The Coalescent and Measurably Evolving Populations Alexei Drummond

The Coalescent and Measurably Evolving Populations Alexei Drummond Department of Computer Science University of Auckland, NZ

The Coalescent and Measurably Evolving Populations Overview 1. Introduction to the Coalescent 2. Hepatitis C in Egypt • An example using the coalescent 3. Measurably evolving populations 4. HIV-1 evolution within and among hosts • An example using MEP concepts 5. Summary + Conclusions

The Coalescent and Measurably Evolving Populations The coalescent • The coalescent is a model of the ancestral relationships of a small sample of individuals taken from a large background population. • The coalescent describes a probability distribution on ancestral genealogies (trees) given a population history. – Therefore the coalescent can convert information from ancestral genealogies into information about population history and vice versa. • The coalescent is a model of ancestral genealogies, not sequences, and its simplest form assumes neutral evolution.

The Coalescent and Measurably Evolving Populations The history of coalescent theory • • • 1930 -40 s: Genealogical arguments well known to Wright & Fisher 1964: Crow & Kimura: Infinite Allele Model 1966: (Hubby & Lewontin) & (Harris) make first surveys of population allele variation by protein electrophoresis 1968: Motoo Kimura proposes neutral explanation of molecular evolution & population variation. So do King & Jukes 1971: Kimura & Ohta proposes infinite sites model. 1975: Watterson makes explicit use of “The Coalescent” 1982: Kingman introduces “The Coalescent”. 1983: Hudson introduces “The Coalescent with Recombination” 1983: Kreitman publishes first major population sequences. 1987: Cann et al. traces human origin and migrations with mitochondrial DNA.

The Coalescent and Measurably Evolving Populations The history of coalescent theory • • 1988: Hughes & Nei: Genes with positive Darwinian Selection. 1989 -90: Kaplan, Hudson, Takahata and others: Selection regimes with coalescent structure (MHC, Incompatibility alleles). 1991: Mac. Donald & Kreitman: Data with surplus of replacement interspecific substitutions. 1994 -95: Griffiths-Tavaré + Kuhner-Yamoto-Felsenstein introduces sampling techniques to estimate parameters in population models. 1997 -98: Krone-Neuhauser introduces Ancestral Selection Graph 1999: Wiuf & Donnelly uses coalescent theory to estimate age of disease allele 2000: Wiuf et al. introduces gene conversion into coalescent. 2000 -: A flood of SNP data & haplotypes are on their way.
The Coalescent and Measurably Evolving Populations Population processes COALESCENT THEORY Genealogy
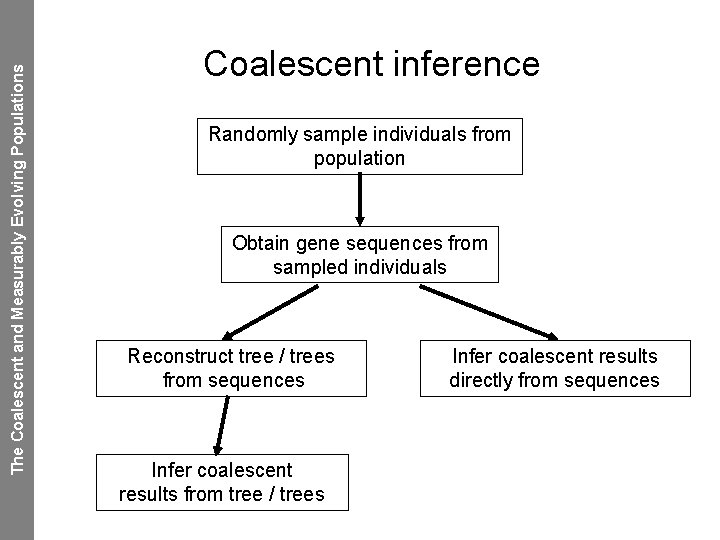
The Coalescent and Measurably Evolving Populations Coalescent inference Randomly sample individuals from population Obtain gene sequences from sampled individuals Reconstruct tree / trees from sequences Infer coalescent results from tree / trees Infer coalescent results directly from sequences

The Coalescent and Measurably Evolving Populations Demographic History • Change in population size through time • Applications include – – Estimating history of human populations Conservation biology Reconstructing infectious disease epidemics Investigating viral dynamics within hosts
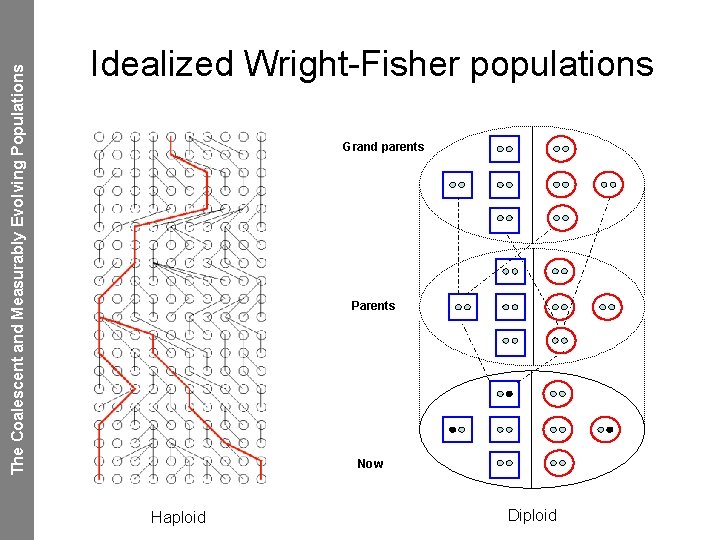
The Coalescent and Measurably Evolving Populations Idealized Wright-Fisher populations Grand parents Parents Now Haploid Diploid

The Coalescent and Measurably Evolving Populations Random mating in an ideal population • A constant population size of N individuals • Each individual in the new generation “chooses” its parent from the previous generation at random
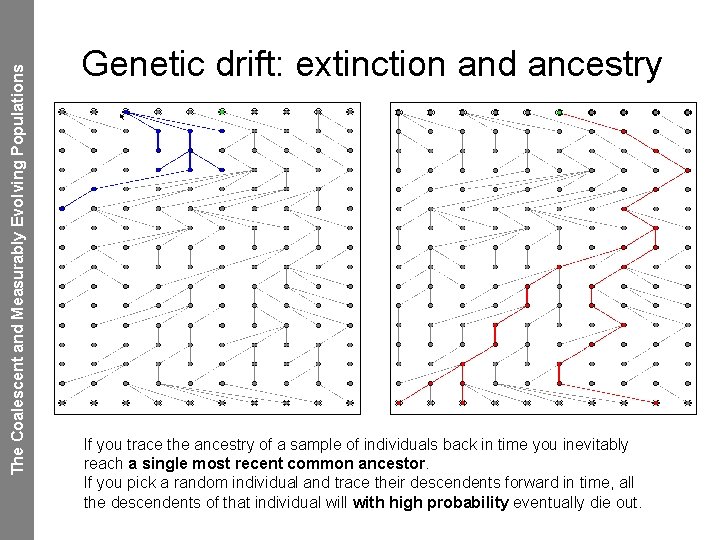
The Coalescent and Measurably Evolving Populations Genetic drift: extinction and ancestry If you trace the ancestry of a sample of individuals back in time you inevitably reach a single most recent common ancestor. If you pick a random individual and trace their descendents forward in time, all the descendents of that individual will with high probability eventually die out.
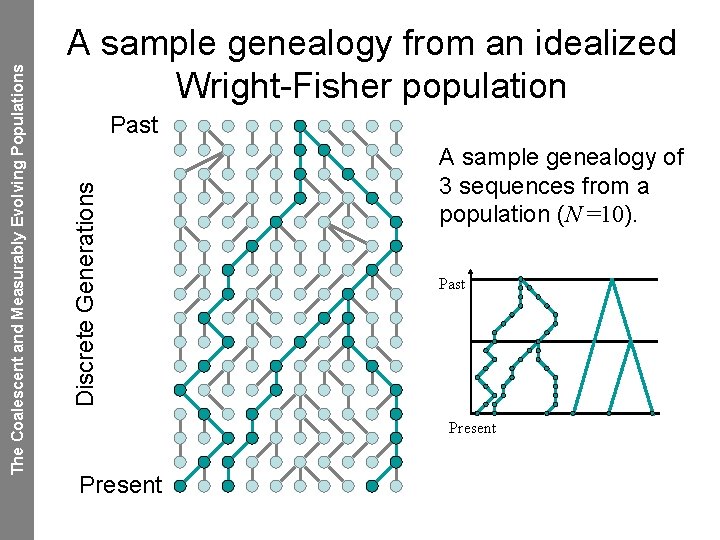
Past Discrete Generations The Coalescent and Measurably Evolving Populations A sample genealogy from an idealized Wright-Fisher population A sample genealogy of 3 sequences from a population (N =10). Past Present

