The experimental system variation Drosophila Egg chamber Eggshell
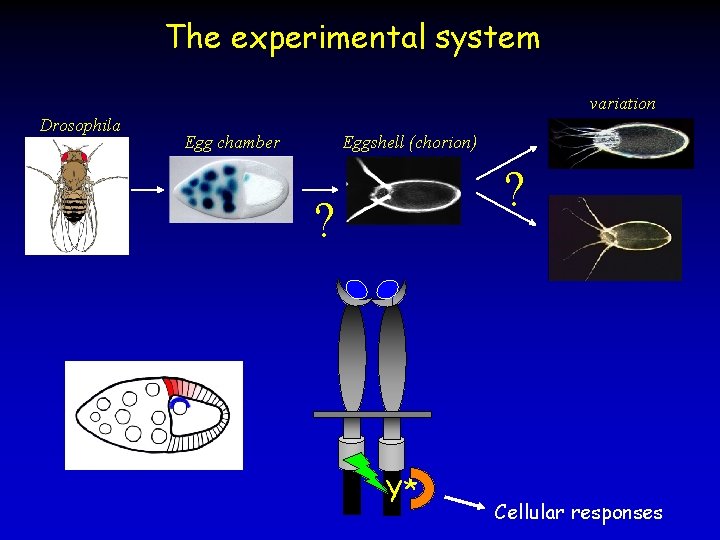
The experimental system variation Drosophila Egg chamber Eggshell (chorion) ? ? Y* Cellular responses
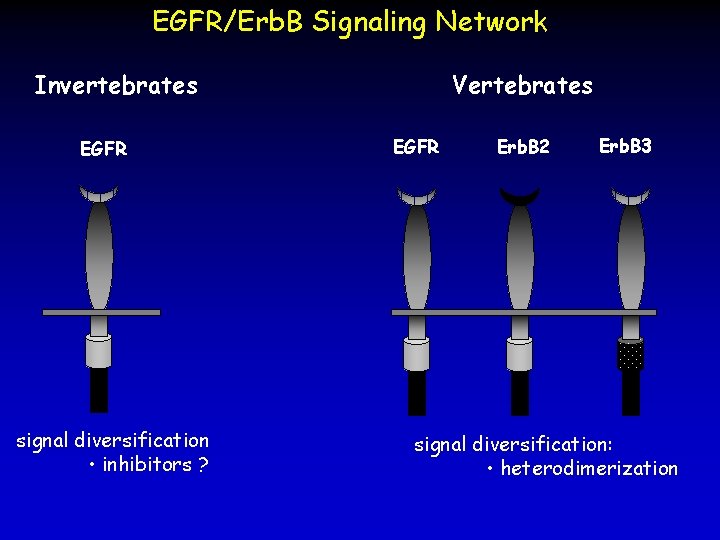
EGFR/Erb. B Signaling Network Invertebrates EGFR signal diversification • inhibitors ? Vertebrates EGFR Erb. B 2 Erb. B 3 signal diversification: • heterodimerization
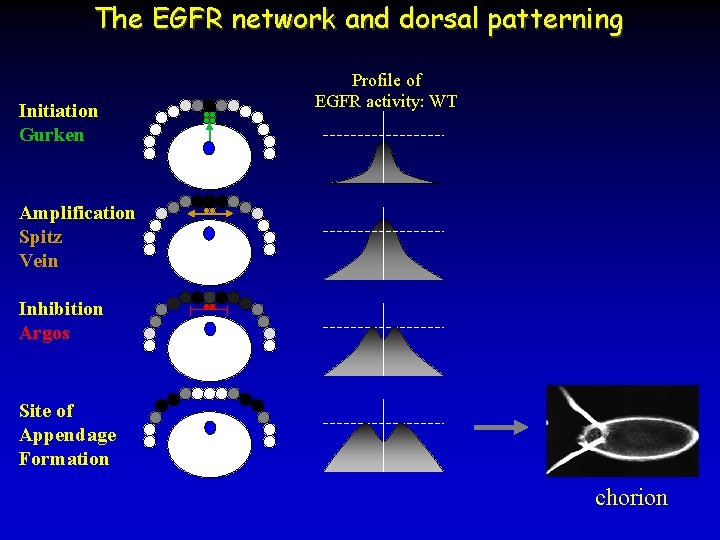
The EGFR network and dorsal patterning Initiation Gurken Profile of EGFR activity: WT Amplification Spitz Vein Inhibition Argos Site of Appendage Formation chorion
