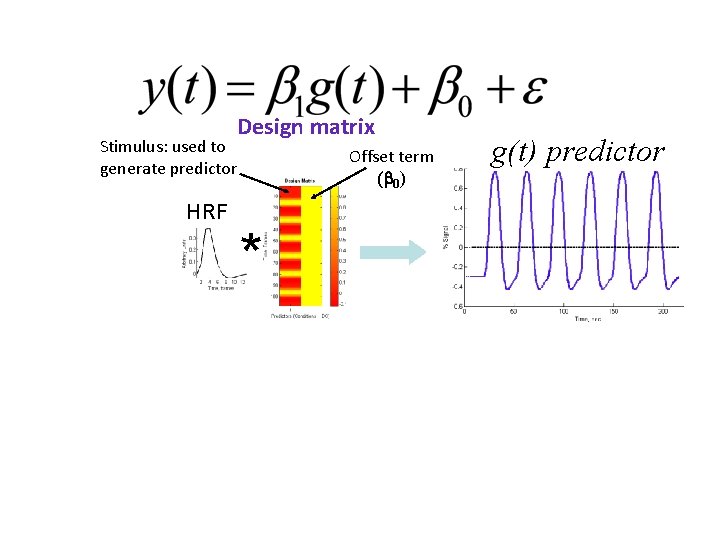
Stimulus: used to generate predictor HRF Design matrix Offset term ( 0) * g(t) predictor
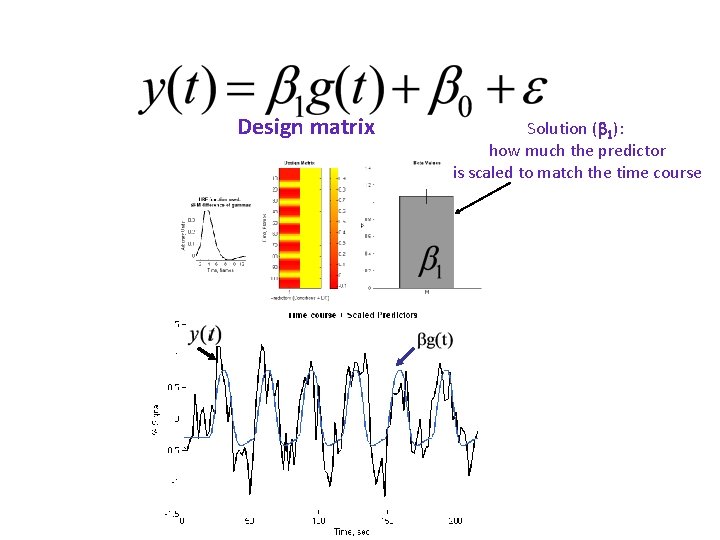
Design matrix Solution ( 1): how much the predictor is scaled to match the time course
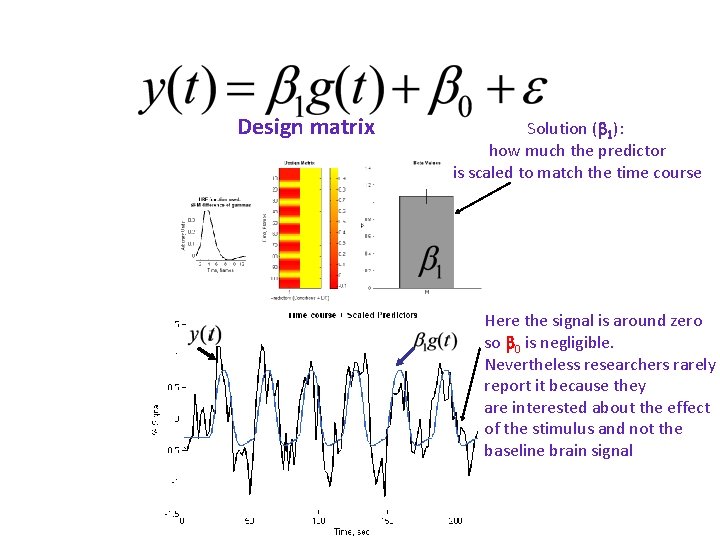
Design matrix Solution ( 1): how much the predictor is scaled to match the time course Here the signal is around zero so 0 is negligible. Nevertheless researchers rarely report it because they are interested about the effect of the stimulus and not the baseline brain signal