The Coalescent and Measurably Evolving Populations The coalescent: distributions and expectations on a sample genealogy Past Present
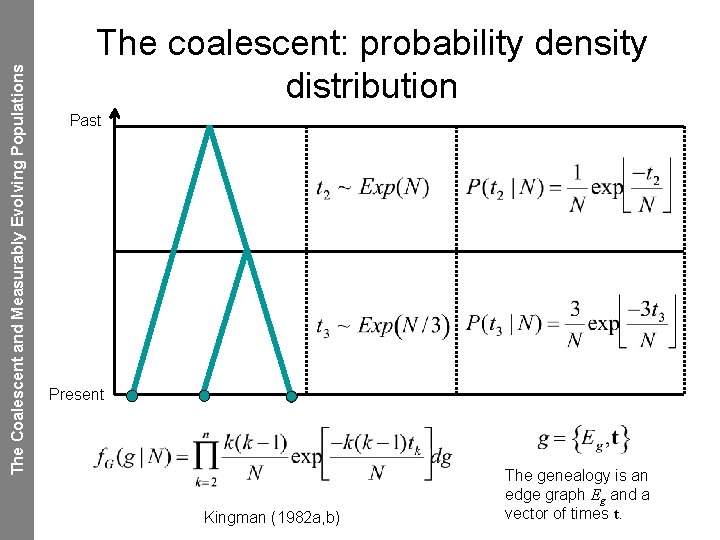
The Coalescent and Measurably Evolving Populations The coalescent: probability density distribution Past Present Kingman (1982 a, b) The genealogy is an edge graph Eg and a vector of times t.
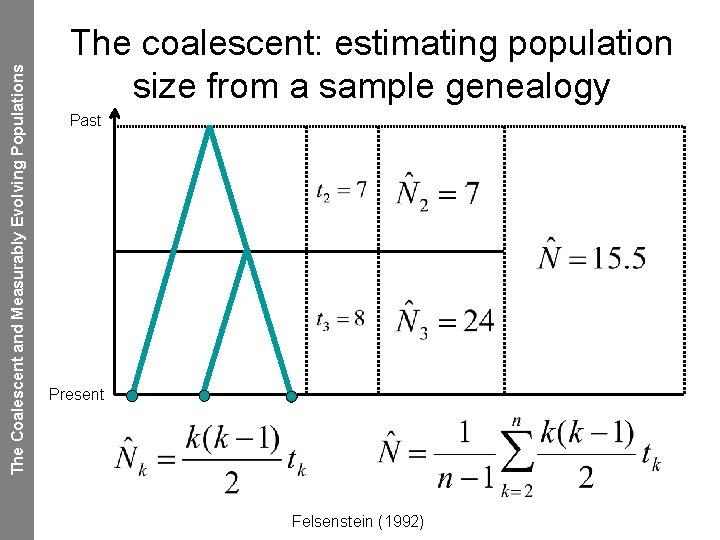
The Coalescent and Measurably Evolving Populations The coalescent: estimating population size from a sample genealogy Past Present Felsenstein (1992)
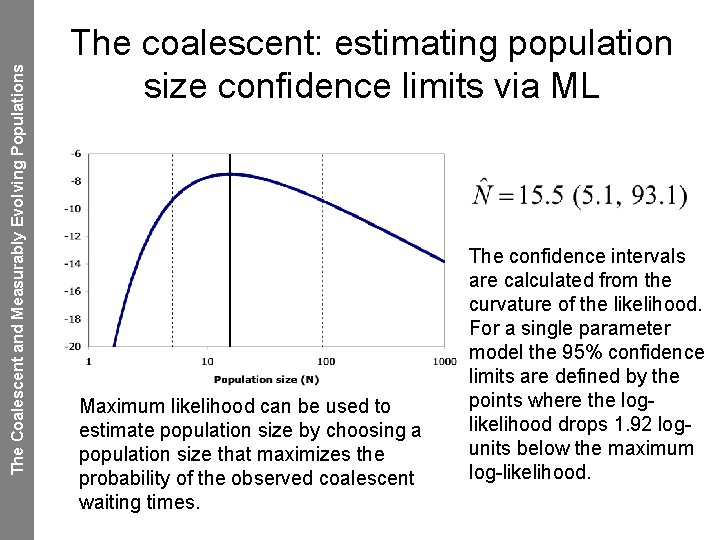
The Coalescent and Measurably Evolving Populations The coalescent: estimating population size confidence limits via ML Maximum likelihood can be used to estimate population size by choosing a population size that maximizes the probability of the observed coalescent waiting times. The confidence intervals are calculated from the curvature of the likelihood. For a single parameter model the 95% confidence limits are defined by the points where the loglikelihood drops 1. 92 logunits below the maximum log-likelihood.
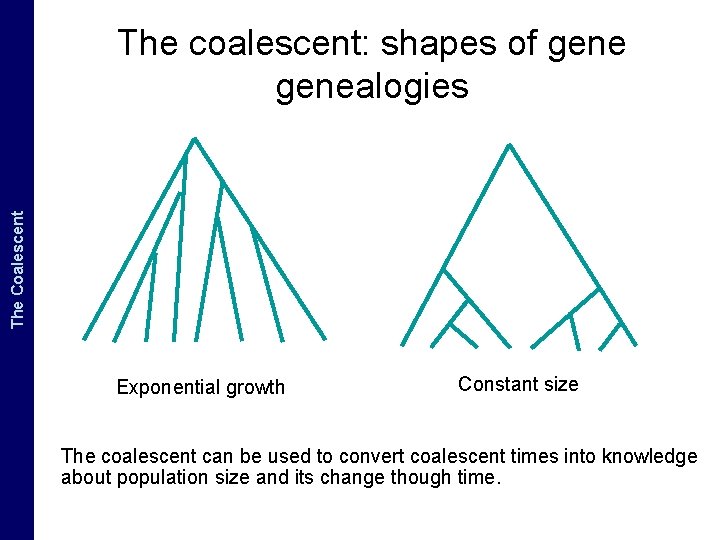
The Coalescent and. The Measurably Evolving Populations Coalescent The coalescent: shapes of genealogies Exponential growth Constant size The coalescent can be used to convert coalescent times into knowledge about population size and its change though time.
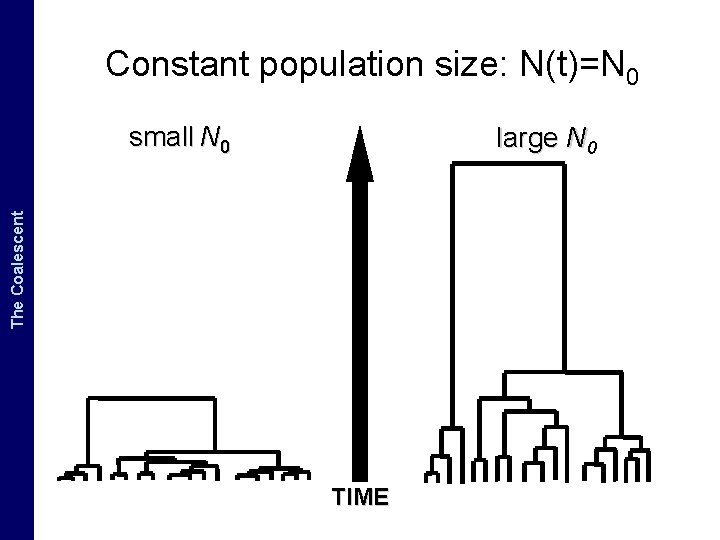
The Coalescent and. The Measurably Evolving Populations Coalescent Constant population size: N(t)=N 0 small N 0 large N 0 TIME
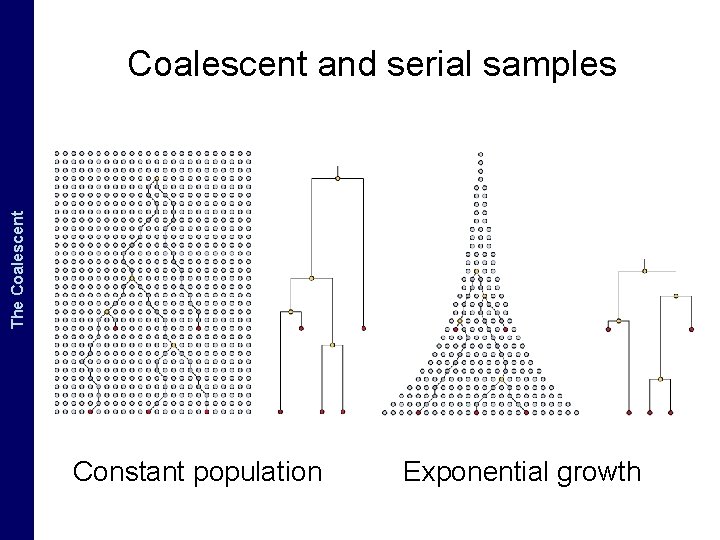
The Coalescent and. The Measurably Evolving Populations Coalescent and serial samples Constant population Exponential growth
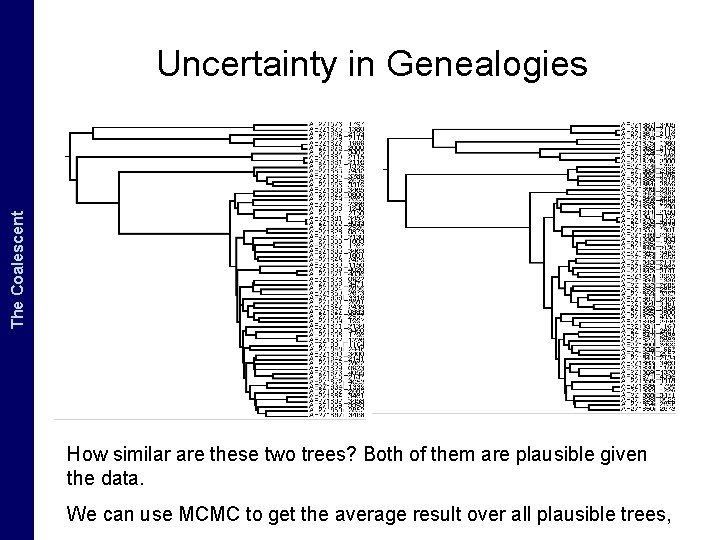
The Coalescent and. The Measurably Evolving Populations Coalescent Uncertainty in Genealogies How similar are these two trees? Both of them are plausible given the data. We can use MCMC to get the average result over all plausible trees,

The Coalescent and. The Measurably Evolving Populations Coalescent Summary • The coalescent provides a theory of how population size is related to the distribution of coalescent events in a tree. • Big populations have old trees • Exponentially growing populations have star-like trees • Given a genealogy the most likely population size can be estimated. • MCMC can be used to get a distribution of trees from which a distribution of population sizes can be estimated.