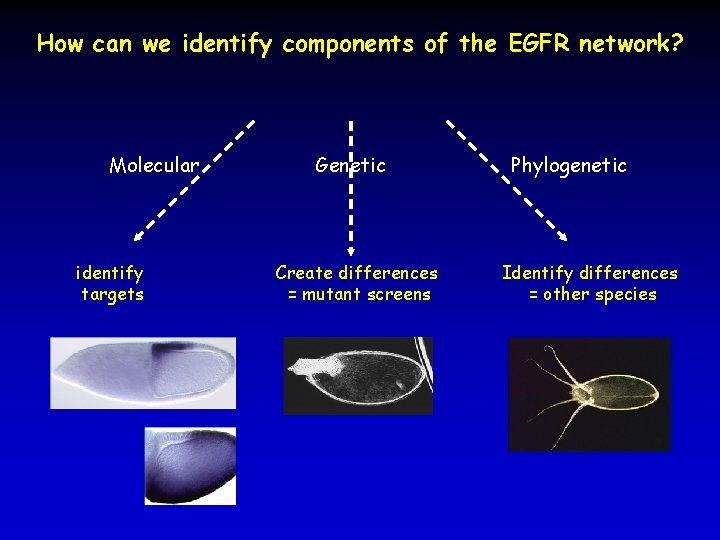
How can we identify components of the EGFR network? Molecular identify targets Genetic Create differences = mutant screens Phylogenetic Identify differences = other species
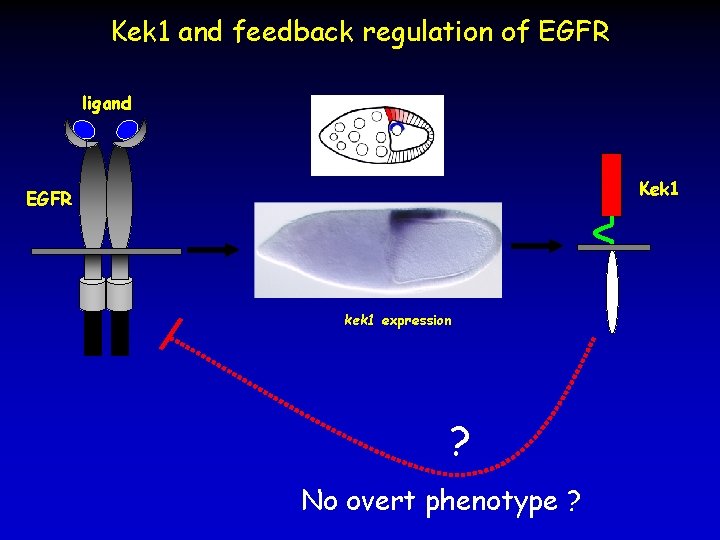
Kek 1 and feedback regulation of EGFR ligand Kek 1 EGFR kek 1 expression ? No overt phenotype ?
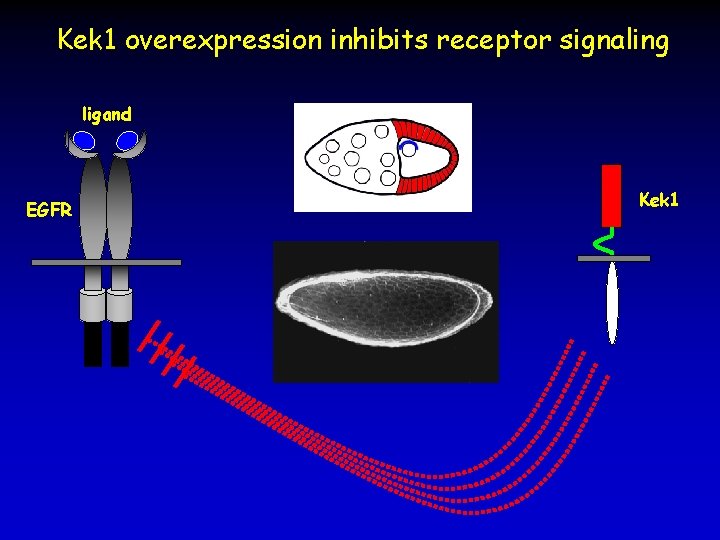
Kek 1 overexpression inhibits receptor signaling ligand EGFR Kek 1
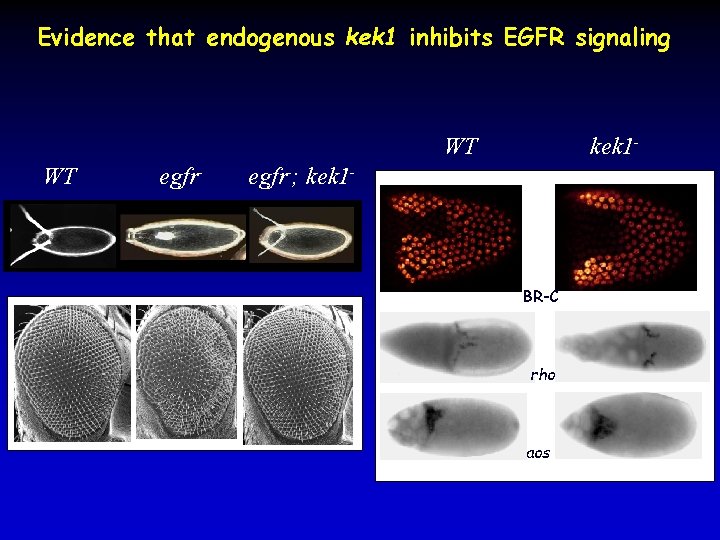
Evidence that endogenous kek 1 inhibits EGFR signaling WT WT egfr- kek 1 - egfr-; kek 1 - BR-C rho aos
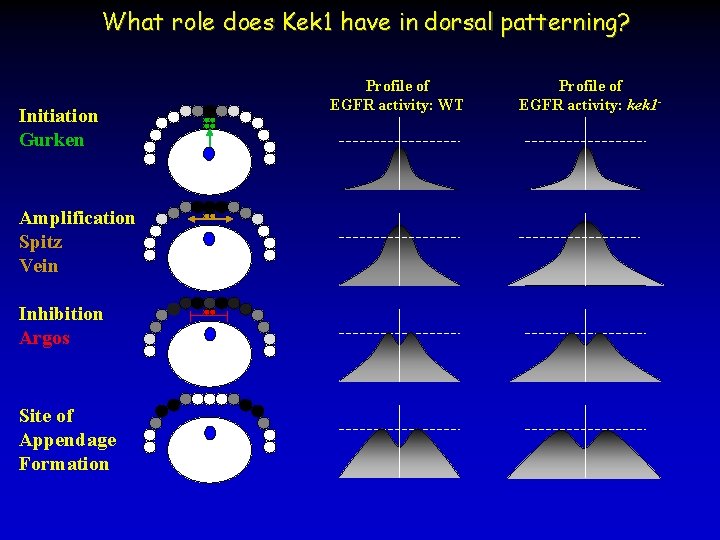