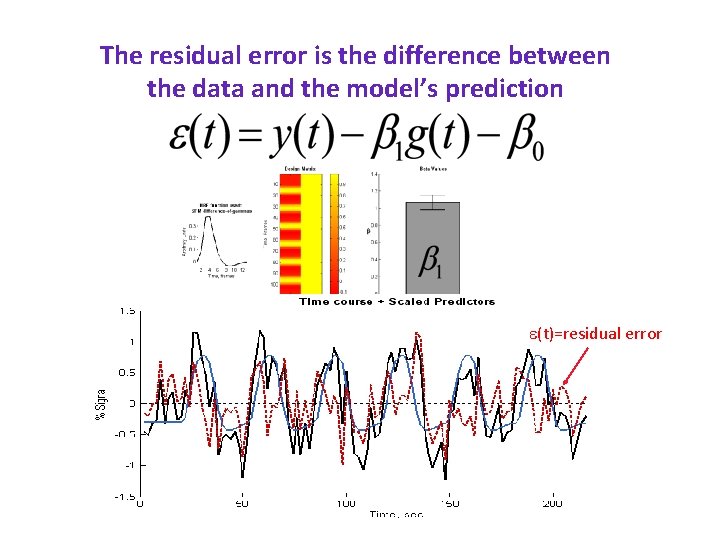
The residual error is the difference between the data and the model’s prediction e(t)=residual error
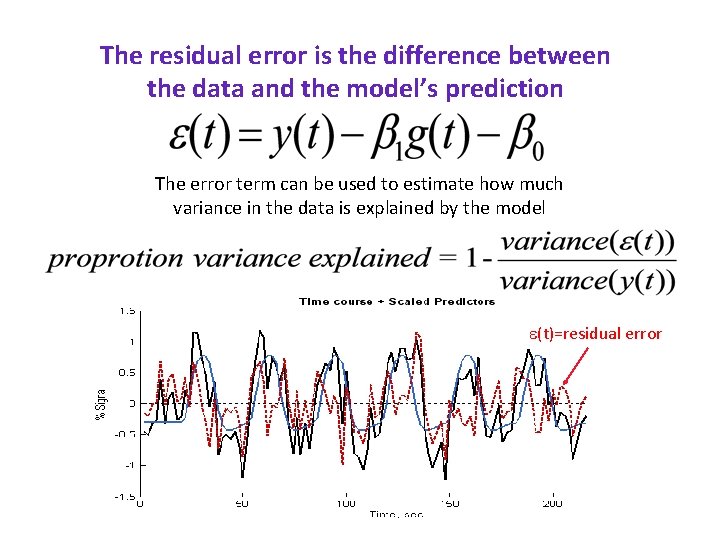
The residual error is the difference between the data and the model’s prediction The error term can be used to estimate how much variance in the data is explained by the model e(t)=residual error
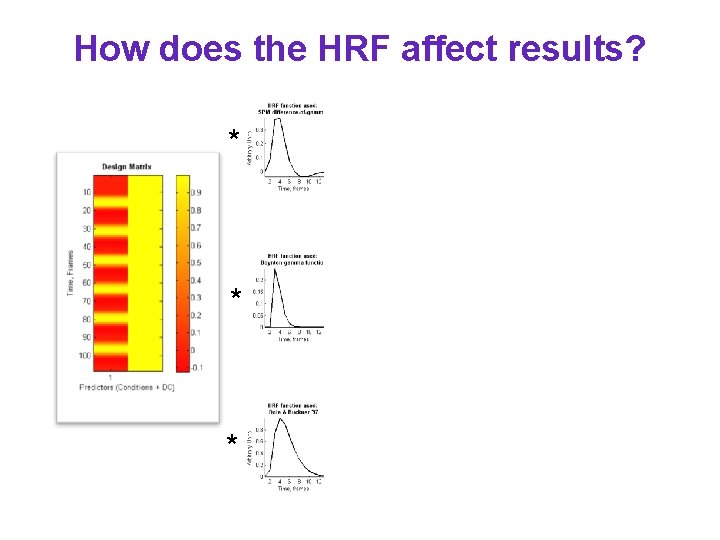
How does the HRF affect results? * * *
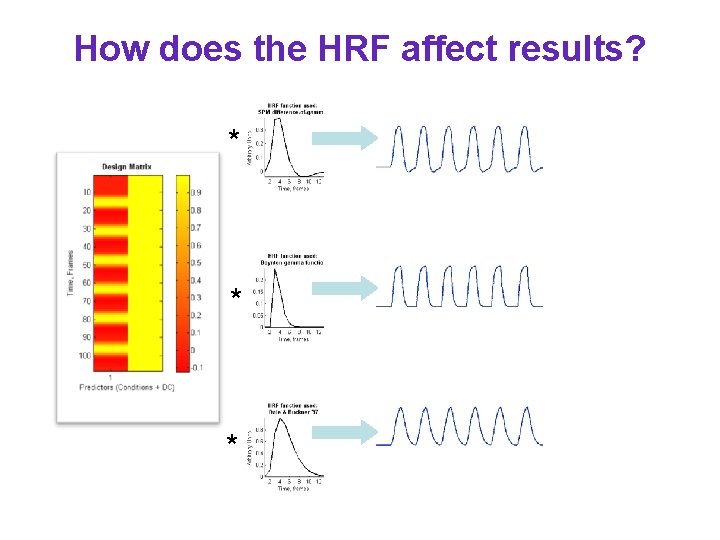
How does the HRF affect results? * * *
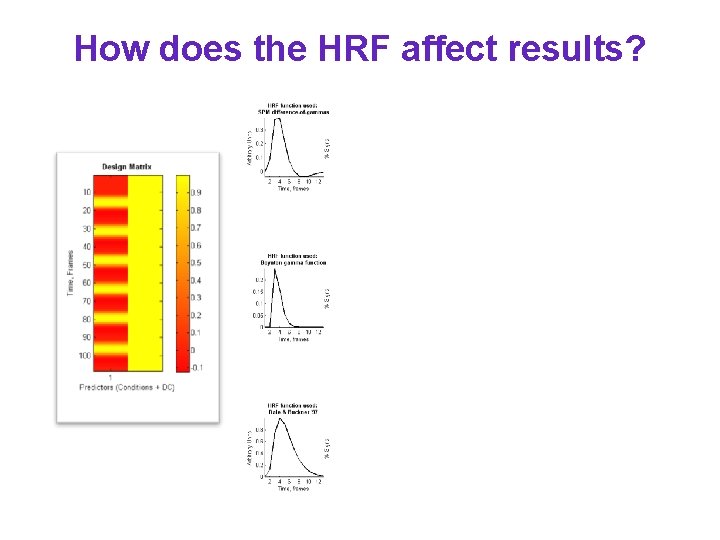
How does the HRF affect results?