The Coalescent and Measurably MCMC Evolving Populations Markov chain Monte Carlo (MCMC) • Imagine you would like to estimate two parameters ( , ) from some data (D). • You want to find values of and that have high probability given the data: p( , |D) • Say you have a likelihood function of the form: Pr{D| , } • Bayes rule tells us that: – p( , |D) = Pr{D| , }p( , ) / Pr{D} – So that p( , |D) Pr{D| , }p( , )

The Coalescent and Measurably MCMC Evolving Populations Markov chain Monte Carlo (MCMC) • p( , |D) is called the posterior probability (density) of , given D • In an ideal world we want to know the posterior density for all possible values of , . • Then we could pick a “credible region” in two dimensions that contained values of , that account for the majority of the posterior probability mass. • This credible region would serve as an estimate that includes incorporates our uncertainty and this credible set could be used to address hypotheses like: is greater than x. • In reality we have to make due with a “sample” of the posterior - so that we evaluate p( , |D) for a finite number (say 10, 000) pairs of , . • So which pairs should we choose?

The Coalescent and Measurably MCMC Evolving Populations Markov chain Monte Carlo (MCMC) • Lets construct a random walk in 2 -dimensional space • In each step of the random walk we propose to make an (unbiased) small jump from our current position ( , ) to a new position ( ’, ’) • If p( ’, ’|D) > p( , |D) then we make the proposed jump • However, if p( ’, ’|D) < p( , |D), then we make the proposed jump with probability = p( ’, ’|D) / p( , |D), otherwise we stay where we are. • It can be shown (trust me!) that if you proceed in this fashion for an infinite time then the equilibrium distribution of this random walk will be p( , |D)! • That is, the random walk will visit a particular region [ 0, 1] x [ 0, 1] of the state space this often:
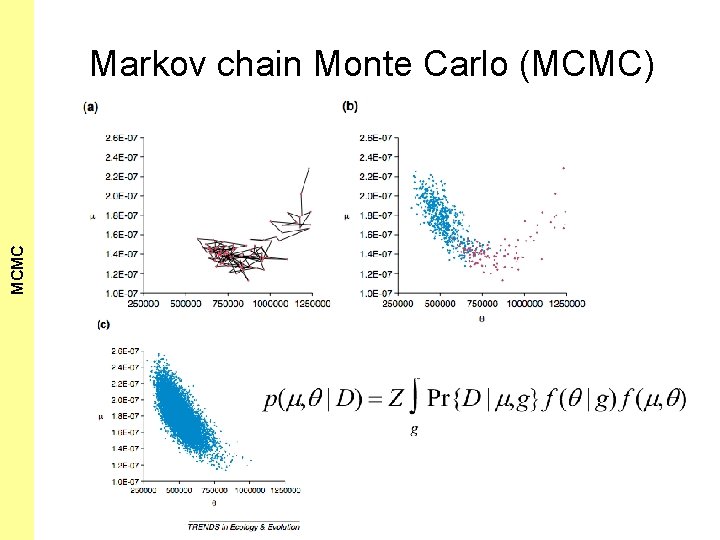
The Coalescent and Measurably MCMC Evolving Populations Markov chain Monte Carlo (MCMC)
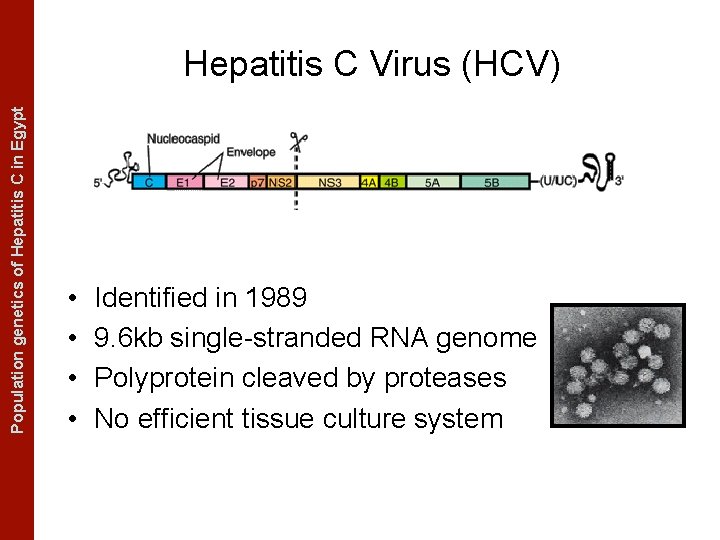
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Hepatitis C Virus (HCV) • • Identified in 1989 9. 6 kb single-stranded RNA genome Polyprotein cleaved by proteases No efficient tissue culture system

The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt How important is HCV? • • 170 m+ infected ~80% infections are chronic Liver cirrhosis & cancer risk 10, 000 deaths per year in USA • No protective immunity?

The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt HCV Transmission Percutaneous exposure to infected blood • Blood transfusion / blood products • Injecting & nasal drug use • Sexual & vertical transmission • Unsafe injections • Unidentified routes
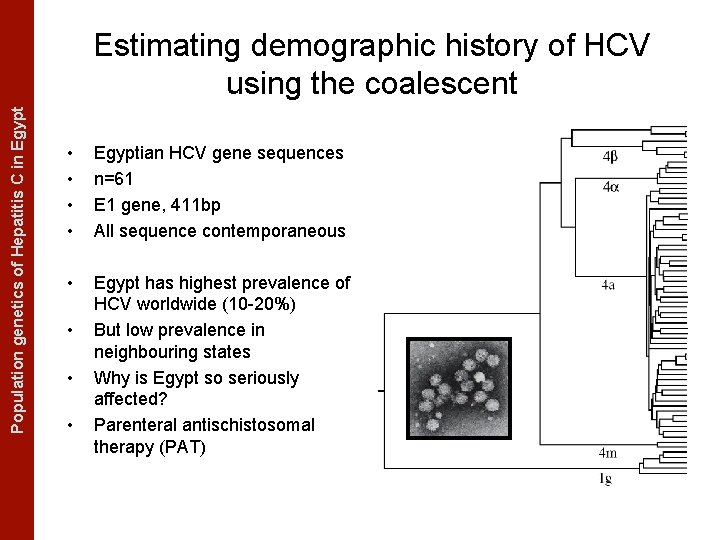
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Estimating demographic history of HCV using the coalescent • • Egyptian HCV gene sequences n=61 E 1 gene, 411 bp All sequence contemporaneous • Egypt has highest prevalence of HCV worldwide (10 -20%) But low prevalence in neighbouring states Why is Egypt so seriously affected? Parenteral antischistosomal therapy (PAT) • • •
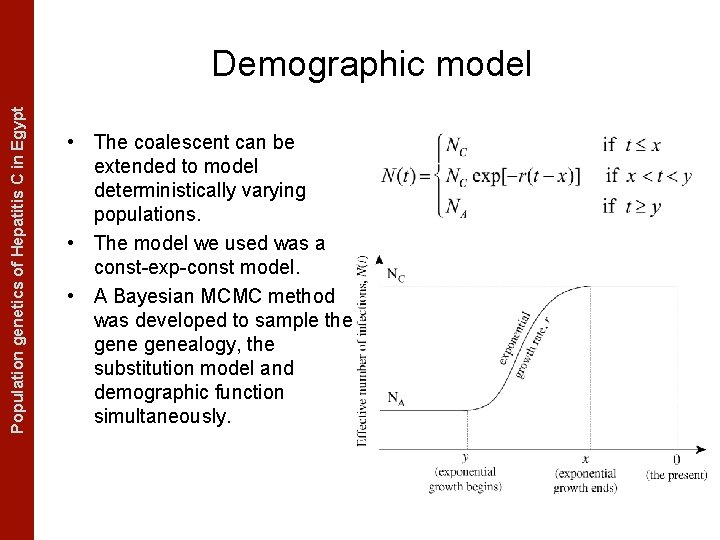
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Demographic model • The coalescent can be extended to model deterministically varying populations. • The model we used was a const-exp-const model. • A Bayesian MCMC method was developed to sample the genealogy, the substitution model and demographic function simultaneously.
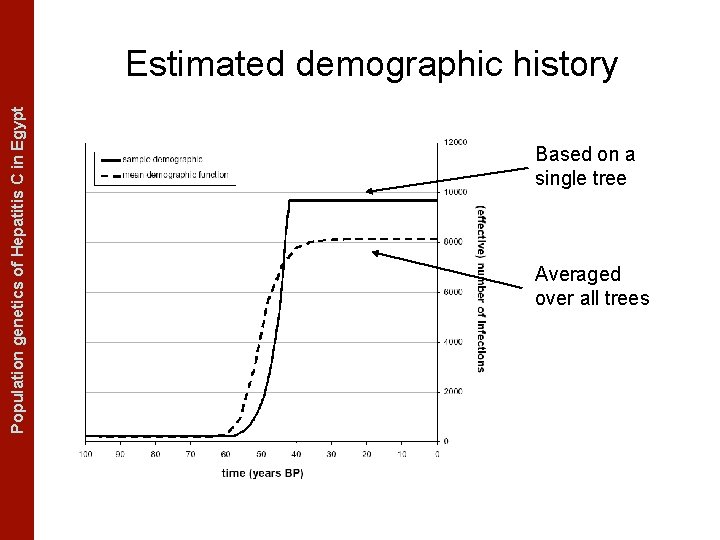
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Estimated demographic history Based on a single tree Averaged over all trees