What role does Kek 1 have in dorsal patterning? Initiation Gurken Amplification Spitz Vein Inhibition Argos Site of Appendage Formation Profile of EGFR activity: WT Profile of EGFR activity: kek 1 -
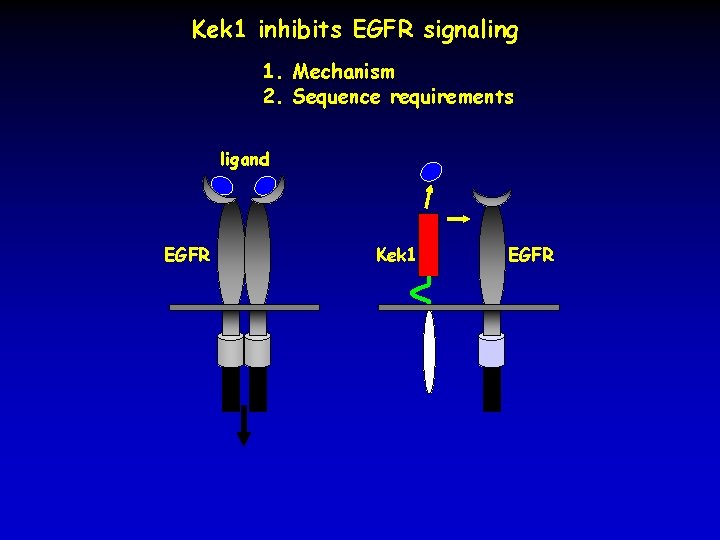
Kek 1 inhibits EGFR signaling 1. Mechanism 2. Sequence requirements ligand EGFR Kek 1 EGFR
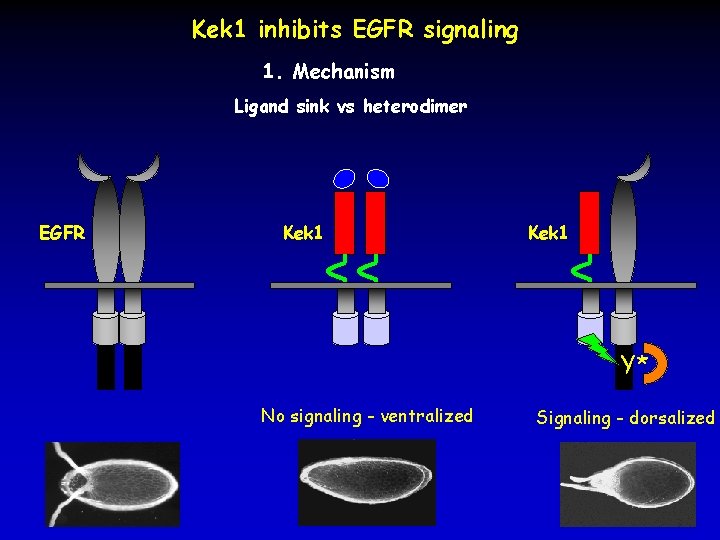
Kek 1 inhibits EGFR signaling 1. Mechanism Ligand sink vs heterodimer EGFR Kek 1 Y* No signaling - ventralized Signaling - dorsalized
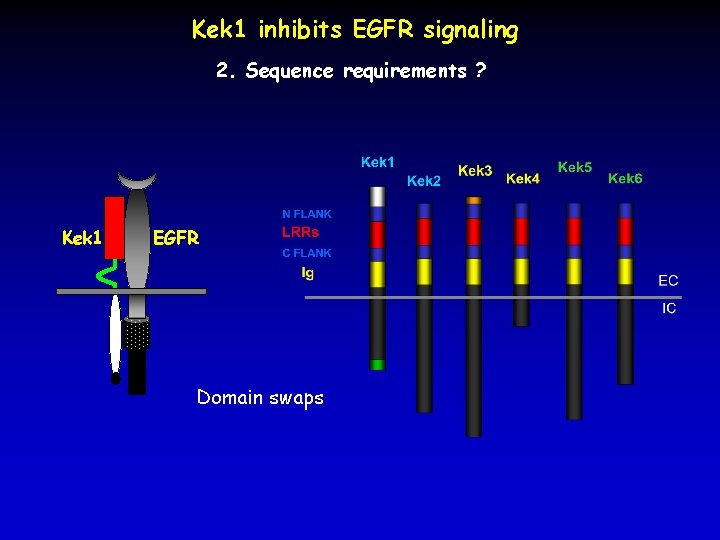
Kek 1 inhibits EGFR signaling 2. Sequence requirements ? Kek 1 EGFR Domain swaps
