The Contribution of Ploidy to Functional Genomic Comparisons

The Contribution of Ploidy to Functional Genomic Comparisons in Yeasts Eric Delgado Regev Group Summer Research Program in Genomics
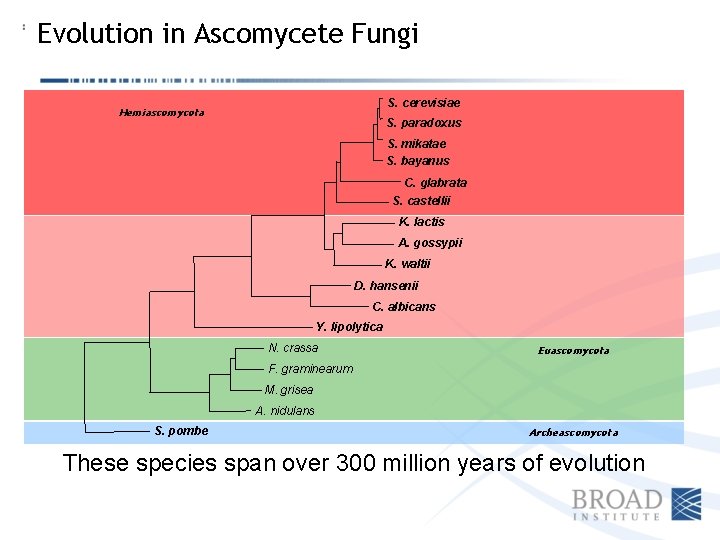
Evolution in Ascomycete Fungi S. cerevisiae Hemiascomycota S. paradoxus S. mikatae S. bayanus C. glabrata S. castellii K. lactis A. gossypii K. waltii D. hansenii C. albicans Y. lipolytica N. crassa Euascomycota F. graminearum M. grisea A. nidulans S. pombe Archeascomycota These species span over 300 million years of evolution
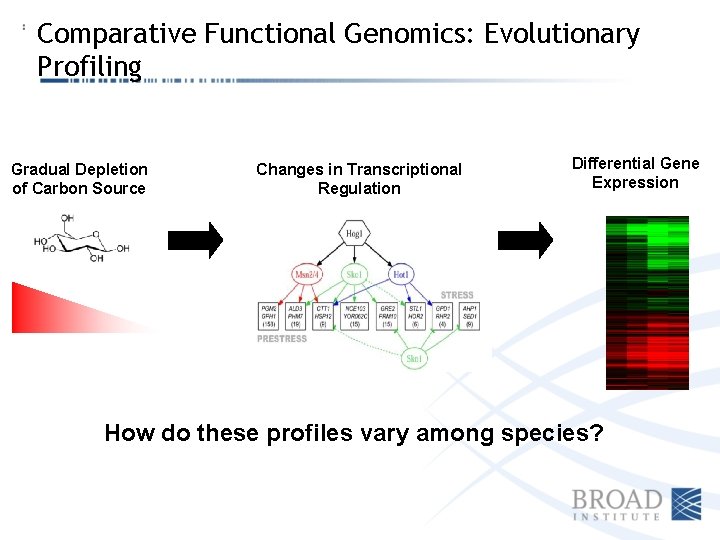
Comparative Functional Genomics: Evolutionary Profiling Gradual Depletion of Carbon Source Changes in Transcriptional Regulation Differential Gene Expression How do these profiles vary among species?
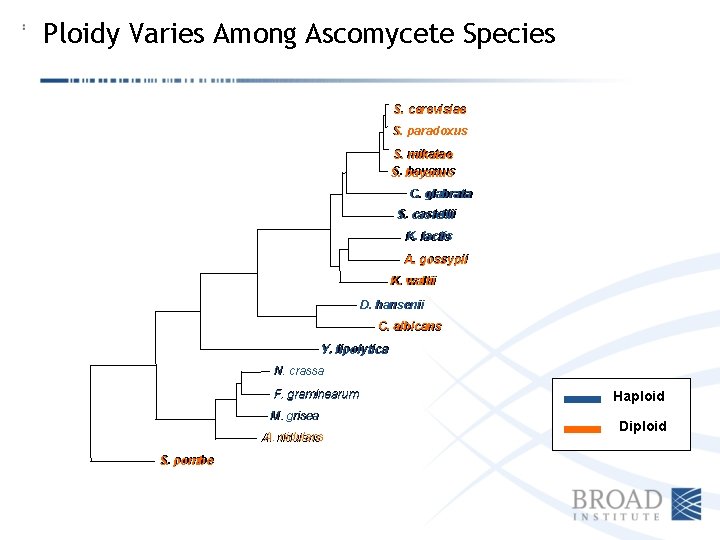
Ploidy Varies Among Ascomycete Species S. cerevisiae S. paradoxus S. mikatae S. bayanus S. C. C. glabrata S. castellii S. K. lactis K. A. gossypii K. K. waltii D. hansenii C. albicans Y. lipolytica Y. N. crassa F. graminearum M. grisea A. nidulans A. S. pombe Haploid Diploid