The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Parameter estimates
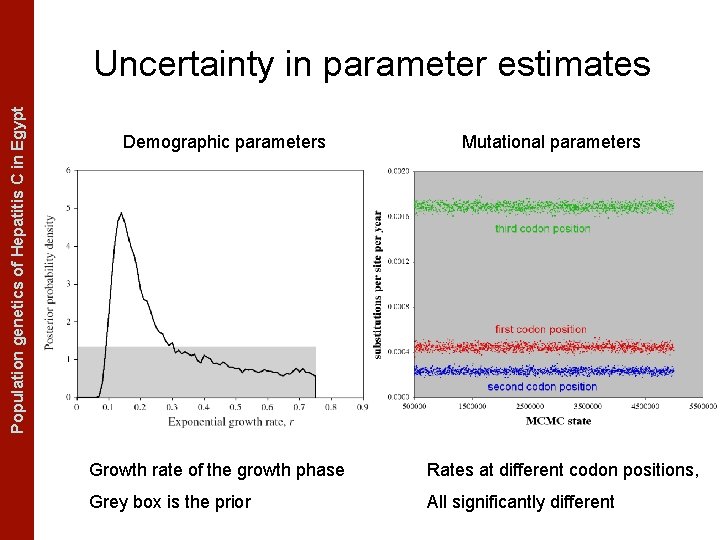
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Uncertainty in parameter estimates Demographic parameters Mutational parameters Growth rate of the growth phase Rates at different codon positions, Grey box is the prior All significantly different
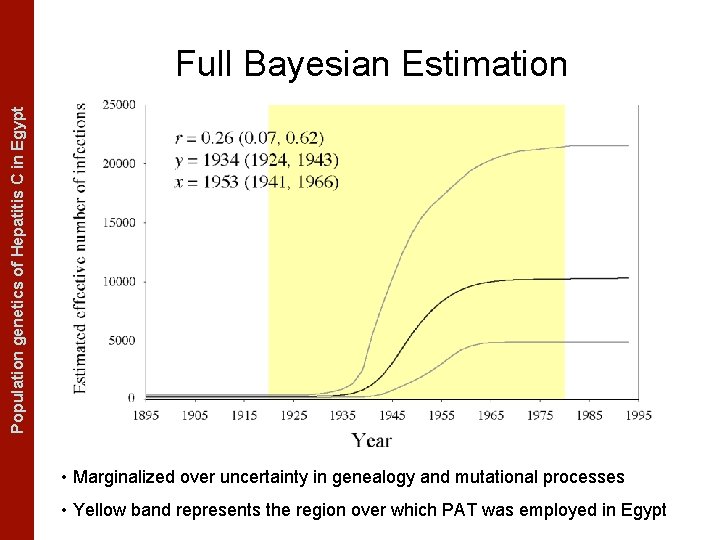
The Coalescent Measurably Evolving Populationand genetics of Hepatitis C in. Populations Egypt Full Bayesian Estimation • Marginalized over uncertainty in genealogy and mutational processes • Yellow band represents the region over which PAT was employed in Egypt
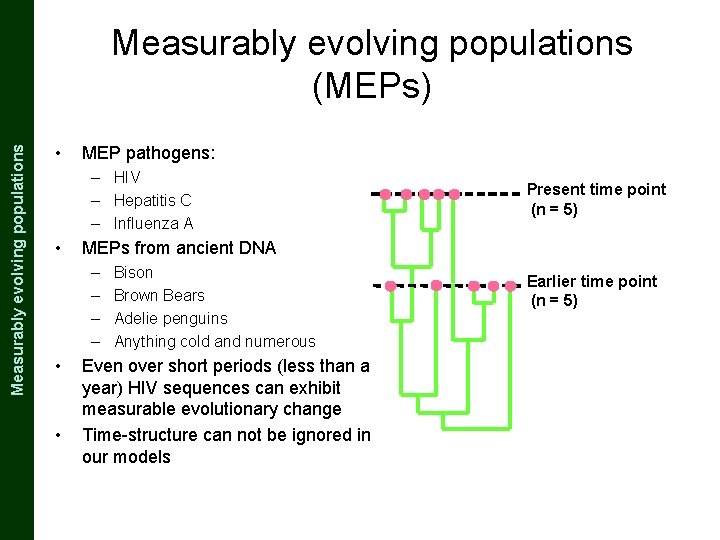
The Coalescent and Measurably Evolving Populations Measurably evolving populations (MEPs) • MEP pathogens: – HIV – Hepatitis C – Influenza A • MEPs from ancient DNA – – • • Present time point (n = 5) Bison Brown Bears Adelie penguins Anything cold and numerous Even over short periods (less than a year) HIV sequences can exhibit measurable evolutionary change Time-structure can not be ignored in our models Earlier time point (n = 5)
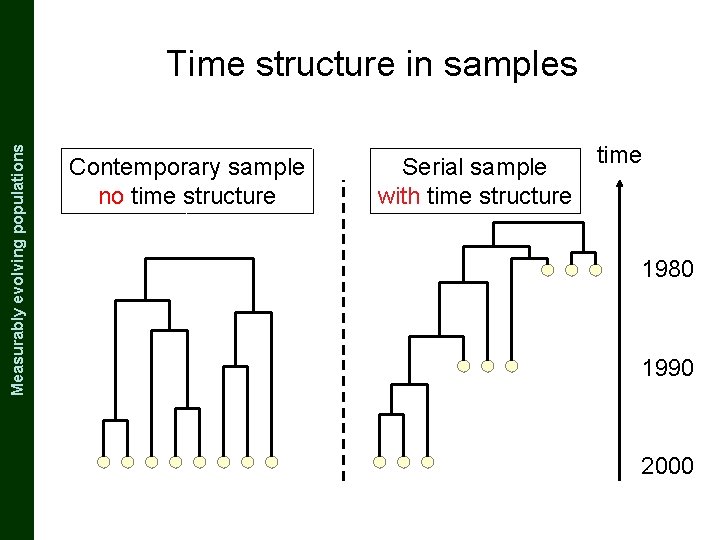
The Coalescent and Measurably Evolving Populations Measurably evolving populations Time structure in samples Contemporary sample no time structure Serial sample with time structure time 1980 1990 2000
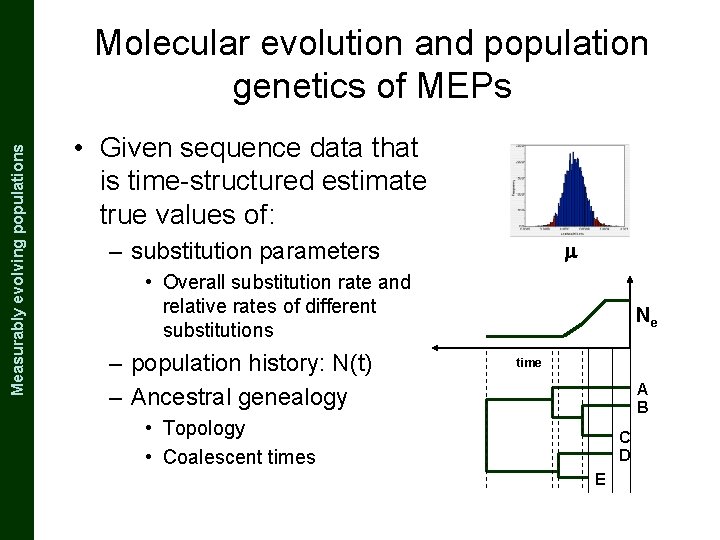
The Coalescent and Measurably Evolving Populations Measurably evolving populations Molecular evolution and population genetics of MEPs • Given sequence data that is time-structured estimate true values of: m – substitution parameters • Overall substitution rate and relative rates of different substitutions – population history: N(t) – Ancestral genealogy Ne time A B • Topology • Coalescent times C D E