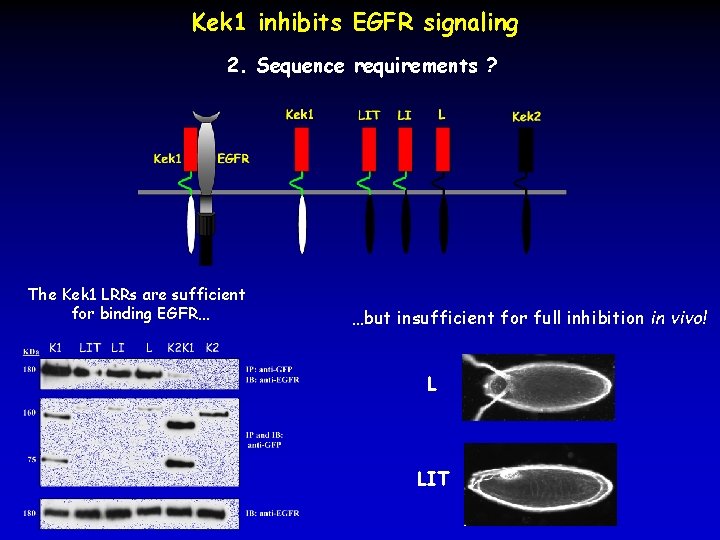
Kek 1 inhibits EGFR signaling 2. Sequence requirements ? The Kek 1 LRRs are sufficient for binding EGFR… …but insufficient for full inhibition in vivo! L LIT
![Identifying residues important for Kek 1/EGFR interaction EMS [GMR-GAL 4] [UAS-kek 1 gfp] __________ Identifying residues important for Kek 1/EGFR interaction EMS [GMR-GAL 4] [UAS-kek 1 gfp] __________](http://slidetodoc.com/presentation_image_h2/cbd67a285a82b2d78d53f2c4c2dd452f/image-13.jpg)
Identifying residues important for Kek 1/EGFR interaction EMS [GMR-GAL 4] [UAS-kek 1 gfp] __________ Cy. O w; iso 2; iso 3 P suppressor F 1 [GMR-GAL 4*] [UAS-kek 1*gfp] __________ +