The General Linear Model: expand the regression model to have more than one predictor • • • y(t) gi(t) i 0 e(t) time course of voxel i-th factor coefficient of i-th factor shift from baseline additive noise

General Linear Model: Matrix Notation • Y time course column vector (nx 1) n: number of time samples • G matrix of concatenated predictors (nxp) p: number of predictors (number of experimental conditions) • • e vector of factor coefficients (px 1) additive noise Least Squares Solution:

General Linear Model: Matrix Notation • Y time course column vector (nx 1) n: number of time samples • G matrix of concatenated predictors (nxp) p: number of predictors (number of experimental conditions) • • e vector of factor coefficients (px 1) additive noise Least Squares Solution: As an experimenter you generate the design matrix, which then gets convolved with the HRF to generator predictors g i

Example GLM with 4 predictors and 2 baselines for run 1 and run 2 F P O T
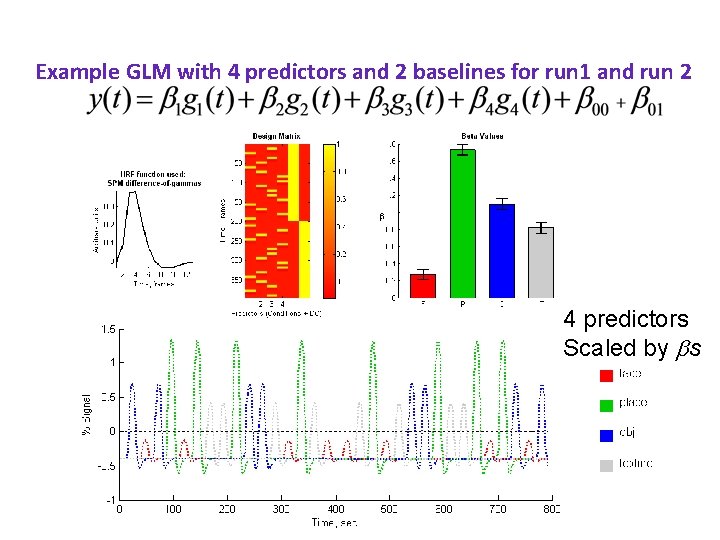
Example GLM with 4 predictors and 2 baselines for run 1 and run 2 4 predictors Scaled by s
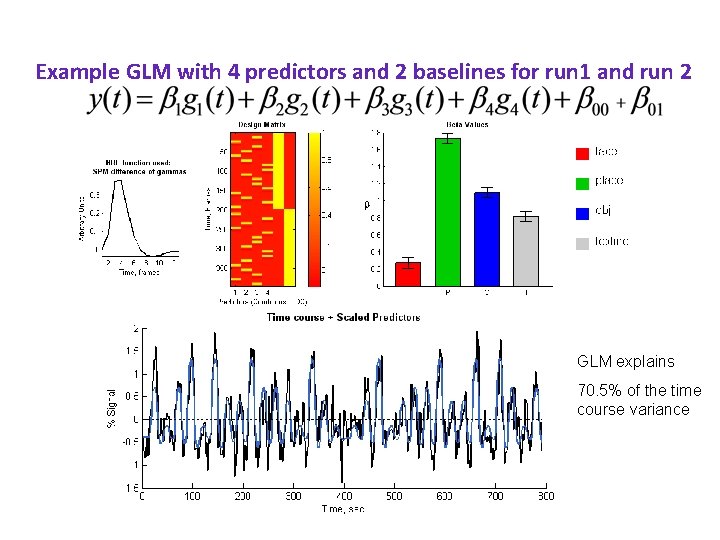
Example GLM with 4 predictors and 2 baselines for run 1 and run 2 GLM explains 70. 5% of the time course variance
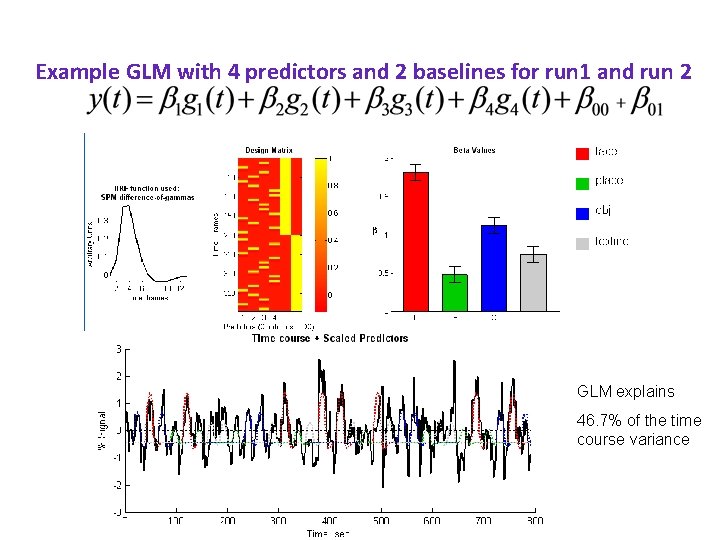
Example GLM with 4 predictors and 2 baselines for run 1 and run 2 GLM explains 46. 7% of the time course variance

Correlated Predictors • Avoid predictors that are correlated with one another • This is why we NEVER include a baseline predictor – baseline predictor is almost completely correlated (r = -1) with the sum of other existing predictors – if we included a baseline predictor, the model would have problems assigning variance to stimulus predictors vs. baseline predictors • for example, the model could not distinguish between two possibilities (e. g. , Beta 1=1, Beta 2=0 vs. Beta 1=0, Beta 2=-1) Stimulus predictor Baseline predictor

Correlated Predictors • Avoid predictors that are correlated with one another • This is why we NEVER include a baseline predictor – baseline predictor is almost completely correlated (r = -1) with the sum of other existing predictors – if we included a baseline predictor, the model would have problems assigning variance to stimulus predictors vs. baseline predictors • for example, the model could not distinguish between two possibilities (e. g. , Beta 1=1, Beta 2=0 vs. Beta 1=0, Beta 2=-1) Stimulus predictor Baseline predictor

Correlated Predictors • Avoid predictors that are correlated with one another • This is why we NEVER include a baseline predictor – baseline predictor is almost completely correlated (r = -1) with the sum of other existing predictors – if we included a baseline predictor, the model would have problems assigning variance to stimulus predictors vs. baseline predictors • for example, the model could not distinguish between two possibilities (e. g. , Beta 1=1, Beta 2=0 vs. Beta 1=0, Beta 2=-1) Stimulus predictor Baseline predictor

Deconvolution Can you solve the inverse problem? If I measure the summed response of several impulses, can I recover the hemodynamic response to a single event?
