The DNA Sequence of chimpanzee chromosome 22 and



















- Slides: 25

The DNA Sequence of chimpanzee chromosome 22 and comparative analysis with its human ortholog, chromosome 21 Bioinformatics Dae-Soo Kim MPL

Comparative analysis of Human and chimpanzee genome Human-chimp comparative genome research is essential for narrowing down the genetic change involved in the acquisitions of unique human features £ We report the high quality DNA sequence of 33. 3 Mb of chimpanzee chromosome 22. £ 1. 44% of the chromosome consisted of single base substitutions in addition to nearly 68, 000 INDEL £ 83% of the 231 coding sequence show difference at the amino acid sequence level. £ BIOINFORMATICS MPL

Introduction £ Estimates of nucleotide substitution rates of aligned sequences were quite ranging from 1. 23% by BAC end sequencing to about 2% by molecular analysis £ Molecular analysis of HSA 21 and its genes is of central medical interest because of trisomy 21, the most common genetic cause of metal retardation in the human population. BIOINFORMATICS MPL

Mapping, sequencing and global view of chimpanzee chromosome 22 £ Genomic DNA origination from three male chimpanzee individuals. £ Sequence coverage of the euchromatic potion of the long arm of chromosome 22 is 98. 6%. £ Accuracy was calculated as 99. 99% from the overlap clone sequence BIOINFORMATICS MPL

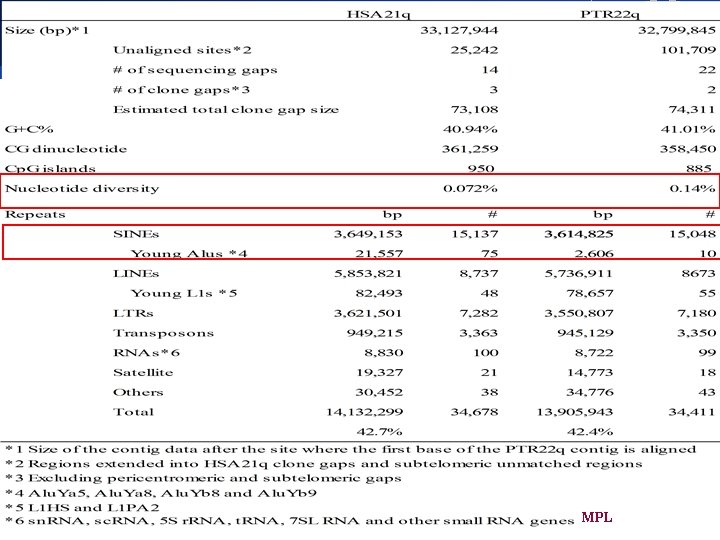
Overall differences The overall structural features of PTR 22 are almost the same as those of HSA 21. £ About a 400 kb or 1. 2% difference in size with HSA 21 being larger then PTR 22 (ISRs; 53. 7% and simple repeats; 9. 54%) £ The pericentromeric copy of a 200 kb region found duplicated in HSA 21 is missing in PTR 22 £ We also detected apparently human specific sequences (first intron PFKL of HSA 21 a) £ BIOINFORMATICS MPL

Two large indel hot spots werw found around 9. 5~11. 5 Mb and 16. 5~17. 5 Mb from the centromere £ We found large human insertion/chimpanzee deletions in the first introns of the NCAM 2(~10 kb)and GRIK 1(~4 kb) (Neural functions) £ One of the largest structural changes identified here is a 54 kb region located at 11. 4 Mb from the centromere in HSA 21 but absent in PTR 22. (flanked by HSAT 5 satellite repeat and consists of 164 fragments from 64 different LTR) £ BIOINFORMATICS MPL

BIOINFORMATICS MPL

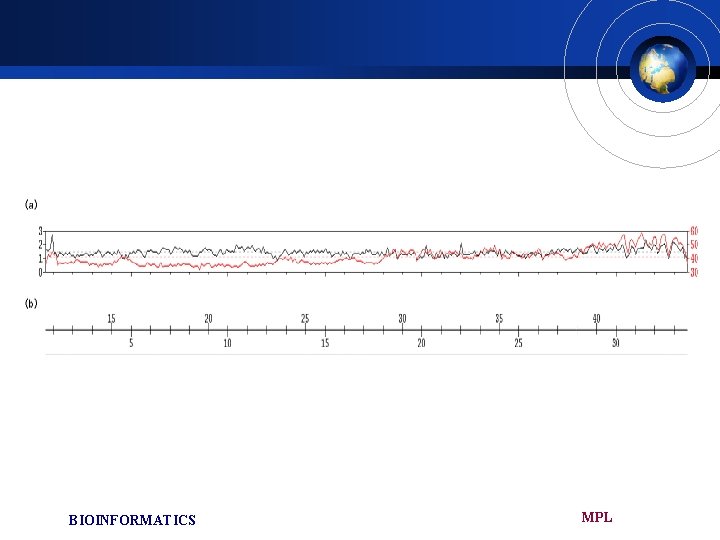
Base substitutions £ The overall nucleotide substitution level in aligned regions between PTR 22 and HSA 21 is about 1. 44%(excluding INDEL) £ The most conserved region was around 12. 5 Mb corresponding to the distal boundary region of the gene desert. BIOINFORMATICS MPL