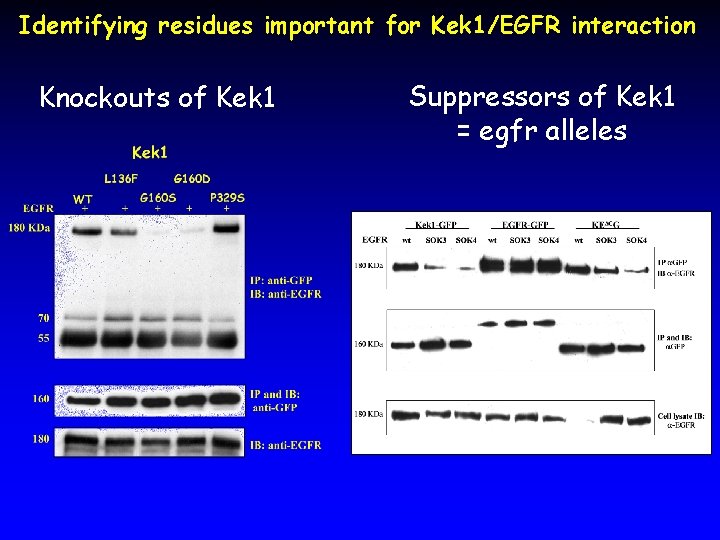
Identifying residues important for Kek 1/EGFR interaction Knockouts of Kek 1 Suppressors of Kek 1 = egfr alleles
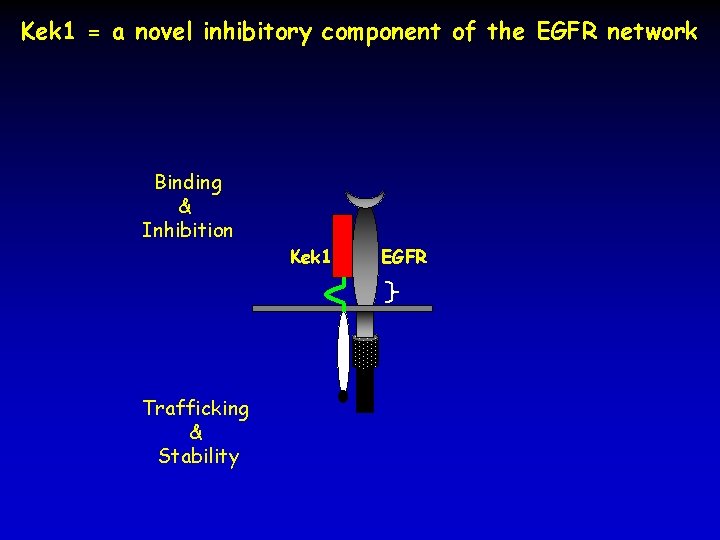
Kek 1 = a novel inhibitory component of the EGFR network Binding & Inhibition Trafficking & Stability Kek 1 EGFR
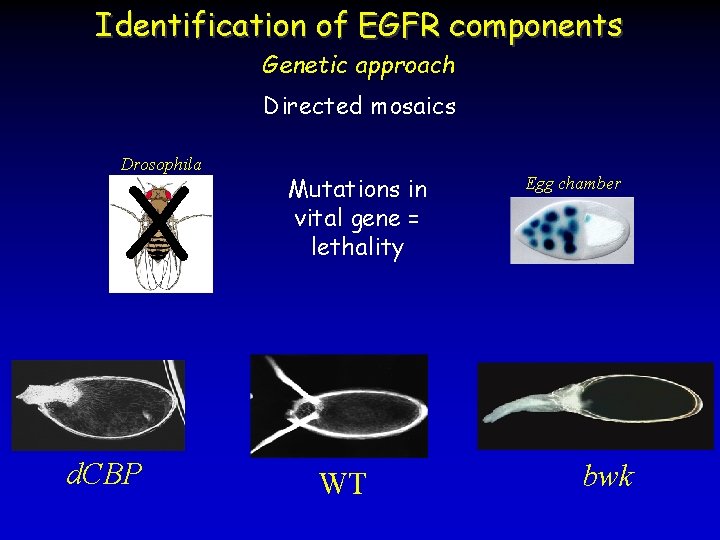
Identification of EGFR components Genetic approach Directed mosaics X Drosophila d. CBP Mutations in vital gene = lethality WT Egg chamber bwk
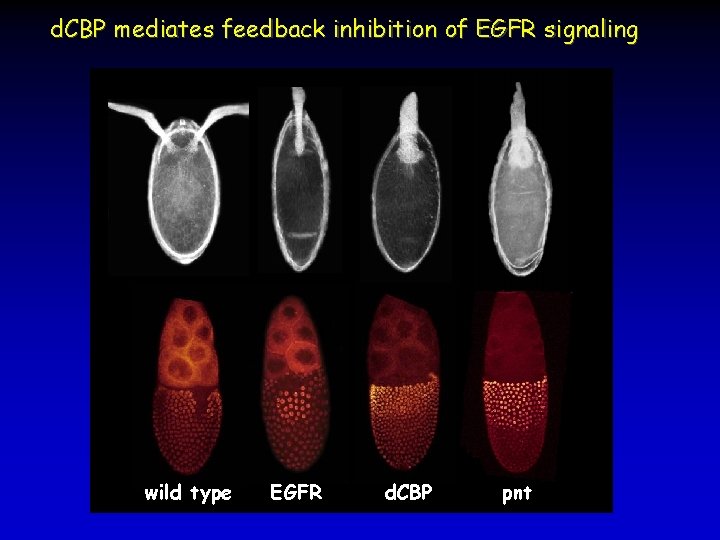
d. CBP mediates feedback inhibition of EGFR signaling wild type EGFR d. CBP pnt