The Coalescent and Measurably Evolving Populations Molecular evolutionary model: Felsenstein’s likelihood (1981) AA b 1 GA b 2 b 4 AC b 3 b 5 GC The probability of the sequence alignment, can be efficiently calculated given a tree and branch lengths (T), and a probabilistic model of mutation represented by an instantaneous rate matrix (Q). In phylogenetics, branch lengths are usually unconstrained.
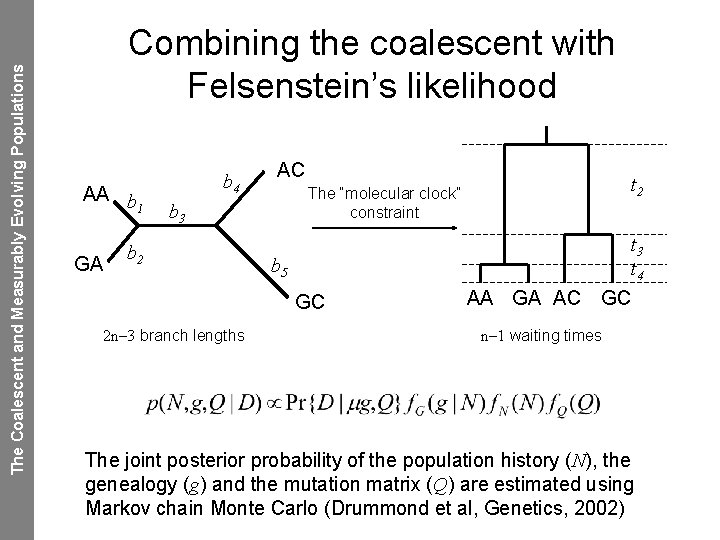
The Coalescent and Measurably Evolving Populations Combining the coalescent with Felsenstein’s likelihood AA b 1 GA b 4 AC b 3 b 2 t 3 t 4 b 5 GC 2 n– 3 branch lengths t 2 The “molecular clock” constraint AA GA AC GC n– 1 waiting times The joint posterior probability of the population history (N), the genealogy (g) and the mutation matrix (Q) are estimated using Markov chain Monte Carlo (Drummond et al, Genetics, 2002)
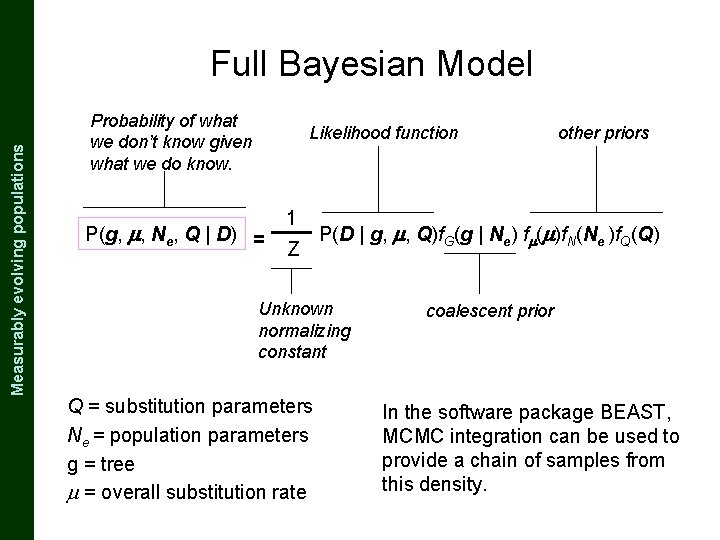
The Coalescent and Measurably Evolving Populations Measurably evolving populations Full Bayesian Model Probability of what we don’t know given what we do know. Likelihood function P(g, , Ne, Q | D) = 1 Z P(D | g, , Q)f. G(g | Ne) f ( )f. N(Ne )f. Q(Q) Unknown normalizing constant Q = substitution parameters Ne = population parameters g = tree = overall substitution rate other priors coalescent prior In the software package BEAST, MCMC integration can be used to provide a chain of samples from this density.
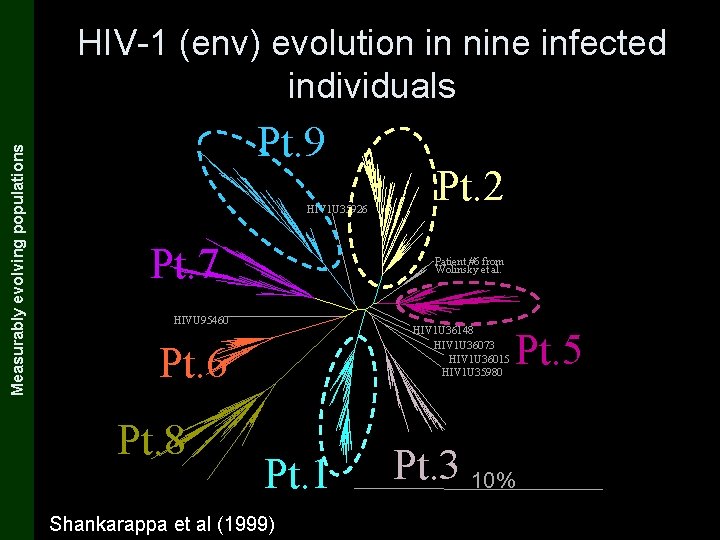
The Coalescent and Measurably Evolving Populations Measurably evolving populations HIV-1 (env) evolution in nine infected individuals Pt. 9 HIV 1 U 35926 Pt. 7 Patient #6 from Wolinsky et al. HIVU 95460 HIV 1 U 36148 HIV 1 U 36073 HIV 1 U 36015 HIV 1 U 35980 Pt. 6 Pt. 8 Pt. 2 Pt. 1 Shankarappa et al (1999) Pt. 3 10% Pt. 5
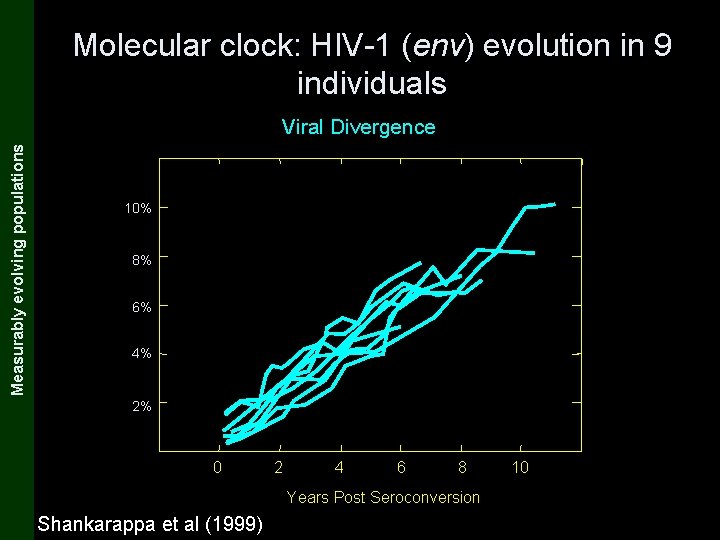
The Coalescent and Measurably Evolving Populations Measurably evolving populations Molecular clock: HIV-1 (env) evolution in 9 individuals Viral Divergence 10% 8% 6% 4% 2% 0 2 4 6 8 Years Post Seroconversion Shankarappa et al (1999) 10

The Coalescent and Measurably Evolving Populations Measurably evolving populations MEP Summary • Most RNA viruses, including HCV and HIV are measurably evolving • Most vertebrate populations that have well-preserved recent fossil records are MEPs. • If sequence data comes from different times the timestructure can’t be ignored • Time structure permits the direct estimation of: – – substitution rate Concerted changes in substitution rate coalescent times in calendar units Demographic function N(t) in calendar units