BIOINFORMATICS MPL

Repetitive elements £ HSA 21 is about 1. 2% longer in size than PTR 22 £ Five LTR subfamilies LTR are more abundant in HSA 21 £ All MER 4 A 1 -int and MER 83 B-int elements are specific to HSA 21 £ All of the seven Alu. Yb 9’s found in HSA 21 and the one in PTR 22 are lineage specific £ Although the Alu. Ya 8 subfamily is though to be a recent derivative of Alu. Ya 5 MPL BIOINFORMATICS

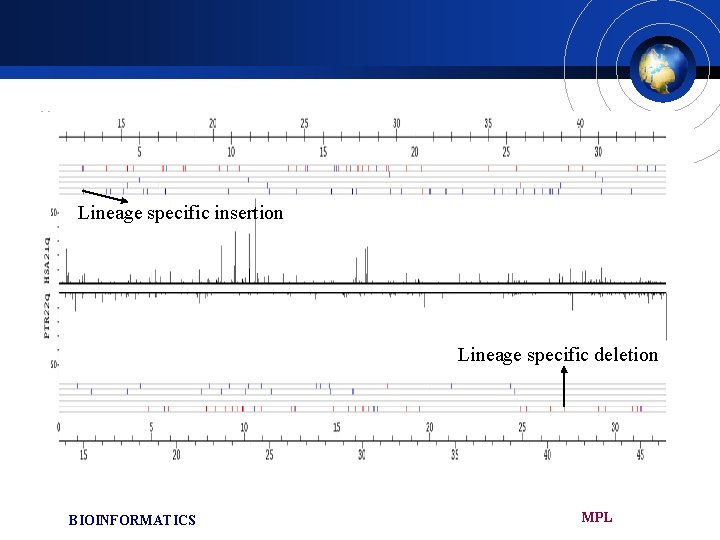
Lineage specific insertions and deletions £ We identified about 68, 000 INDEL is total £ Greater than 99% of the INDELs were shorter than 300 bp £ These site should be produced either through h-ins/p-dels or p-ins/h-dels £ We tested 567 INDEL larger than 300 bp in size using DNA samples from 5 human , 5 chimpanzee , 1 gorilla, 2 orangutan £ Insertions being mostly produced by the integration of Alu and L 1 elements MPL BIOINFORMATICS

BIOINFORMATICS MPL

Lineage specific insertion Lineage specific deletion BIOINFORMATICS MPL

BIOINFORMATICS MPL

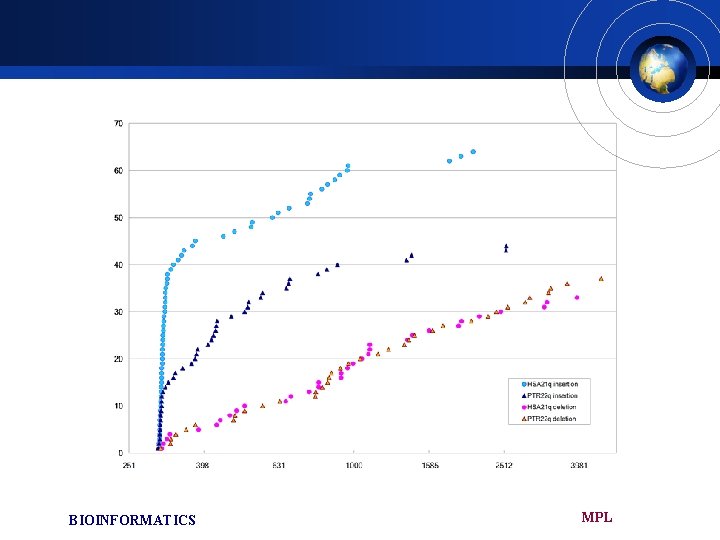
£ Deletions not being related to particular repetitive structures except for a few cases. £ We found that most of the insertions 300350 bp in length were members of Alu. Y family in both chromosome £ Between 370 -1000 bp only a smaller number of insertions mostly L 1 and LTR £ We observed that the distribution of newly integrated Alu are quit different between HSA 21 and PTR 22 (HSA 21; 56% high G+C , PTR 22; 70% low G+C) MPL BIOINFORMATICS

Unlike the insertion, deletions do not exactly correspond to any ISR elements, indicating that deletion events are independent of ISRs. £ The deletion of these elements may have also been generated by homologous recombination between these relatively short identical or similar flanking segments. £ HSA 21 gained 32 kb but lost 39 kb while PTR 22 gained 25 kb and lost 53 kb (INDEL 300~5000 bp) £ PTR 22 has suffered more losses than HSA 21 since speciation. £ BIOINFORMATICS MPL

£A neighbor joining analysis show that such Alu. Y elements can be largely separated into chimp and human groups as expected (Alu. Y was inserted after speciation) £ Humans seem to have experienced such expansions more frequently and more recently than chimp BIOINFORMATICS MPL