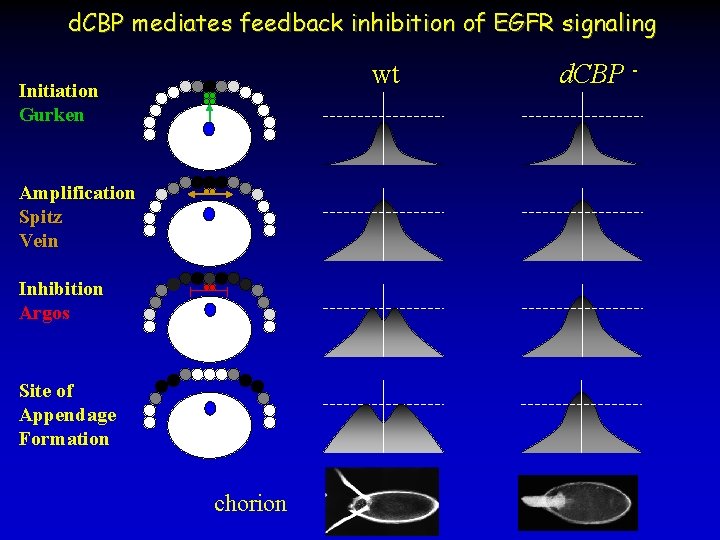
d. CBP mediates feedback inhibition of EGFR signaling wt Initiation Gurken Amplification Spitz Vein Inhibition Argos Site of Appendage Formation chorion d. CBP -
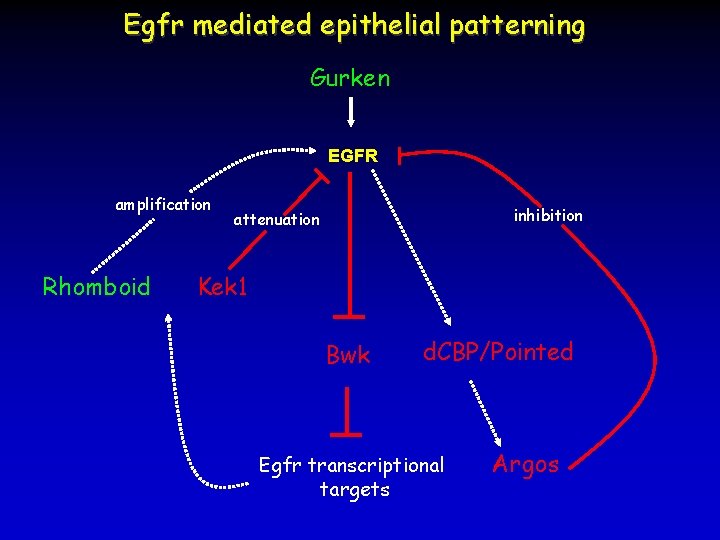
Egfr mediated epithelial patterning Gurken EGFR amplification Rhomboid inhibition attenuation Kek 1 Bwk d. CBP/Pointed Egfr transcriptional targets Argos
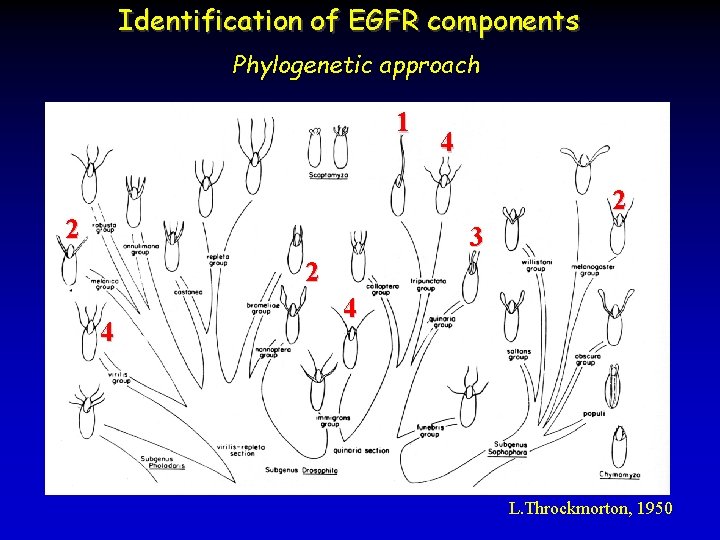
Identification of EGFR components Phylogenetic approach 1 4 2 2 3 2 4 4 latifasciaformis L. Throckmorton, 1950
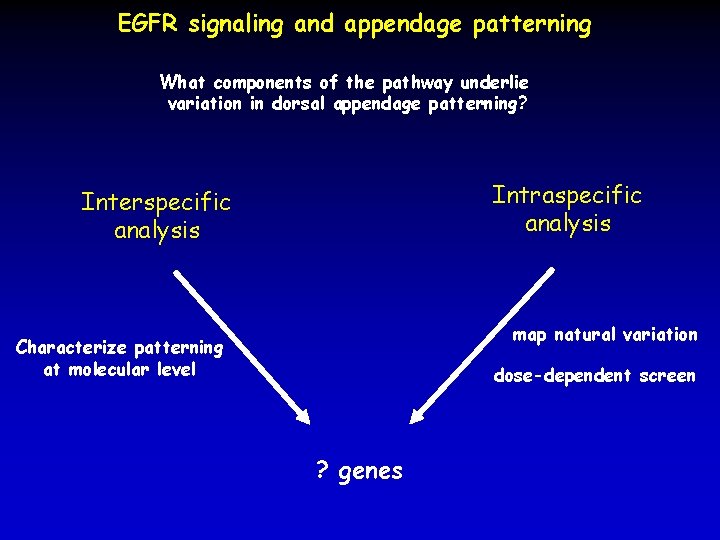
