The db system and data analysis tools data

The db system and data analysis tools • data extraction, averaging, exporting; • using templates for statistical analyses and plots, • documentation and FAQs

Data extraction How do I get data from one or more instruments into a data file? Need to know what data you want! (a) what instrument(s)? (b) what averaging time? (c) what processing (e. g. , raw/edited)? (d) what format? How to figure out instrument names in the db system data. instruments. get stn_id station flow diagram cpx 2 Data Records and Data Variables http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/cpd 2 record. html http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/cpd 2 variables. html Types of archives Raw, edited, clean, averaged
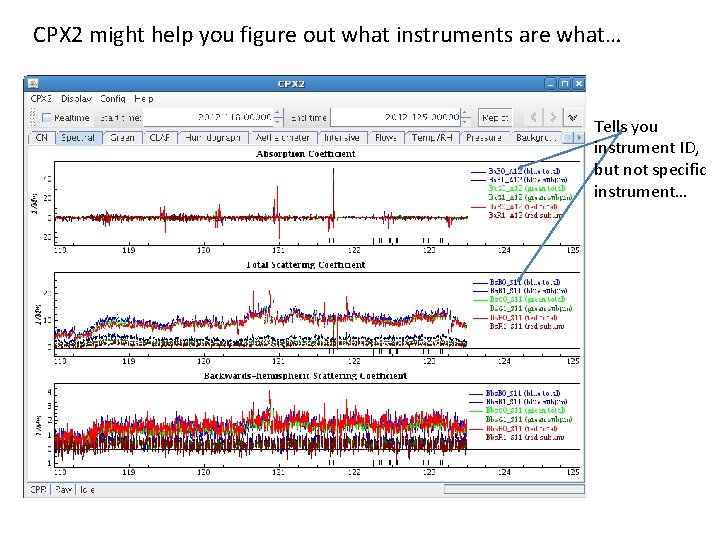
CPX 2 might help you figure out what instruments are what… Tells you instrument ID, but not specific instrument…
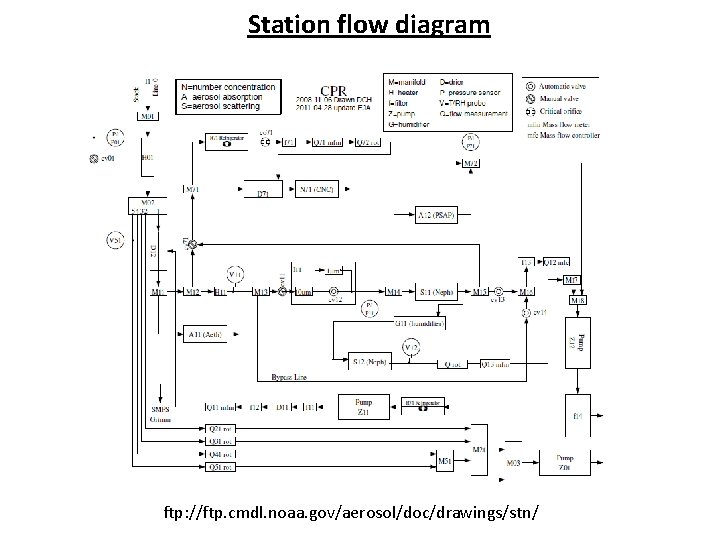
Station flow diagram ftp: //ftp. cmdl. noaa. gov/aerosol/doc/drawings/stn/

Best way to figure out what’s what! In a terminal window in Aer_vm, type: data. instruments. get stn 2009 Q 4 2010 Q 1 2010 Q 2 2010 Q 3 2010 Q 4 2011 Q 1 2011 Q 2 2011 Q 3 2011 Q 4 2012 Q 1 2012 Q 2 S 11 N 71 A 12 S 12 A 13 A 11 | | | N 71 | | | S 11 | | | A 12 | | | 2004 -11 -24 T 00: 00: 00 Z 2004 -11 -24 T 00: 00 Z 2006 -05 -27 T 00: 00 Z 2011 -04 -10 T 00: 00 Z 2012 -03 -30 T 18: 05: 07 Z S 12 A 13 | | | Where stn is your station’s 3 -letter id (e. g. , mlo, app, cpr, etc. ) | | TSI; 3563; 70417325: Installed TSI; 3022; 608: Installed Magee; Aethalometer-AE 31; 0202: Installed RR; PSAP-3 W; 92: Installed RR; M 903; 424: Installed GMD; CLAP-3 W; 18: Installed Current Instruments: S 11 TSI; 3563; 70417325 N 71 TSI; 3022; 608 A 11 Magee; Aethalometer-AE 31; 0202 A 12 RR; PSAP-3 W; 92 S 12 RR; M 903; 424 A 13 GMD; CLAP-3 W; 18