BIOINFORMATICS MPL

Gene catalogue and structural characterization of coding sequences £ £ £ We have annotated 284 protein coding genes and 98 pseudogenes for HSA 21 and 272 genes and 89 pseudogenes for PTR 22 All the conserved pseudogenes showed the same size except for KRTAP 21 P 1 which is non processed in HSA 21 but processed in PTR 22 Six HSA 21 genes showing hallmarks of retrogenes were not found in PTR 22 and are likely to have inserted during human evolution (H 2 BFS; histon family S, 5 keratin associated protein) The minimum nucleotide sequence identity is 83%(KRTAP 6 -3) and the maximum is 100% We compared the human and chimp coding sequences in 231 genes (omitted 41) BIOINFORMATICS MPL

Among the 231 genes associated to a canonical ORF 179 show a coding sequence of identical length in human and chimpanzee and exhibit similar intron-exon boundaries £ 39 genes shown an identical amino acid and nucleotide sequence between human and chimp (biological process 5, metabolic enzymes 5, signal transduction 8, protein folding 2) £ One hundred and forty out of these 179 genes show amino acid replacements but no gross structural changes and expected. £ BIOINFORMATICS MPL

Ka/Ks analysis £ 10% of the genes had Ka/Ks rations >1 with the highest value being 3. 37 for the human hair keratine associated protein £ Relatively rapidly evolving genes may be estimated from Ka, Ka+Ks or just nucleotide divergence values. (3 KRTAP gene, KCNE 1; potassium channel protein , TCP 10 L; complex protein, B 3 GALT 5; galctocyltransferase, IGSF 5; immu noglobulin) BIOINFORMATICS MPL

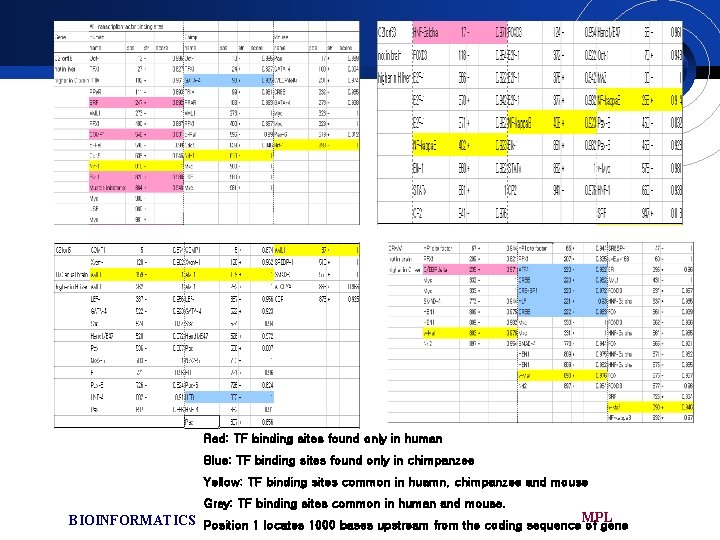
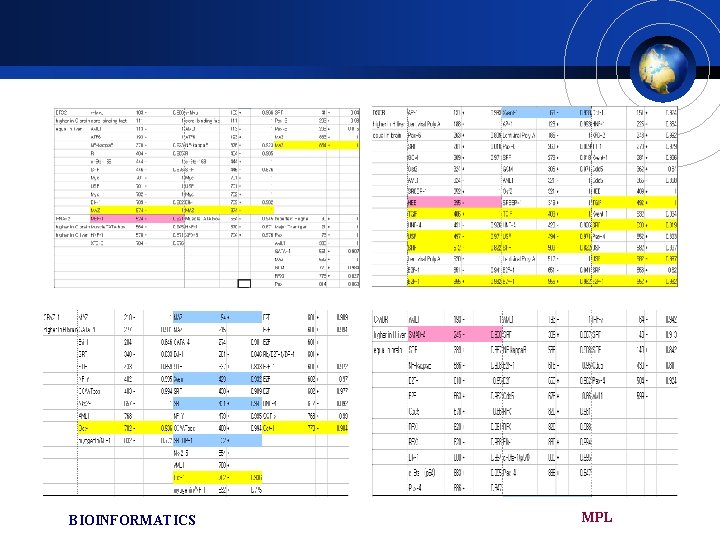
Promoter analysis £ Computation analysis of the transcription factor binding site within the l-kb upstream region of each gene. £ All of the specific TFBSs were caused by base substitution in either human or chimpanzee £ These may mot clearly account for the expression changes observed in this study BIOINFORMATICS MPL

Red: TF binding sites found only in human Blue: TF binding sites found only in chimpanzee Yellow: TF binding sites common in huamn, chimpanzee and mouse Grey: TF binding sites common in human and mouse. BIOINFORMATICS Position 1 locates 1000 bases upstream from the coding sequence MPL of gene

BIOINFORMATICS MPL

Conclusion £ This study shows for the first time a chromosome wide comparison between human and chimpanzee using high quality sequence. BIOINFORMATICS MPL