The Coalescent and Measurably Evolving Populations Intermission My brain is fried!
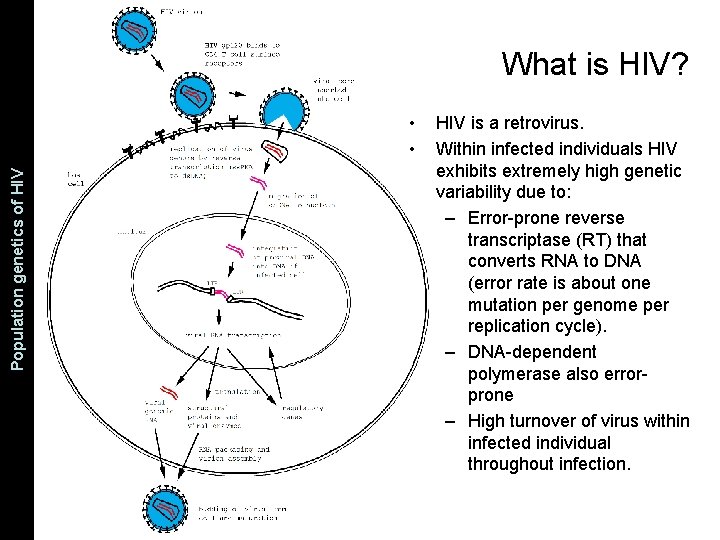
The Coalescent and Measurably Population genetics. Evolving of HIV Populations What is HIV? • • HIV is a retrovirus. Within infected individuals HIV exhibits extremely high genetic variability due to: – Error-prone reverse transcriptase (RT) that converts RNA to DNA (error rate is about one mutation per genome per replication cycle). – DNA-dependent polymerase also errorprone – High turnover of virus within infected individual throughout infection.
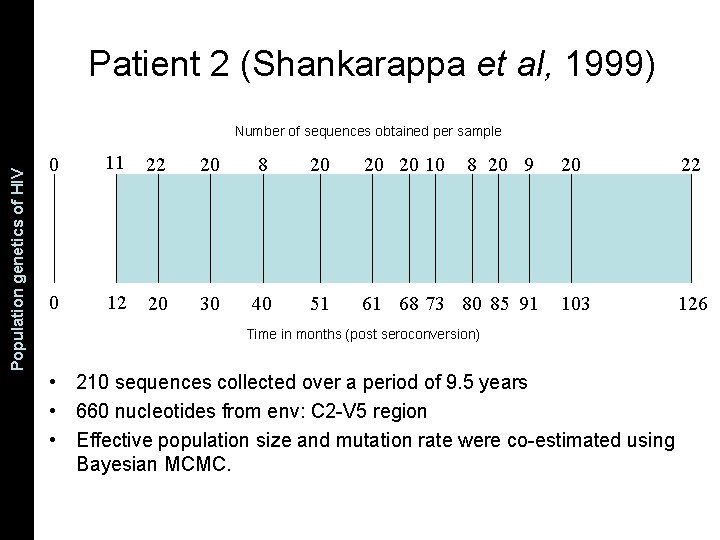
The Coalescent and Measurably Population genetics. Evolving of HIV Populations Patient 2 (Shankarappa et al, 1999) Number of sequences obtained per sample 0 11 22 20 8 20 20 20 10 0 12 20 30 40 51 61 68 73 80 85 91 8 20 9 20 22 103 126 Time in months (post seroconversion) • 210 sequences collected over a period of 9. 5 years • 660 nucleotides from env: C 2 -V 5 region • Effective population size and mutation rate were co-estimated using Bayesian MCMC.
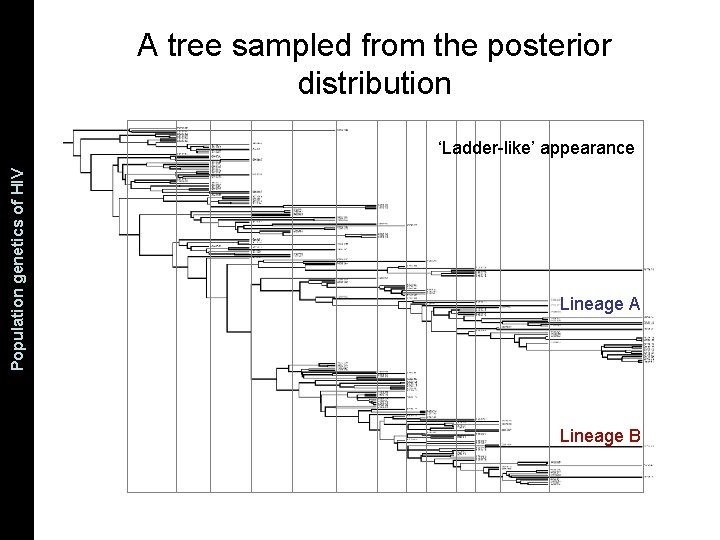
The Coalescent and Measurably Population genetics. Evolving of HIV Populations A tree sampled from the posterior distribution ‘Ladder-like’ appearance Lineage A Lineage B
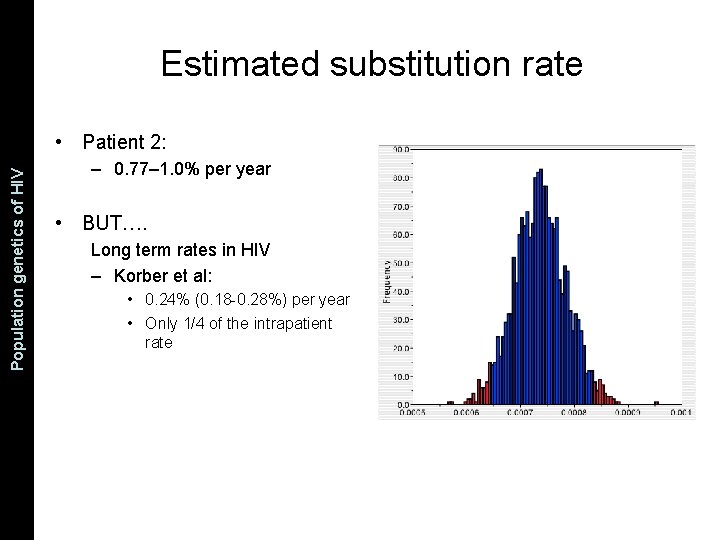
The Coalescent and Measurably Population genetics. Evolving of HIV Populations Estimated substitution rate • Patient 2: – 0. 77– 1. 0% per year • BUT…. Long term rates in HIV – Korber et al: • 0. 24% (0. 18 -0. 28%) per year • Only 1/4 of the intrapatient rate
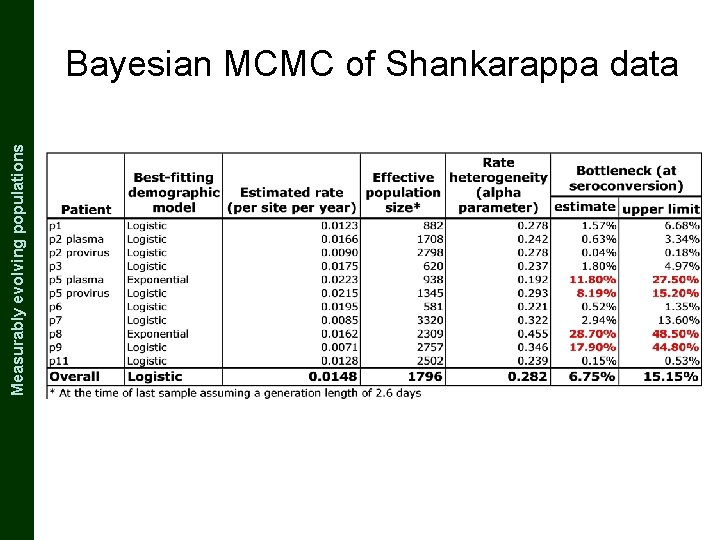
The Coalescent and Measurably Evolving Populations Measurably evolving populations Bayesian MCMC of Shankarappa data
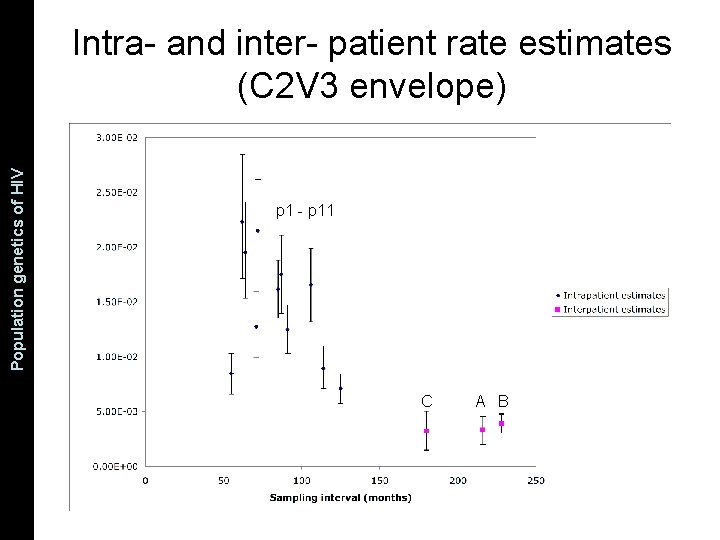
The Coalescent and Measurably Population genetics. Evolving of HIV Populations Intra- and inter- patient rate estimates (C 2 V 3 envelope) p 1 - p 11 C A B

The Coalescent and Measurably Population genetics. Evolving of HIV Populations Summary: HIV intra-patient evolution • HIV evolutionary rates appear to be faster intrapatient then across pandemic – Different selection pressure at transmission? – Transmitted viruses undergoing less rounds of replication? – Latent viruses? – Reversion of escape mutants? • Effective population size is changing over time (bottleneck in envelope at least)