Record names in cpd 2 and the DB-system • • • Two-part naming convention First part is instrument identifier ("S 11") Second part is record type Most common record type to extract – a high-frequency – m monitor (low-frequency, housekeeping) – r raw – Others… Example: TSI Neph “S” records a High frequency. Scattering, backscattering, temperature, RH and pressure. m Low frequency. Status: voltages and reference. s Statewise. Flags and status string. c Zero results. Background k Spancheck results. So extracting “S 11 k” records would get nephelometer span check records Details at: http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/cpd 2 record. html

Variable names in cpd 2 and the DB-system • Two part names Optical property Number Conc. – variable identifier • first letter is variable category (e. g. , "B", "N", “Q") • various modifiers are used – wavelength, size cut, and more… Flow – instrument identifier • first character is instrument category ("S") • second character is "line" identifier • third character is sequence number in "line“ Bs. G 0_S 11 = optical property scattering, green wavelength, 10 um, neph S 11 Bbs. R_S 12 = optical property backscattering, red wavelength, neph S 12 Q_A 11 = flow, absorption instrument A 11 – details at: http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/cpd 2 variables. html

Types of data in the db system raw - no corrections, no edits. Raw data exists as a physical archive. edited - edits and corrections, but has not (necessarily) been approved and put into the clean archive. Edited data does not exist as a physical archive clean - edits and corrections and has been approved by the station mentor. Clean data exists as a physical archive and is the source of most final data products. avg[HDM] averaged clean data, excluding records flagged as contaminated, with one record for each hour/day/month, with everything on line one split on cut size, and including standard deviations and counts for each variable. avg. He , avg. De, avg. Me As above, except from edited data instead of clean. how to get averaged data with corrections applied even if you haven’t had a chance to edit it yet…

Where can I find help? Software webpages: http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/ (these webpages are also accessible under help menu on aer_vm) Tutorial ftp: //ftp. cmdl. noaa. gov/aerosol/doc/software/ How_to_Get_Data_Out_of_the_Aero_Database. pptx FAQ http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/dbfaq. htm

HOW TO: Extract data in Excel format • Constant format files 'xt 2' creates files in comma-separated format with default name xt 2 --header --clean -h cpr 2005 2006 Makes a file called ‘al_HX. cpr’ in the current directory Advantages has neph, PSAP, CN data always has same format simple to remember Disadvantages limited to neph, PSAP, CN limited to min. , H, D, M can’t use other data base tools