The Ploidy Contribution • Does Ploidy Affect Comparisons? • Species-Specific Traits or Ploidy Dependent Differences • Over a Series of Time Points and Different Species
Addressing the Ploidy Question 2 N Diploid N Haploid
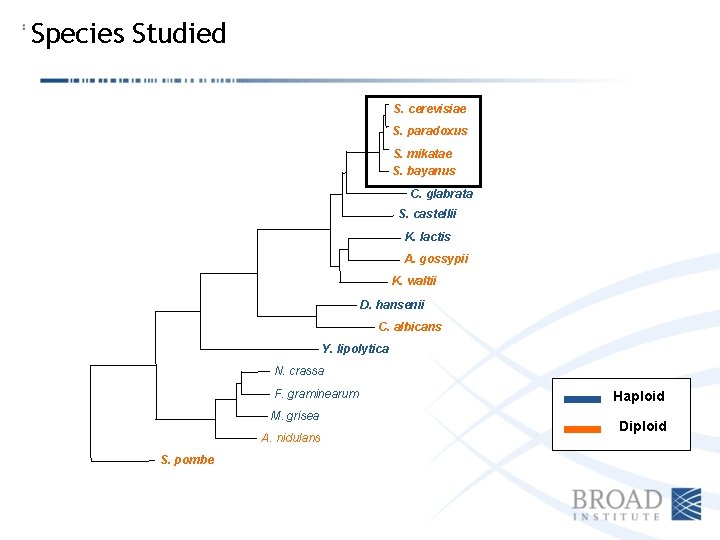
Species Studied S. cerevisiae S. paradoxus S. mikatae S. bayanus C. glabrata S. castellii K. lactis A. gossypii K. waltii D. hansenii C. albicans Y. lipolytica N. crassa F. graminearum M. grisea A. nidulans S. pombe Haploid Diploid
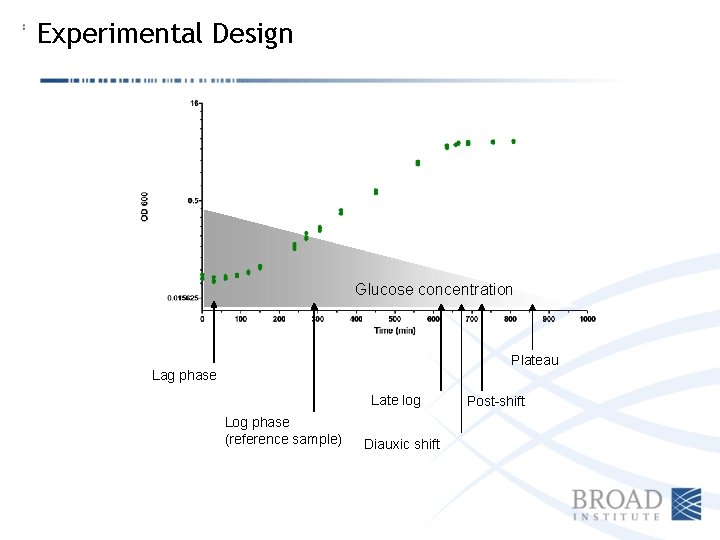
Experimental Design Glucose concentration Plateau Lag phase Late log Log phase (reference sample) Diauxic shift Post-shift
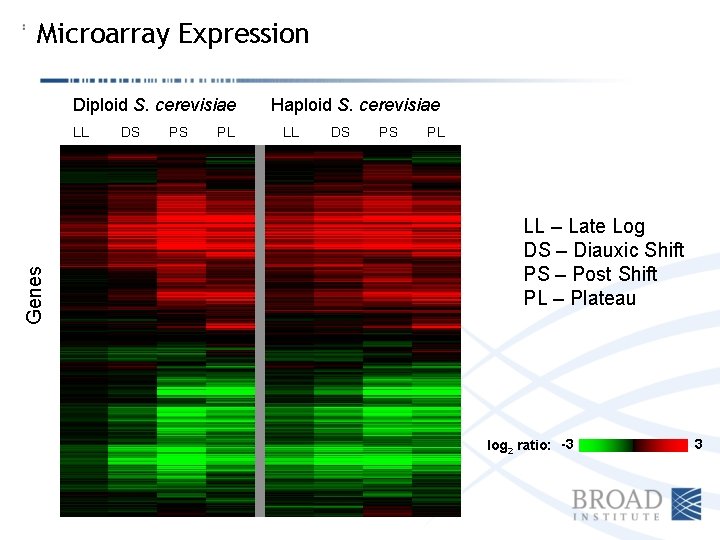
Microarray Expression Diploid S. cerevisiae Genes LL DS PS PL Haploid S. cerevisiae LL DS PS PL LL – Late Log DS – Diauxic Shift PS – Post Shift PL – Plateau log 2 ratio: -3 3
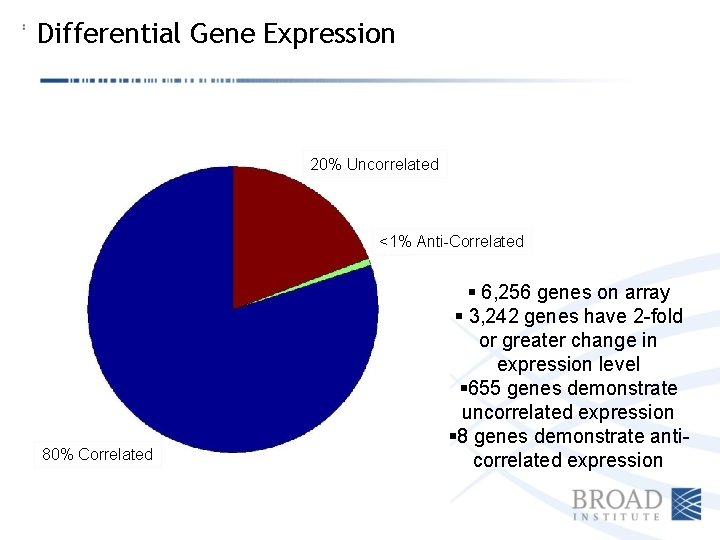
Differential Gene Expression 20% Uncorrelated <1% Anti-Correlated 80% Correlated § 6, 256 genes on array § 3, 242 genes have 2 -fold or greater change in expression level § 655 genes demonstrate uncorrelated expression § 8 genes demonstrate anticorrelated expression