The Coalescent and Measurably Evolving Populations Goodness-of-fit tests But how good is our best model? • We can use standard statistical model-choice criteria to choose between different models of substitution and demography, but are any of the models we consider any good at all? • One way to look at this is ask the following question: – Does our real data look anything like what we would expect data from our model to look like? • So what aspect of the data should we look at? • And what should we expect?
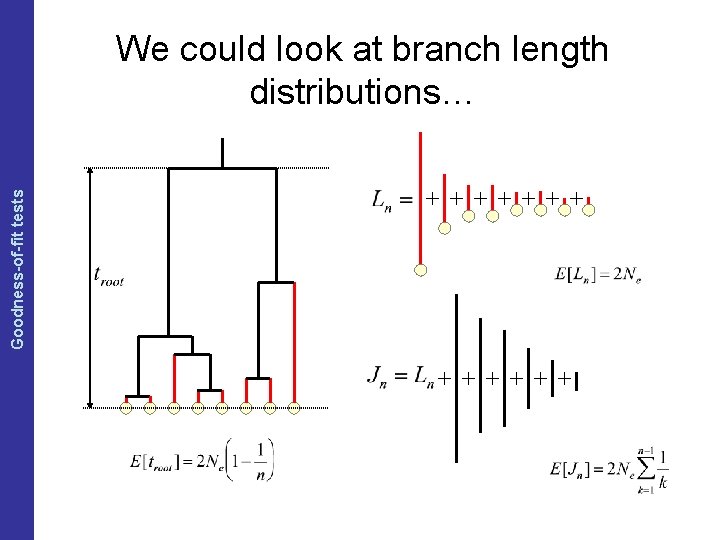
The Coalescent and Measurably Evolving Populations Goodness-of-fit tests We could look at branch length distributions…
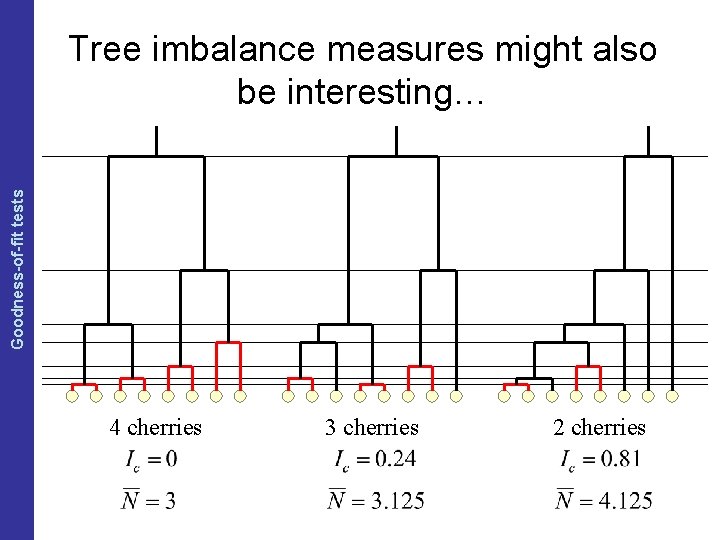
The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Tree imbalance measures might also be interesting… 4 cherries 3 cherries 2 cherries

The Coalescent and Measurably Evolving Populations Posterior predictive simulation • A method of testing the goodness-of-fit of a Bayesian model. 1. Run a Bayesian MCMC analysis on the data 2. Calculate the value of your favourite summary statistic, T(. ) from the data, D 3. For each state in the chain 1. Simulate a synthetic dataset, Di, using the parameter values of state i. 2. Calculate T(Di) from the simulated data set. 4. Compare the T(D) value with predictive distribution of T(Di)

The Coalescent and Measurably Evolving Populations Posterior predictive simulation So we need some summary statistics • Summary statistics that can be measured directly from an genealogy: from sequence alignment: – Mean pairwise distance ( ) – Tajima’s D – Fu & Li’s D – Number of segregating sites (S) – … – Genealogical mean pairwise distance ( ) – Genealogical Tajima’s D – Genealogical Fu & Li’s D – Tree-imbalance statistics – Age of the root – Length of the tree

The Coalescent and Measurably Evolving Populations Posterior predictive simulation (2) • Testing the goodness-of-fit of the neutral coalescent model under variable demographic functions. 1. Run a Bayesian MCMC analysis on the data 2. For each state in the chain 1. Simulate a coalescent genealogy (Gi. S) using the population parameter values of state i. 2. Calculate T(Gi. S) from the ith simulated genealogy 3. Calculate T(Gi. P) from the ith posterior genealogy 3. Calculate the predictive probability by comparing the posterior distribution of T(. ) with predictive distribution of T(. ):

The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Human influenza A (HA gene) trees State 5 m State 10 m Posterior genealogy Predictive simulations