HOW TO: Extract data in Excel format New style "cpd 2" formats Combine 'data. get' to select data with 'data. export' to get data file in desired format: “pipe” data. get cpr S 11 a 2008: 10 2008: 11 | data. export --mode=excel > data. csv db command record station Start time End time Advantages data from any instrument any averaging time can use db tools db command File format option File name Disadvantages complicated to remember format changes depending on instrument(s) and extraction modes
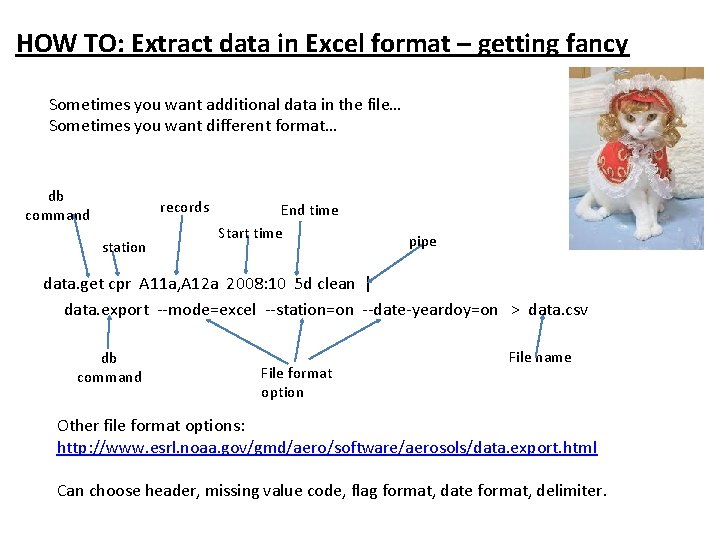
HOW TO: Extract data in Excel format – getting fancy Sometimes you want additional data in the file… Sometimes you want different format… db command records station End time Start time pipe data. get cpr A 11 a, A 12 a 2008: 10 5 d clean | data. export --mode=excel --station=on --date-yeardoy=on > data. csv db command File format option File name Other file format options: http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/data. export. html Can choose header, missing value code, flag format, date format, delimiter.
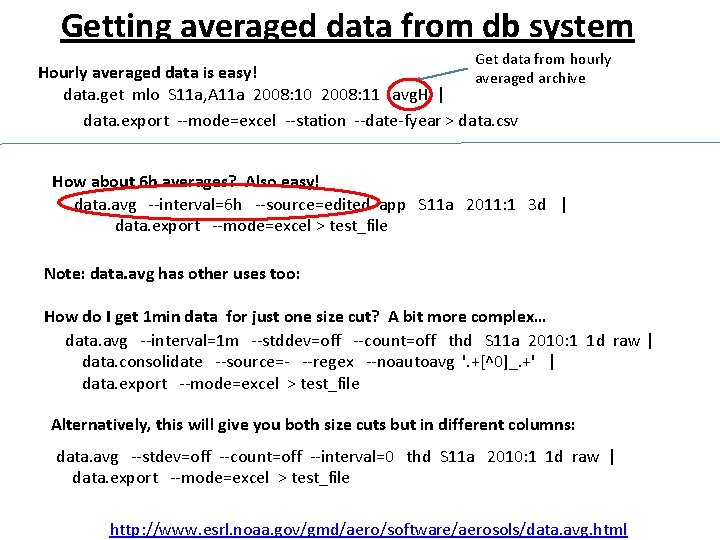
Getting averaged data from db system Get data from hourly Hourly averaged data is easy! averaged archive data. get mlo S 11 a, A 11 a 2008: 10 2008: 11 avg. H | data. export --mode=excel --station --date-fyear > data. csv How about 6 h averages? Also easy! data. avg --interval=6 h --source=edited app S 11 a 2011: 1 3 d | data. export --mode=excel > test_file Note: data. avg has other uses too: How do I get 1 min data for just one size cut? A bit more complex… data. avg --interval=1 m --stddev=off --count=off thd S 11 a 2010: 1 1 d raw | data. consolidate --source=- --regex --noautoavg '. +[^0]_. +' | data. export --mode=excel > test_file Alternatively, this will give you both size cuts but in different columns: data. avg --stdev=off --count=off --interval=0 thd S 11 a 2010: 1 1 d raw | data. export --mode=excel > test_file http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/data. avg. html