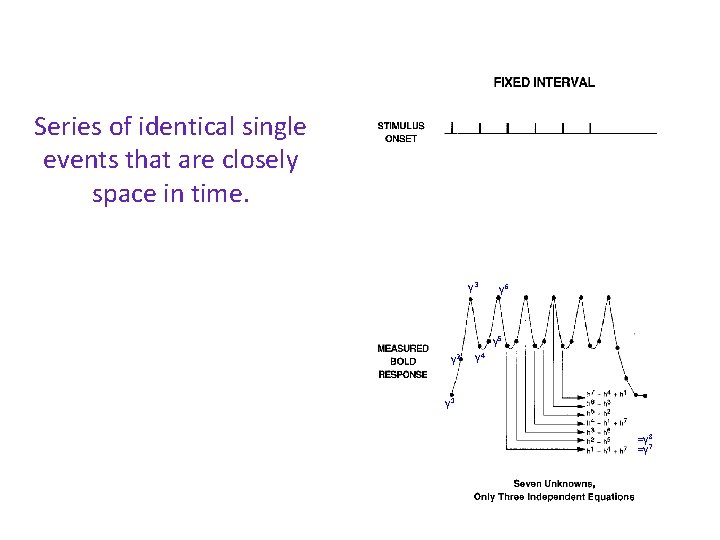
Series of identical single events that are closely space in time. h 3 h 4 h 2 Can I recover the hemodynamic response of a single event? h 5 h 6 h 1 h 7 y 3 y 6 y 5 y 2 y 4 y 1 =y 8 =y 7
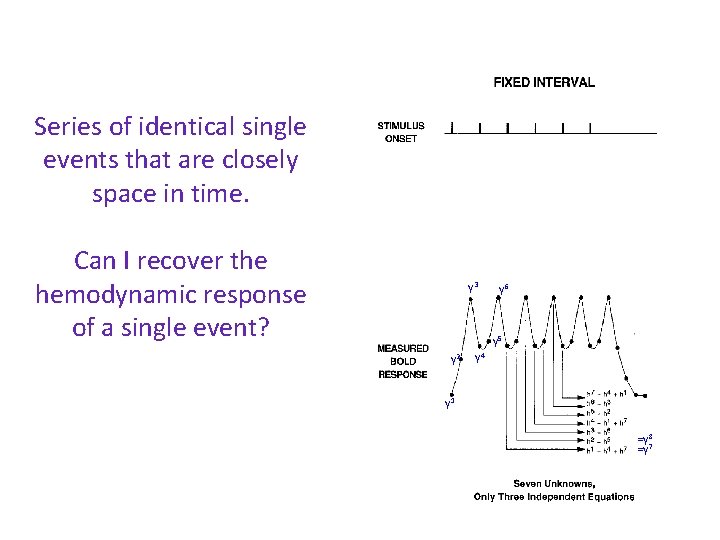
Series of identical single events that are closely space in time. h 3 h 4 h 2 Can I recover the hemodynamic response of a single event? h 5 h 6 h 1 h 7 y 3 y 6 y 5 y 2 y 4 y 1 =y 8 =y 7
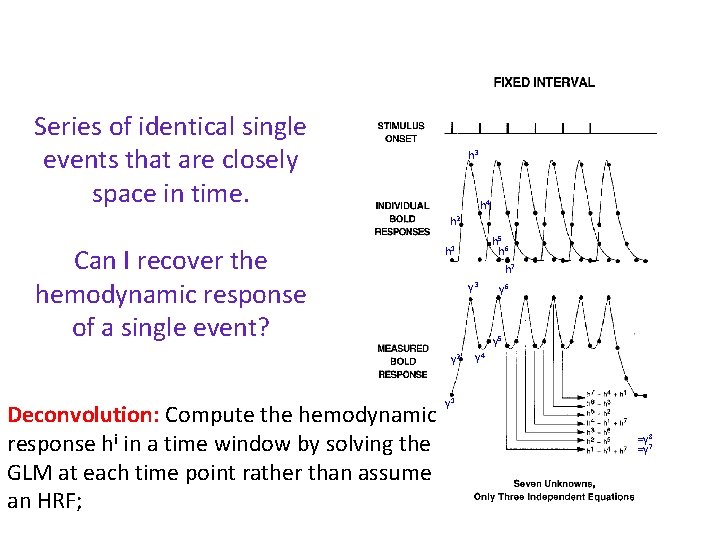
Series of identical single events that are closely space in time. h 3 h 4 h 2 Can I recover the hemodynamic response of a single event? h 7 y 3 y 6 y 5 y 2 Deconvolution: Compute the hemodynamic response hi in a time window by solving the GLM at each time point rather than assume an HRF; h 5 h 6 h 1 y 4 y 1 =y 8 =y 7
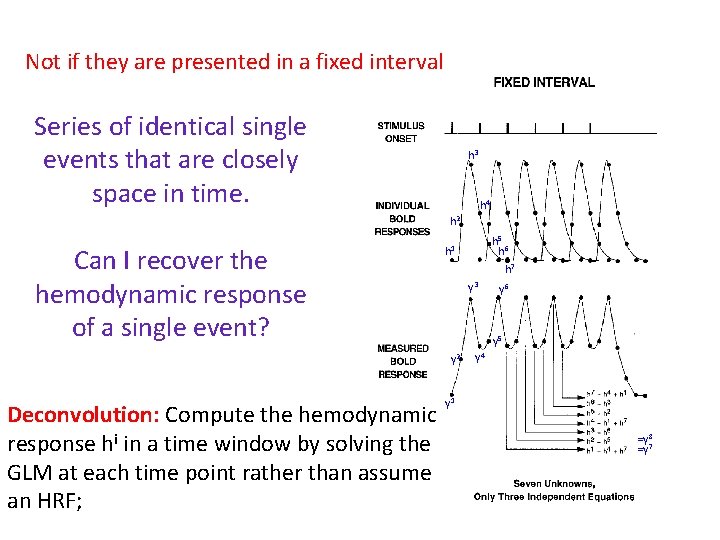
Not if they are presented in a fixed interval Series of identical single events that are closely space in time. h 3 h 4 h 2 Can I recover the hemodynamic response of a single event? h 7 y 3 y 6 y 5 y 2 Deconvolution: Compute the hemodynamic response hi in a time window by solving the GLM at each time point rather than assume an HRF; h 5 h 6 h 1 y 4 y 1 =y 8 =y 7