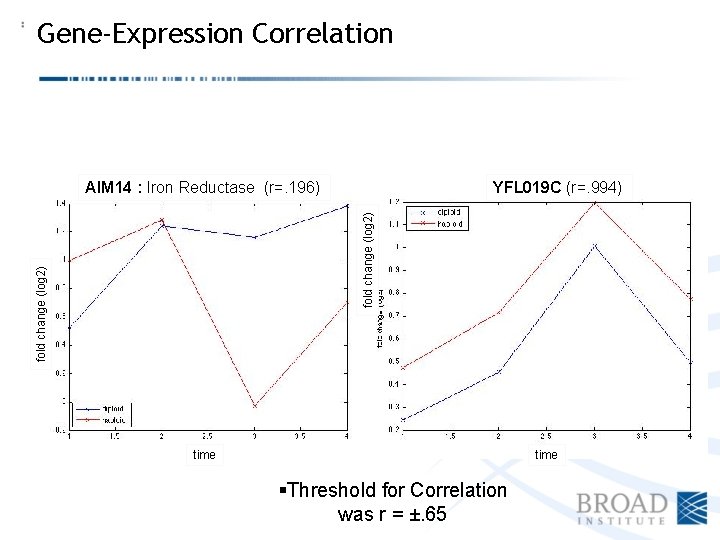
Gene-Expression Correlation YFL 019 C (r=. 994) fold change (log 2) AIM 14 : Iron Reductase (r=. 196) time §Threshold for Correlation was r = ±. 65
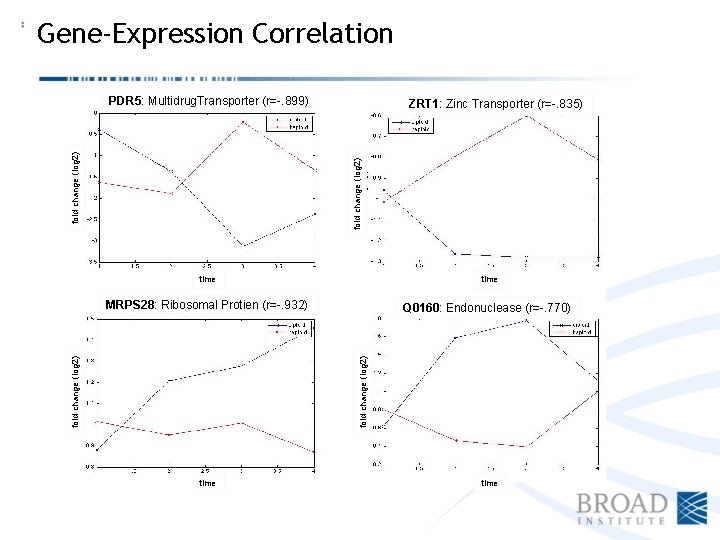
Gene-Expression Correlation ZRT 1: Zinc Transporter (r=-. 835) fold change (log 2) PDR 5: Multidrug. Transporter YKL 106 -C (r=-. 943) (r=-. 899) MRPS 28: Ribosomal Protien (r=-. 932) Q 0160: Endonuclease (r=-. 770) fold change (log 2) time

Functional Enrichment §Ribosome biogenesis was most affected §Haploid cells are smaller than diploid cells §No genes in common with the findings of Glatizsky et al.

Conclusions § Ploidy has an effect on ribosome biogenesis §These differences will be taken into account in the future §Future Work: §Work focusing on more species should be pursued §Engineering a haploid strain to become diploid would offer more insights

Acknowledgements The Regev Group Dawn Thomspon Jenna Pfifner Courtney French Mark Styczynski Michelle Chan Aviv Regev Summer Research Program In Genomics Shawna Young Lucia Vielma Maura Silverstien Bruce Birren
- Slides: 15