data. get cpr A 11 a, A 12 a, A 13 a 2012: 116 1 d avg. He | data. export --mode=excel > test_file Date. Time. UTC, F 1_A 12, F 1_A 13, F 2_A 12, F 2_A 13, Ba. B 0_A 12, Ba. B 0_A 13, Ba. G 0_A 12, Ba. G 0_A 13, Ba. R 0_A 12, Ba. R 0_A 13, Ba. O 0_A 12, Ba. O 0_A 13, Ba. B 1_A 12, Ba. B 1_A 13, Ba. G 1_A 12, Ba. G 1_A 13, Ba. R 1_A 12, Ba. R 1_A 13, Ba. O 1_A 12, Ba. O 1_A 13, Ba. B 0 g_A 12, Ba. B 0 g_A 13, Ba. G 0 g_A 12, Ba. G 0 g_A 13, Ba. R 0 g_A 12, Ba. R 0 g_A 13, Ba. O 0 g_A 12, Ba. O 0 g_A 13, Ba. B 1 g_A 12, Ba. B 1 g_A 13, Ba. G 1 g_A 12, Ba. G 1 g_A 13, Ba. R 1 g_A 12, Ba. R 1 g_A 13, Ba. O 1 g_A 12, Ba. O 1 g_A 13, Ba. B 0 N_A 12, Ba. B 0 N_A 13, Ba. G 0 N_A 12, Ba. G 0 N_A 13, Ba. R 0 N_A 12, Ba. R 0 N_A 13, Ba. O 0 N_A 12, Ba. O 0 N_A 13, Ba. B 1 N_A 12, Ba. B 1 N_A 13, Ba. G 1 N_A 12, Ba. G 1 N_A 13, Ba. R 1 N_A 12, Ba. R 1 N_A 13, Ba. O 1 N_A 12, Ba. O 1 N_A 13, If 1_A 11, If 2_A 11, If 3_A 11, If 4_A 11, If 5_A 11, If 6_A 11, If 7_A 11, Ifz 1_A 11, Ifz 2 _A 11, Ifz 3_A 11, Ifz 4_A 11, Ifz 5_A 11, Ifz 6_A 11, Ifz 7_A 11, Ip 1_A 11, Ip 2_A 11, Ip 3_A 11, Ip 4_A 11, Ip 5_A 11, Ip 6_A 11, Ip 7_A 11, Ipz 1_A 11, Ipz 2_A 11, Ipz 3_A 11, Ipz 4_A 11, Ipz 5_A 11, Ipz 6_A 11, Ipz 7_A 11, PCT_A 11, Q_A 11, X 1 c_A 11, X 2 c_A 11, X 3 c_A 11, X 4 c_A 11, X 5 c_A 11, X 6 c_A 11, X 7 c_A 11, ZIr 1_A 11, ZIr 2_A 11, ZIr 3_A 11, ZIr 4_A 11, ZIr 5_A 11, ZIr 6_A 11, ZIr 7_A 11, Ff 0_A 12, Ff 0_A 13, Fn 0_A 13, Ir. B 0_A 12, Ir. B 0_A 13, Ir. G 0_A 12, Ir. G 0_A 13, Ir. R 0_A 12, Ir. R 0 _A 13, L 0_A 12, L 0_A 13, Ff 1_A 12, Ff 1_A 13, Fn 1_A 13, Ir. B 1_A 12, Ir. B 1_A 13, Ir. G 1_A 12, Ir. G 1_A 13, Ir. R 1_A 12, Ir. R 1_A 13, L 1_A 12, L 1_A 13, If 1 g_A 11, If 2 g _A 11, If 3 g_A 11, If 4 g_A 11, If 5 g_A 11, If 6 g_A 11, If 7 g_A 11, Ifz 1 g_A 11, Ifz 2 g_A 11, Ifz 3 g_A 11, Ifz 4 g_A 11, Ifz 5 g_A 11, Ifz 6 g_A 11, Ifz 7 g_A 11, Ip 1 g_A 11, I p 2 g_A 11, Ip 3 g_A 11, Ip 4 g_A 11, Ip 5 g_A 11, Ip 6 g_A 11, Ip 7 g_A 11, Ipz 1 g_A 11, Ipz 2 g_A 11, Ipz 3 g_A 11, Ipz 4 g_A 11, Ipz 5 g_A 11, Ipz 6 g_A 11, Ipz 7 g_A 11, P CTg_A 11, Qg_A 11, X 1 cg_A 11, X 2 cg_A 11, X 3 cg_A 11, X 4 cg_A 11, X 5 cg_A 11, X 6 cg_A 11, X 7 cg_A 11, ZIr 1 g_A 11, ZIr 2 g_A 11, ZIr 3 g_A 11, ZIr 4 g_A 11, ZIr 5 g_A 11, ZIr 6 g_A 11, ZIr 7 g_A 11, Ff 0 g_A 12, Ff 0 g_A 13, Fn 0 g_A 13, Ir. B 0 g_A 12, Ir. B 0 g_A 13, Ir. G 0 g_A 12, Ir. G 0 g_A 13, Ir. R 0 g_A 12, Ir. R 0 g_A 13, L 0 g_A 12, L 0 g _A 13, Ff 1 g_A 12, Ff 1 g_A 13, Fn 1 g_A 13, Ir. B 1 g_A 12, Ir. B 1 g_A 13, Ir. G 1 g_A 12, Ir. G 1 g_A 13, Ir. R 1 g_A 12, Ir. R 1 g_A 13, L 1 g_A 12, L 1 g_A 13, F 2 N_A 12, F 2 N_A 1 3, If 1 N_A 11, If 2 N_A 11, If 3 N_A 11, If 4 N_A 11, If 5 N_A 11, If 6 N_A 11, If 7 N_A 11, Ifz 1 N_A 11, Ifz 2 N_A 11, Ifz 3 N_A 11, Ifz 4 N_A 11, Ifz 5 N_A 11, Ifz 6 N_A 11, If z 7 N_A 11, Ip 1 N_A 11, Ip 2 N_A 11, Ip 3 N_A 11, Ip 4 N_A 11, Ip 5 N_A 11, Ip 6 N_A 11, Ip 7 N_A 11, Ipz 1 N_A 11, Ipz 2 N_A 11, Ipz 3 N_A 11, Ipz 4 N_A 11, Ipz 5 N_A 1 1, Ipz 6 N_A 11, Ipz 7 N_A 11, PCTN_A 11, QN_A 11, X 1 c. N_A 11, X 2 c. N_A 11, X 3 c. N_A 11, X 4 c. N_A 11, X 5 c. N_A 11, X 6 c. N_A 11, X 7 c. N_A 11, ZIr 1 N_A 11, ZIr 2 N _A 11, ZIr 3 N_A 11, ZIr 4 N_A 11, ZIr 5 N_A 11, ZIr 6 N_A 11, ZIr 7 N_A 11, Ff 0 N_A 12, Ff 0 N_A 13, Fn 0 N_A 13, Ir. B 0 N_A 12, Ir. B 0 N_A 13, Ir. G 0 N_A 12, Ir. G 0 N_A 1 3, Ir. R 0 N_A 12, Ir. R 0 N_A 13, L 0 N_A 12, L 0 N_A 13, F 1 N_A 12, F 1 N_A 13, Ff 1 N_A 12, Ff 1 N_A 13, Fn 1 N_A 13, Ir. B 1 N_A 12, Ir. B 1 N_A 13, Ir. G 1 N_A 12, Ir. G 1 N_A 13, Ir. R 1 N_A 12, Ir. R 1 N_A 13, L 1 N_A 12, L 1 N_A 13