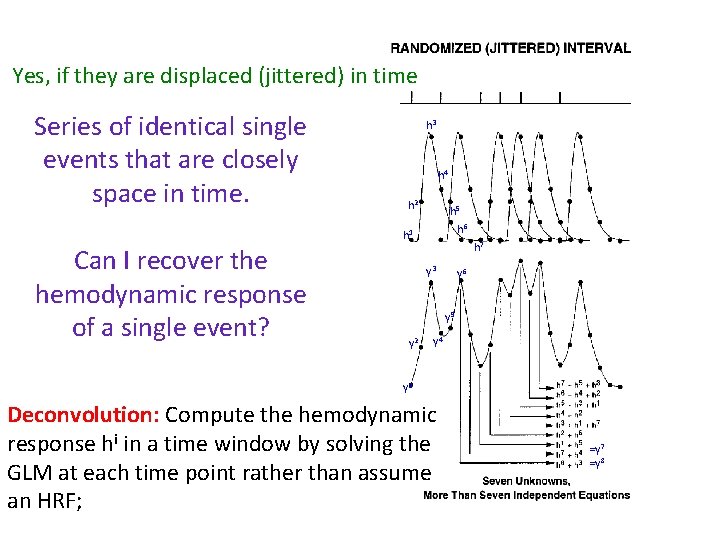
Yes, if they are displaced (jittered) in time Series of identical single events that are closely space in time. h 3 h 4 h 2 h 5 h 6 h 1 Can I recover the hemodynamic response of a single event? h 7 y 3 y 6 y 5 y 2 y 4 y 1 Deconvolution: Compute the hemodynamic response hi in a time window by solving the GLM at each time point rather than assume an HRF; =y 7 =y 8
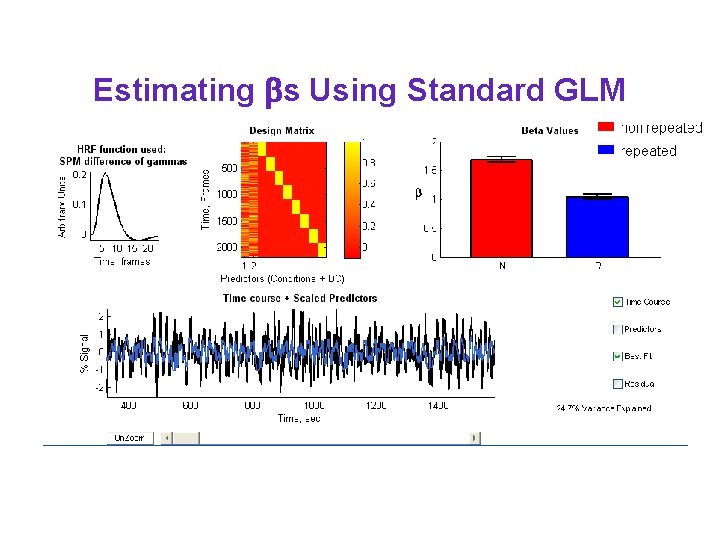
Estimating s Using Standard GLM
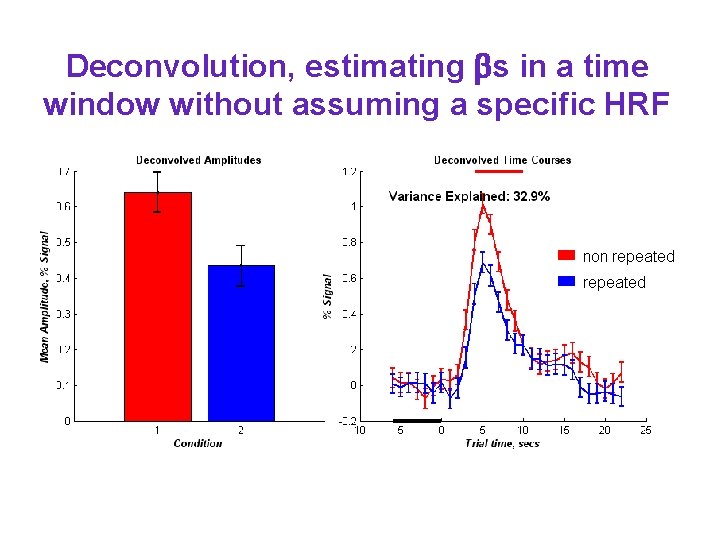
Deconvolution, estimating s in a time window without assuming a specific HRF
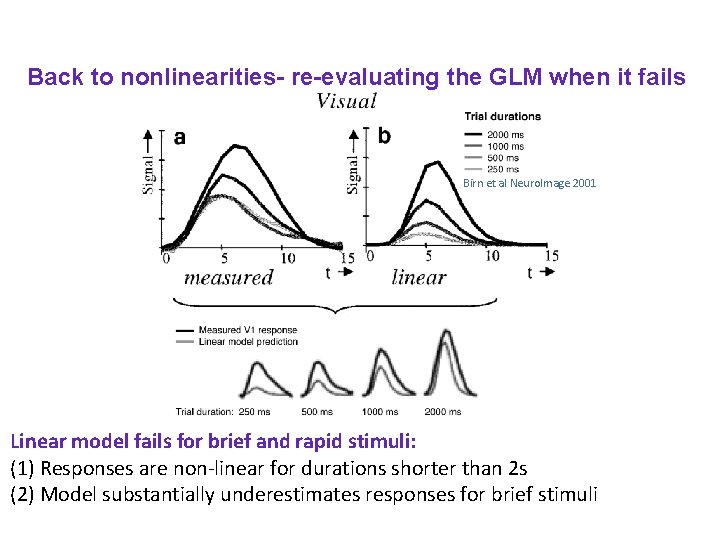
Back to nonlinearities- re-evaluating the GLM when it fails Birn et al Neuro. Image 2001 Linear model fails for brief and rapid stimuli: (1) Responses are non-linear for durations shorter than 2 s (2) Model substantially underestimates responses for brief stimuli
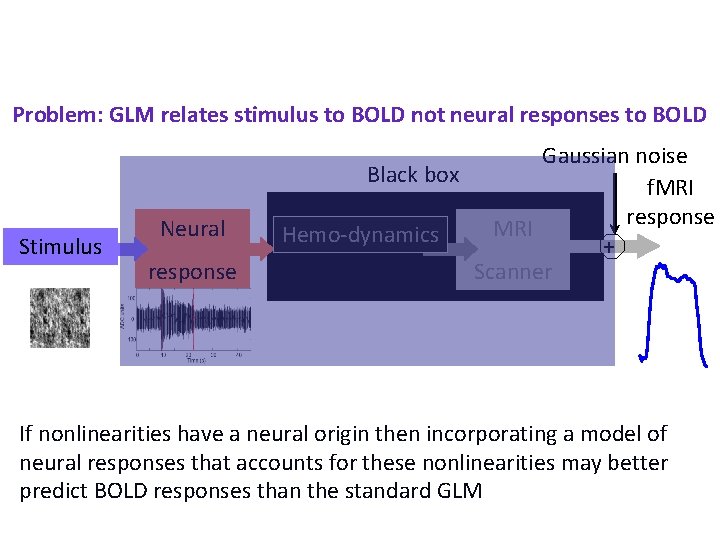