EGFR signaling and appendage patterning What components of the pathway underlie variation in dorsal appendage patterning? Intraspecific analysis Interspecific analysis map natural variation Characterize patterning at molecular level dose-dependent screen ? genes
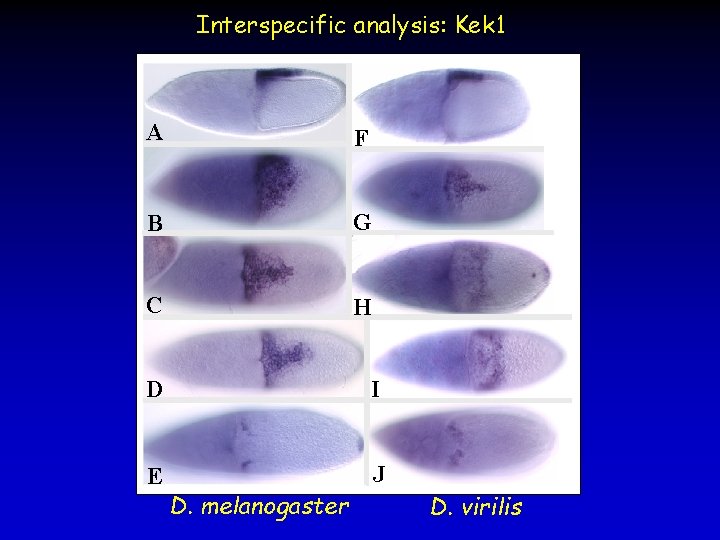
Interspecific analysis: Kek 1 A F B G C H D I E J D. melanogaster D. virilis
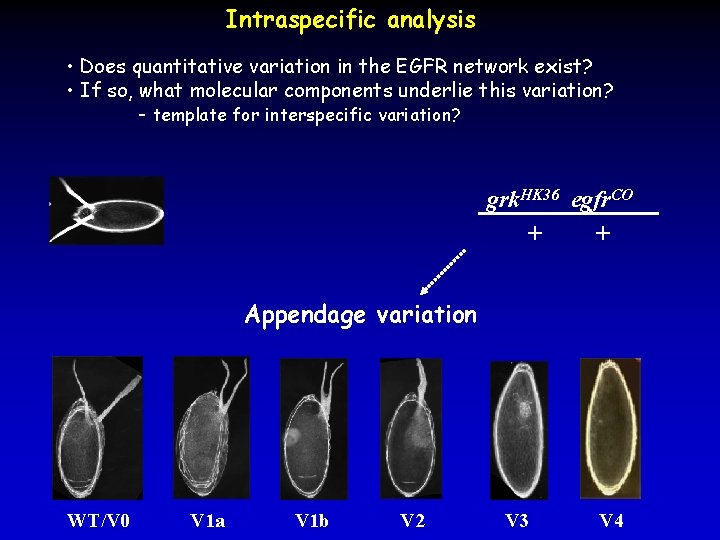
Intraspecific analysis • Does quantitative variation in the EGFR network exist? • If so, what molecular components underlie this variation? - template for interspecific variation? grk. HK 36 egfr. CO + + Appendage variation WT/V 0 V 1 a V 1 b V 2 V 3 V 4