How do I get only absorption and not any of the crazy variables that I won’t look at anyway? data. consolidate --source=avg. H cpr 115 116 'Ba*_A 12' | data. export --mode=excel > myfile Date. Time. UTC, Ba. B 0_A 12, Ba. G 0_A 12, Ba. R 0_A 12, Ba. O 0_A 12, Ba. B 1_A 12, Ba. G 1_A 12, Ba. R 1_A 12, Ba. O 1_A 12, Ba. B 0 g_A 12, Ba. G 0 g_A 12, Ba. R 0 g_A 12, Ba. O 0 g_A 12, Ba. B 1 g_A 12, Ba. G 1 g_A 12, Ba. R 1 g_A 12, Ba. O 1 g_A 12, Ba. B 0 N_A 12, Ba. G 0 N_A 12, Ba. R 0 N_A 12, Ba. O 0 N_A 12, Ba. B 1 N_A 12, Ba. G 1 N_A 12, B a. R 1 N_A 12, Ba. O 1 N_A 12 POP-QUIZ! What does variable Ba. R 0_A 12 represent? ? What about Ba. O 0 g_A 12?

I don’t use excel…microsoft is the evil empire! data. get cpr A 12 a 115 116 avg. He | data. export --mode=idl > test_file Julian. Day: 0 Year: 1 DOY: 2 F 1_A 12_Bit 0: 3 F 1_A 12_Bit 1: 4 F 1_A 12_Bit 2: 5 F 1_A 12_Bit 3: 6 F 1_A 12_Bit 4: 7 F 1_A 12_Bit 5: 8 F 1_A 12_Bit 6: 9 F 1_A 12_Bit 7: 10 F 1_A 12_Bit 8: 11 F 1_A 12_Bit 9: 12 F 1_A 12_Bit 10: 13 F 1_A 12_Bit 11: 14 F 1_A 12_Bit 12: 15 F 1_A 12_Bit 13: 16 F 1_A 12_Bit 14: 17 F 1_A 12_Bit 15: 18 F 2_A 12_Bit 0: 19 F 2_A 12_Bit 1: 20 F 2_A 12_Bit 2: 21 F 2_A 12_Bit 3: 22 F 2_A 12_Bit 4: 23 F 2_A 12_Bit 5: 24 F 2_A 12_Bit 6: 25 F 2_A 12_Bit 7: 26 F 2_A 12_Bit 8: 27 F 2_A 12_Bit 9: 28 F 2_A 12_Bit 10: 29 F 2_A 12_Bit 11: 30 F 2_A 12_Bit 12: 31 F 2_A 12_Bit 13: 32 F 2_A 12_Bit 14: 33 F 2_A 12_Bit 15: 34 Ba. B 0_A 12: 35 Ba. B 0_A 12_MVC: 36 Ba. G 0_A 12: 37 Ba. G 0_A 12_MVC: 38 Ba. R 0_A 12: 39 Ba. R 0_A 12_MVC: 40 Ba. O 0_A 12: 41 Ba. O 0_A 12_MVC: 42 Ba. B 1_A 12: 43 Ba. B 1_A 12_MVC: 44 Ba. G 1_A 12: 45 Ba. G 1_A 12_MVC: 46 Ba. R 1_A 12: 47 Ba. R 1_A 12_MVC: 48 Ba. O 1_A 12: 49 Ba. O 1_A 12_MVC: 50 Ba. B 0 g_A 12: 51 Ba. B 0 g_A 12_MVC: 52 Ba. G 0 g_A 12: 53 Ba. G 0 g_A 12_MVC: 54 Ba. R 0 g_A 12: 55 Ba. R 0 g_A 12_MVC: 56 Ba. O 0 g_A 12: 57 