Problem: GLM relates stimulus to BOLD not neural responses to BOLD Stimulus Neural response Gaussian noise Black box f. MRI response MRI Hemo-dynamics + Scanner If nonlinearities have a neural origin then incorporating a model of neural responses that accounts for these nonlinearities may better predict BOLD responses than the standard GLM
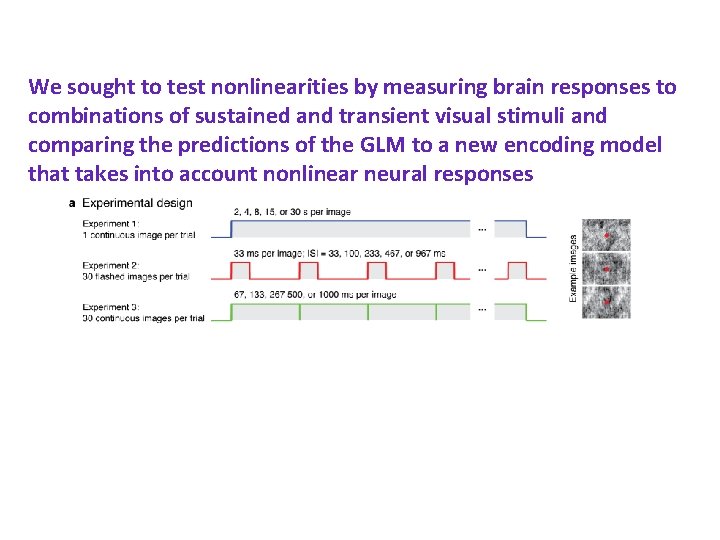
We sought to test nonlinearities by measuring brain responses to combinations of sustained and transient visual stimuli and comparing the predictions of the GLM to a new encoding model that takes into account nonlinear neural responses
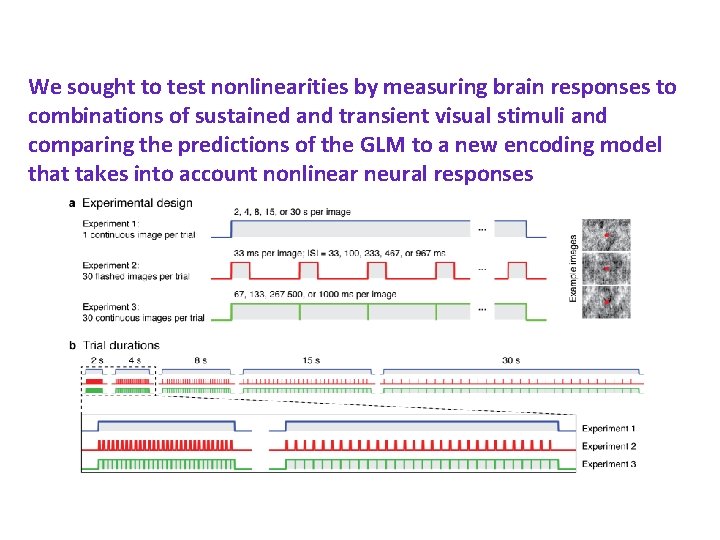
We sought to test nonlinearities by measuring brain responses to combinations of sustained and transient visual stimuli and comparing the predictions of the GLM to a new encoding model that takes into account nonlinear neural responses
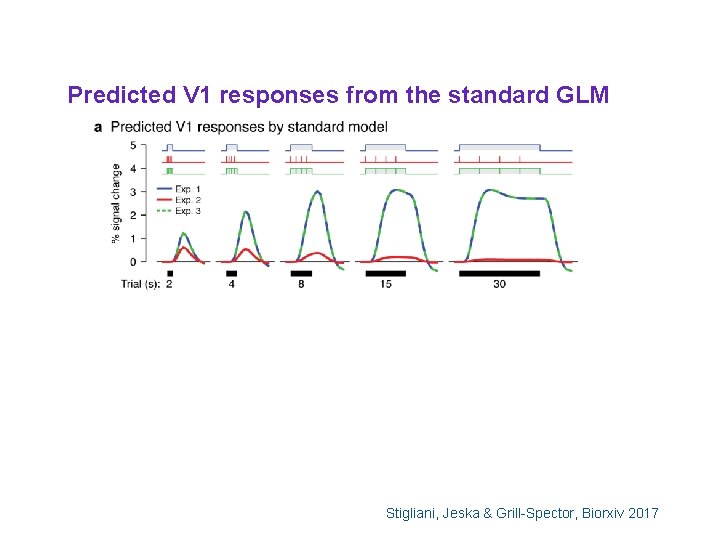
Predicted V 1 responses from the standard GLM Stigliani, Jeska & Grill-Spector, Biorxiv 2017
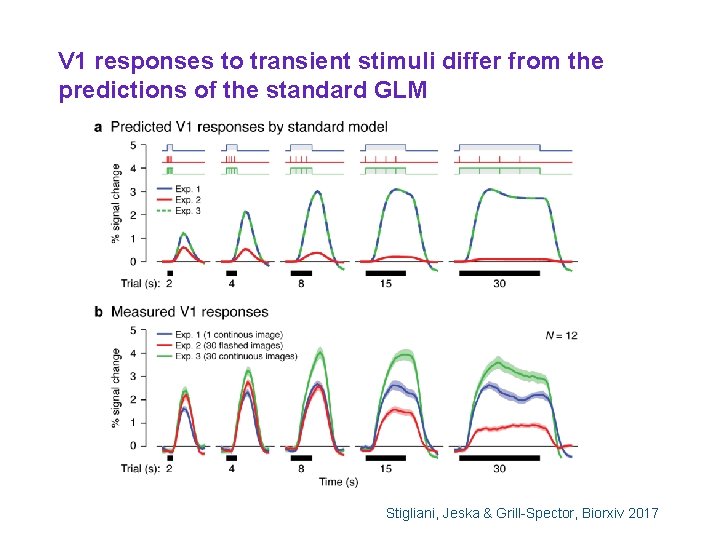