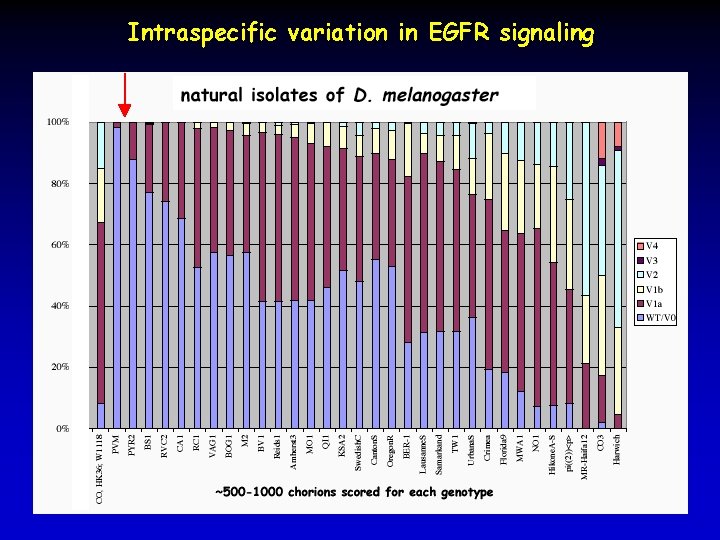
Intraspecific variation in EGFR signaling
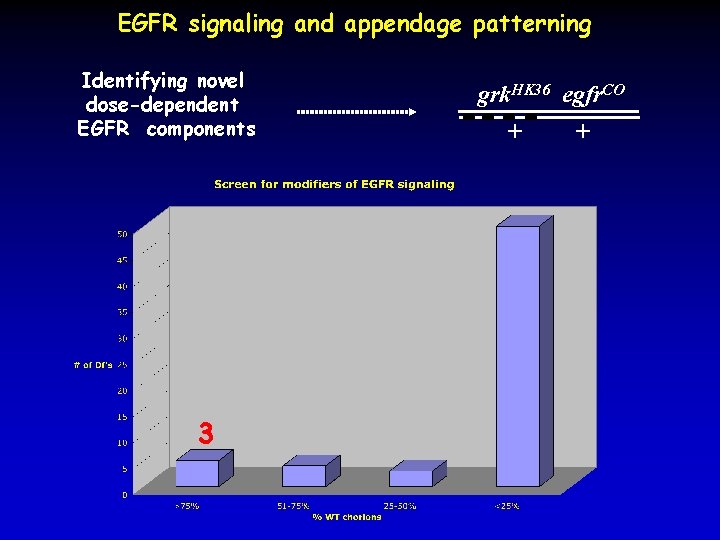
EGFR signaling and appendage patterning Identifying novel dose-dependent EGFR components 3 grk. HK 36 egfr. CO + +
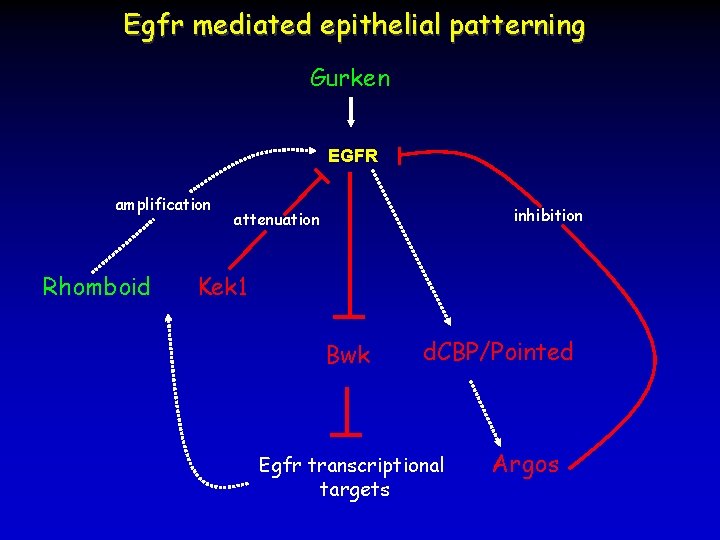
Egfr mediated epithelial patterning Gurken EGFR amplification Rhomboid inhibition attenuation Kek 1 Bwk d. CBP/Pointed Egfr transcriptional targets Argos

Diego Alvarado Alex Doetsch Tim Evans Christina Mac. Laren Amy Rice Anne Scherer Brandon Weasner Andrea Cowden Omar Hasnie Matt Rubin Kyle Payne Fred Derheimer Libbie Eakin Paul Miller Christopher Skipwith
- Slides: 27