Ba. O 0 g_A 12_MVC: 58 Ba. B 1 g_A 12: 59 Ba. B 1 g_A 12_MVC: 60 Ba. G 1 g_A 12: 61 Ba. G 1 g_A 12_MVC: 62 Ba. R 1 g_A 12: 63 Ba. R 1 g_A 12_MVC: 64 Ba. O 1 g_A 12: 65 Ba. O 1 g_A 12_MVC: 66 Ba. B 0 N_A 12: 67 Ba. B 0 N_A 12_MVC: 68 Ba. G 0 N_A 12: 69 Ba. G 0 N_A 12_MVC: 70 Ba. R 0 N_A 12: 71 Ba. R 0 N_A 12_MVC: 72 Ba. O 0 N_A 12: 73 Ba. O 0 N_A 12_MVC: 74 Ba. B 1 N_A 12: 75 Ba. B 1 N_A 12_MVC: 76 Ba. G 1 N_A 12: 77 Ba. G 1 N_A 12_MVC: 78 Ba. R 1 N_A 12: 79 Ba. R 1 N_A 12_MVC: 80 Ba. O 1 N_A 12: 81 Ba. O 1 N_A 12_MVC: 82 Ff 0_A 12: 83 Ff 0_A 12_MVC: 84 Ir. B 0_A 12: 85 Ir. B 0_A 12_MVC: 86 Ir. G 0_A 12: 87 Ir. G 0_A 12_MVC: 88 Ir. R 0_A 12: 89 Ir. R 0_A 12_MVC: 90 L 0_A 12: 91 L 0_A 12_MVC: 92 Ff 1_A 12: 93 Ff 1_A 12_MVC: 94 Ir. B 1_A 12: 95 Ir. B 1_A 12_MVC: 96 Ir. G 1_A 12: 97 Ir. G 1_A 12_MVC: 98 Ir. R 1_A 12: 99 Ir. R 1_A 12_MVC: 100 L 1_A 12: 101 L 1_A 12_MVC: 102 Ff 0 g_A 12: 103 Ff 0 g_A 12_MVC: 104 Ir. B 0 g_A 12: 105 Ir. B 0 g_A 12_MVC: 106 Ir. G 0 g_A 12: 107 Ir. G 0 g_A 12_MVC: 108 Ir. R 0 g_A 12: 109 Ir. R 0 g_A 12_MVC: 110 L 0 g_A 12: 111 L 0 g_A 12_MVC: 112 Ff 1 g_A 12: 113 Ff 1 g_A 12_MVC: 114 Ir. B 1 g_A 12: 115 Ir. B 1 g_A 12_MVC: 116 Ir. G 1 g_A 12: 117 Ir. G 1 g_A 12_MVC: 118 Ir. R 1 g_A 12: 119 Ir. R 1 g_A 12_MVC: 120 L 1 g_A 12: 121 L 1 g_A 12_MVC: 122 F 2 N_A 12: 123 F 2 N_A 12_MVC: 124 Ff 0 N_A 12: 125 Ff 0 N_A 12_MVC: 126 Ir. B 0 N_A 12: 127 Ir. B 0 N_A 12_MVC: 128 Ir. G 0 N_A 12: 129 Ir. G 0 N_A 12_MVC: 130 Ir. R 0 N_A 12: 131 Ir. R 0 N_A 12_MVC: 132 L 0 N_A 12: 133 L 0 N_A 12_MVC: 134 F 1 N_A 12: 135 F 1 N_A 12_MVC: 136 Ff 1 N_A 12: 137 Ff 1 N_A 12_MVC: 138 Ir. B 1 N_A 12: 139 Ir. B 1 N_A 12_MVC: 140 Ir. G 1 N_A 12: 141 Ir. G 1 N_A 12_MVC: 142 Ir. R 1 N_A 12: 143 Ir. R 1 N_A 12_MVC: 144 L 1 N_A 12: 145 L 1 N_A 12_MVC: 146 Tells you what columns data are in, file has no header Splits flags column into individual bits