V 1 responses to transient stimuli differ from the predictions of the standard GLM Stigliani, Jeska & Grill-Spector, Biorxiv 2017
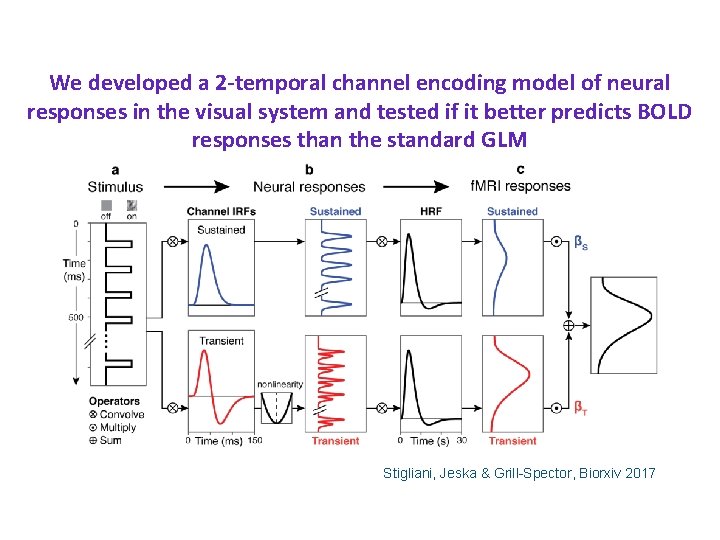
We developed a 2 -temporal channel encoding model of neural responses in the visual system and tested if it better predicts BOLD responses than the standard GLM Stigliani, Jeska & Grill-Spector, Biorxiv 2017
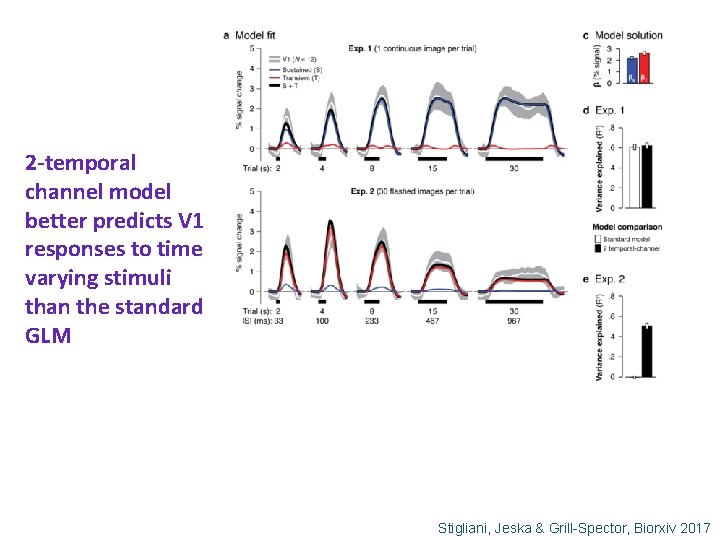
2 -temporal channel model better predicts V 1 responses to time varying stimuli than the standard GLM Stigliani, Jeska & Grill-Spector, Biorxiv 2017
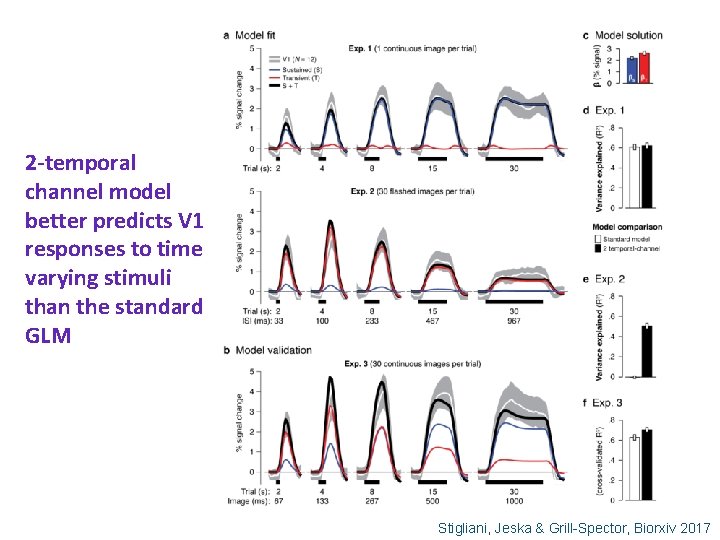
2 -temporal channel model better predicts V 1 responses to time varying stimuli than the standard GLM Stigliani, Jeska & Grill-Spector, Biorxiv 2017
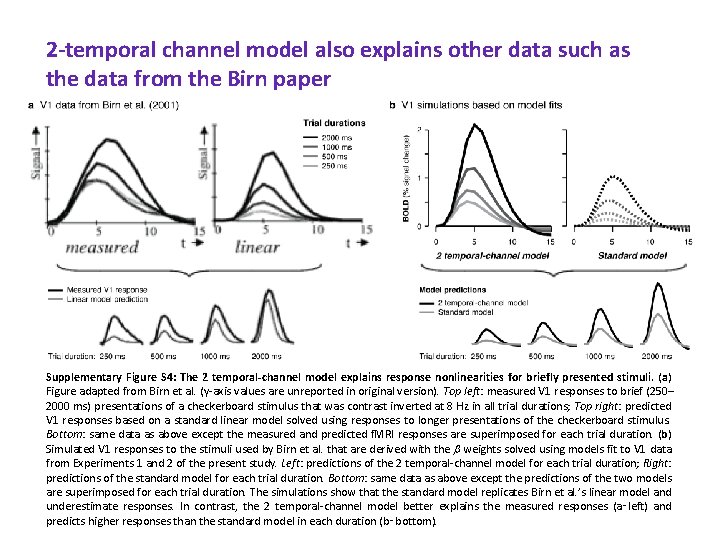
2 -temporal channel model also explains other data such as the data from the Birn paper Supplementary Figure S 4: The 2 temporal-channel model explains response nonlinearities for briefly presented stimuli. (a) Figure adapted from Birn et al. (y-axis values are unreported in original version). Top left: measured V 1 responses to brief (250– 2000 ms) presentations of a checkerboard stimulus that was contrast inverted at 8 Hz in all trial durations; Top right: predicted V 1 responses based on a standard linear model solved using responses to longer presentations of the checkerboard stimulus. Bottom: same data as above except the measured and predicted f. MRI responses are superimposed for each trial duration. (b) Simulated V 1 responses to the stimuli used by Birn et al. that are derived with the weights solved using models fit to V 1 data from Experiments 1 and 2 of the present study. Left: predictions of the 2 temporal-channel model for each trial duration; Right: predictions of the standard model for each trial duration. Bottom: same data as above except the predictions of the two models are superimposed for each trial duration. The simulations show that the standard model replicates Birn et al. ’s linear model and underestimate responses. In contrast, the 2 temporal-channel model better explains the measured responses (a‑left) and predicts higher responses than the standard model in each duration (b‑bottom).
- Slides: 62