Intensive parameters – can I get those too? All those useful intensive parameters… single scattering albedo, Ångström exponent, etc. Do I really have to do the calculations myself? Nope…there’s a script for that! data. aggregate. intensives sgp XIs 2008: 10 2008: 11 XIs is the ‘short’ intensives record - mostly properties at 550 nm + blue/red Ångström exponent XIl is the ‘long’ intensives record – properties at all the wavelengths Go here to see specifically what is in the intensives record: http: //www. esrl. noaa. gov/gmd/aero/software/aerosols/cpd 2 variables. html#intensives_record

I’ve made this data file – Now what? Move it from the AER_VM to your computer and do with it what you will… --excel --idl --matlab , igor, … --send it to someone… Another option: Use ‘R’ scripts on AER_VM to make several ‘canned’ statistical plots Scripts are in /aer/prg/r/ and /aer/prg/r/examples/ Some useful scripts are: density-plot. r multi-station-box-month. r quantile. r Note: the R scripts are set up to do the data extraction also – so you don’t need to extract data before using the scripts – you need to edit the script to tell it what data to extract. . .

This is what you type to run the script This is the stuff you can change to get the plot you want There is more stuff further down in the script that can be changed too.


Modifying the R script OLD NEW


Practical R stuff Opening the script to edit it on AER_VM use nedit or vi e. g. , type: nedit multi-station-box-month. r <enter> The pound symbol “#” in an R script indicates a comment – the computer ignores everything on the same line after a ‘#’ Getting help with some basic R Get an ‘R’ book using the ‘R’ documentation online info: type: R <enter> at a command line prompt in a AER_VM terminal then type the name of the command you need help with for example type: ? POSIXlt <enter> at the prompt in the R interface to see stuff about time or type: ? read. csv<enter> to see stuff about reading files to get out of the R interface: type: q() <enter> Get a Derek (can’t have ours!)
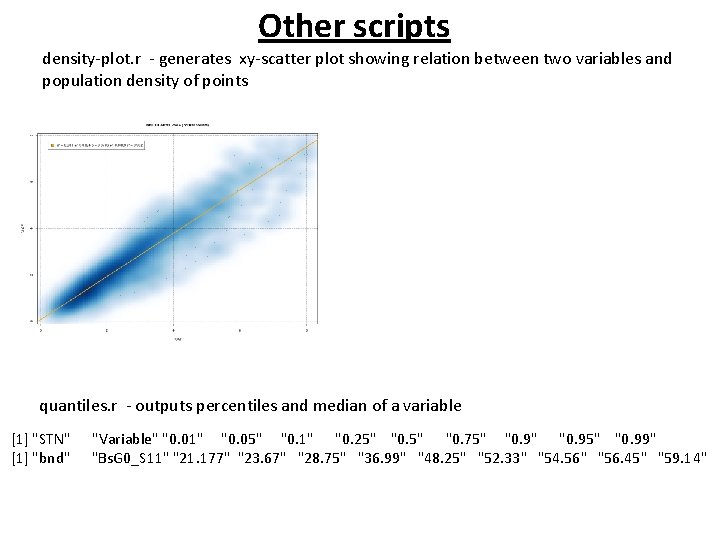
Other scripts density-plot. r - generates xy-scatter plot showing relation between two variables and population density of points quantiles. r - outputs percentiles and median of a variable [1] "STN" [1] "bnd" "Variable" "0. 01" "0. 05" "0. 1" "0. 25" "0. 75" "0. 95" "0. 99" "Bs. G 0_S 11" "21. 177" "23. 67" "28. 75" "36. 99" "48. 25" "52. 33" "54. 56" "56. 45" "59. 14"

When you submit data to the WDCA, where does it go? Website: ebas. nilu. no Note: website is painfully slow. Need to be patient. - Public data Most of the data stored in EBAS are originating from programs encouraging an unlimited and open data policy for non-commercial use. For scientific purposes, access to these data is unlimited and provided without charge. By their use you accept that an offer of co -authorship will be made through personal contact with the data providers or owners whenever substantial use is made of their data. In all cases, an acknowledgement must be made to the data providers or owners and to the project name when these data are used within a publication. Public data are available without login.

How do I get data out of the WDCA? To get to this page need to click ‘accept terms’ on start-up page. Data policy is also here

Select group to get all parameters Select date range Download!
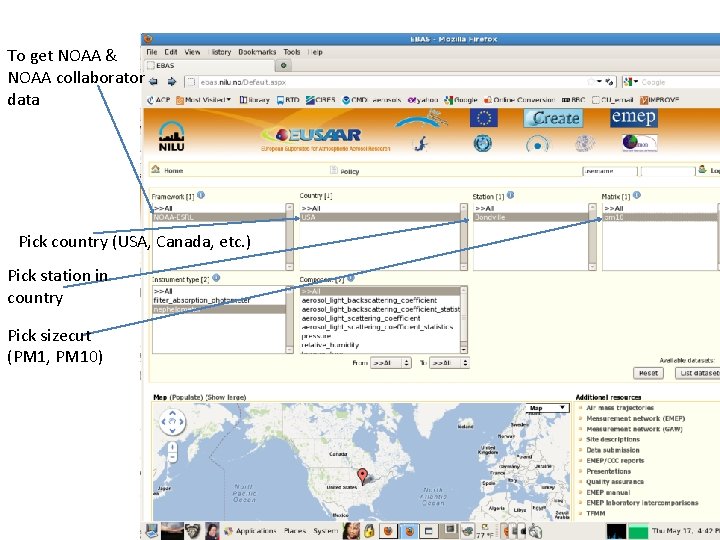
To get NOAA & NOAA collaborator data Pick country (USA, Canada, etc. ) Pick station in country Pick sizecut (PM 1, PM 10)

Info/Caveats about data downloaded from NILU/EBAS There will be one file for each year, size cut, and instrument. If it’s NOAA collaborator file, the header will be huge and descriptive. The data files will have consistent format from year to year (so long as there wasn’t an instrument change). If you download data from a non-NOAA site the header may not be as informative. Non-NOAA station files may switch format from year to year. You will need to look in the header to see what data are in what column. Example list of TSI neph files for a station: PR 0100 C. 20080101000000. 20120201192555. aerosol_light_scattering_coefficient. pm 10. 1 y. 1 h. lev 2. nas PR 0100 C. 20080101000000. 20120201192555. aerosol_light_scattering_coefficient. pm 1. 1 y. 1 h. lev 2. nas PR 0100 C. 20090101000000. 20120201194805. aerosol_light_scattering_coefficient. pm 10. 1 y. 1 h. lev 2. nas PR 0100 C. 20090101000000. 20120201194805. aerosol_light_scattering_coefficient. pm 1. 1 y. 1 h. lev 2. nas PR 0100 C. 20100101000000. 20120202190241. aerosol_light_scattering_coefficient. pm 10. 1 y. 1 h. lev 2. nas PR 0100 C. 20100101000000. 20120202190241. aerosol_light_scattering_coefficient. pm 1. 1 y. 1 h. lev 2. nas PR 0100 C. 20110101000000. 20120201160604. aerosol_light_scattering_coefficient. pm 10. 1 y. 1 h. lev 2. nas PR 0100 C. 20110101000000. 20120201160604. aerosol_light_scattering_coefficient. pm 1. 1 y. 1 h. lev 2. nas
- Slides: 29