The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Human influenza A trees: Genealogical Fu & Li’s D statistic
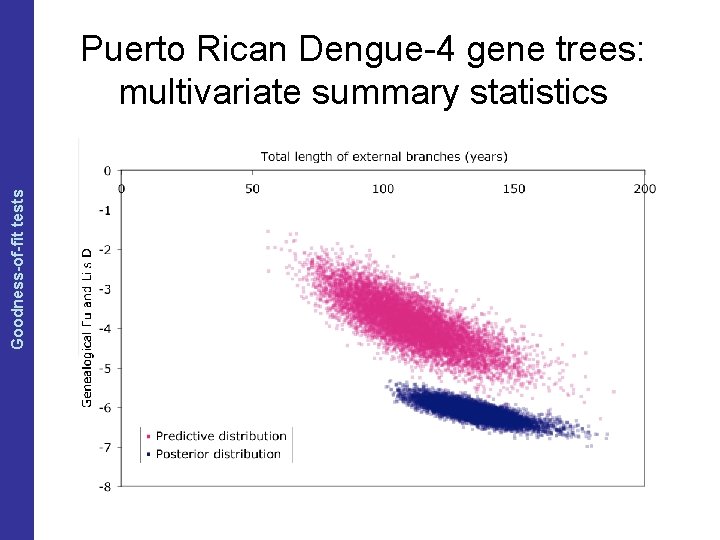
The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Puerto Rican Dengue-4 gene trees: multivariate summary statistics
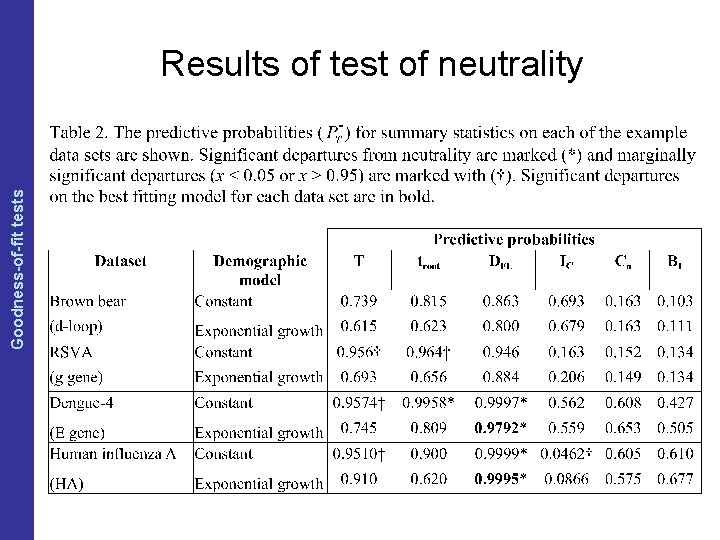
The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Results of test of neutrality
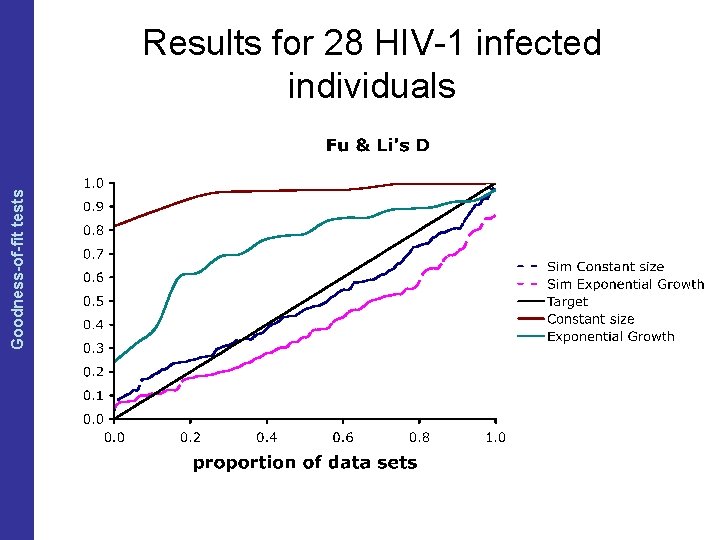
The Coalescent and Measurably Evolving Populations Goodness-of-fit tests Results for 28 HIV-1 infected individuals
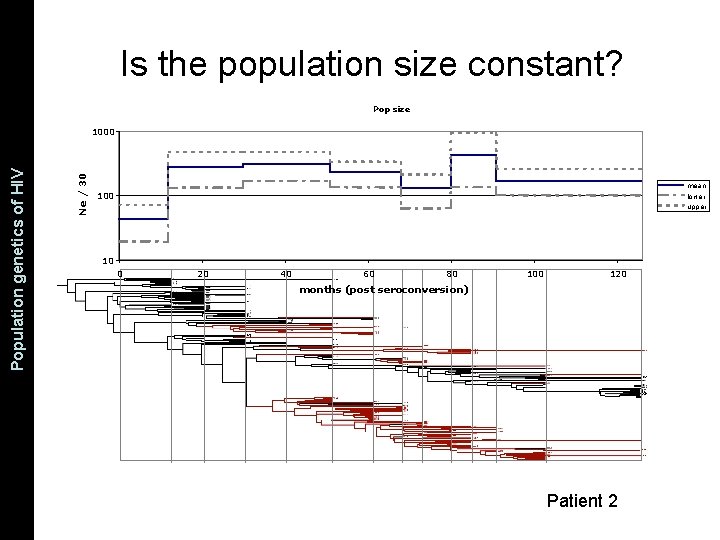
Pop size 1000 Ne / 30 The Coalescent and Measurably Population genetics. Evolving of HIV Populations Is the population size constant? mean lower upper 100 10 0 20 40 60 80 100 120 months (post seroconversion) Patient 2
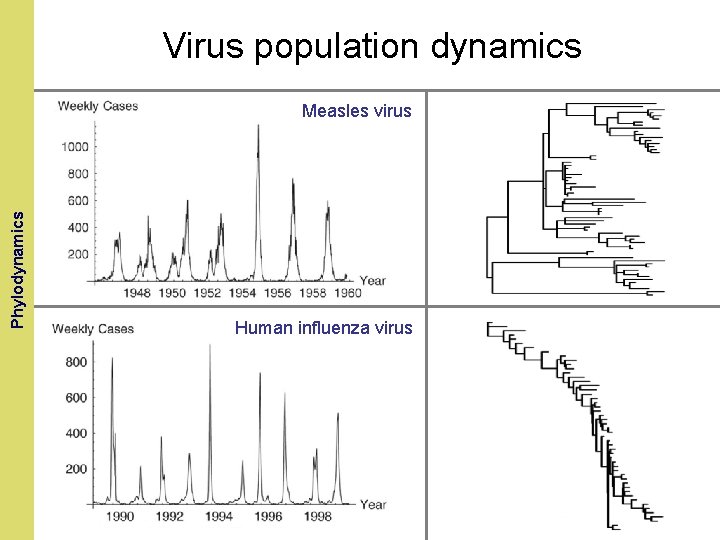
The Coalescent and. Phylodynamics Measurably Evolving Populations Virus population dynamics Measles virus Human influenza virus
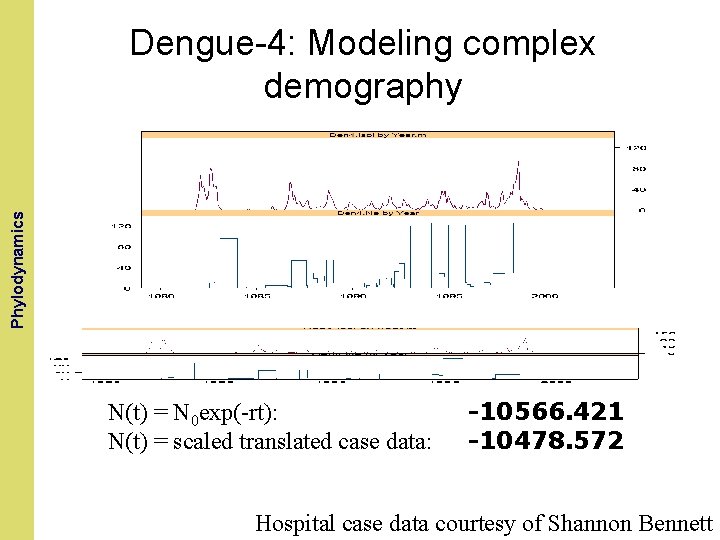
The Coalescent and. Phylodynamics Measurably Evolving Populations Dengue-4: Modeling complex demography N(t) = N 0 exp(-rt): N(t) = scaled translated case data: -10566. 421 -10478. 572 Hospital case data courtesy of Shannon Bennett
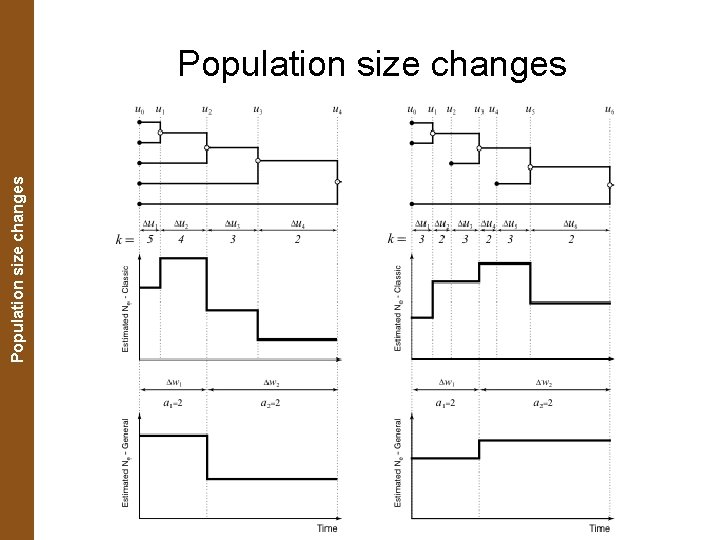
The Coalescent. Population and Measurably Evolving Populations size changes Population size changes
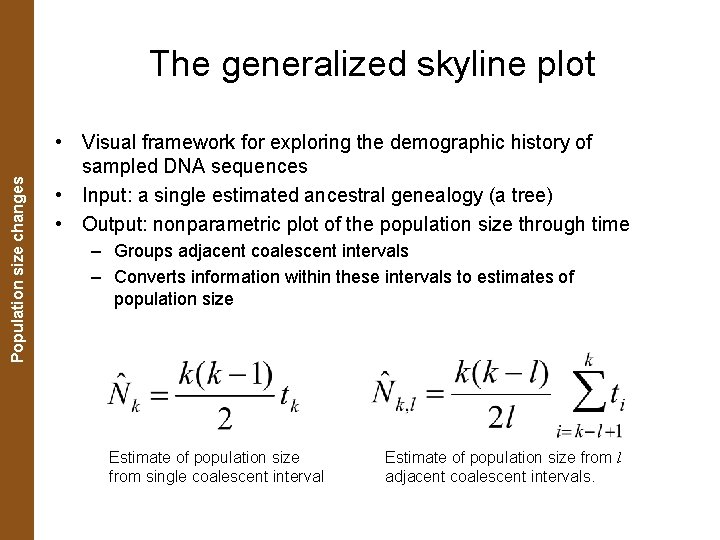
The Coalescent. Population and Measurably Evolving Populations size changes The generalized skyline plot • Visual framework for exploring the demographic history of sampled DNA sequences • Input: a single estimated ancestral genealogy (a tree) • Output: nonparametric plot of the population size through time – Groups adjacent coalescent intervals – Converts information within these intervals to estimates of population size Estimate of population size from single coalescent interval Estimate of population size from l adjacent coalescent intervals.
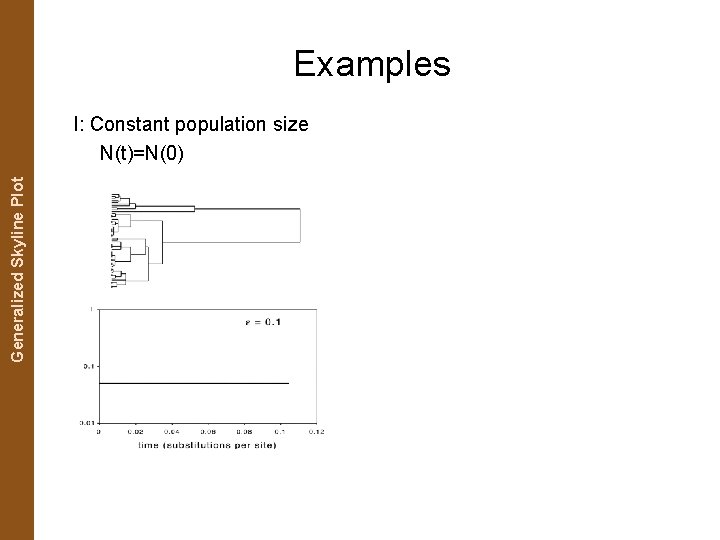
The Coalescent. Generalized and Measurably Evolving Skyline Plot Populations Examples I: Constant population size N(t)=N(0)
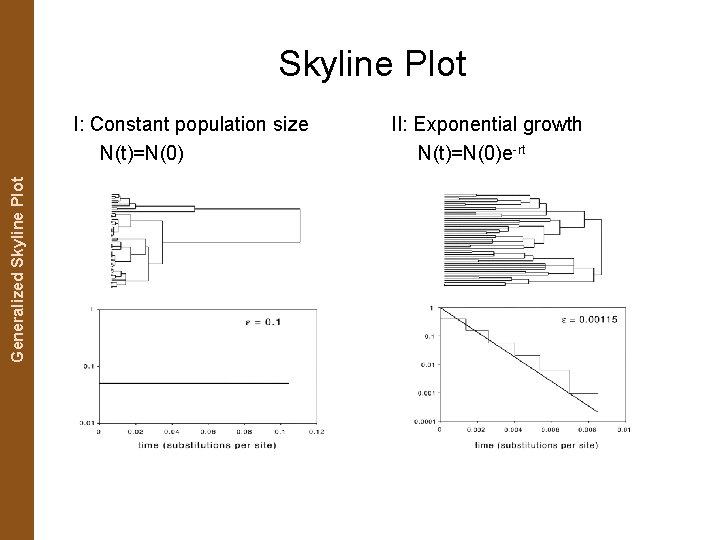
The Coalescent. Generalized and Measurably Evolving Skyline Plot Populations Skyline Plot I: Constant population size N(t)=N(0) II: Exponential growth N(t)=N(0)e-rt
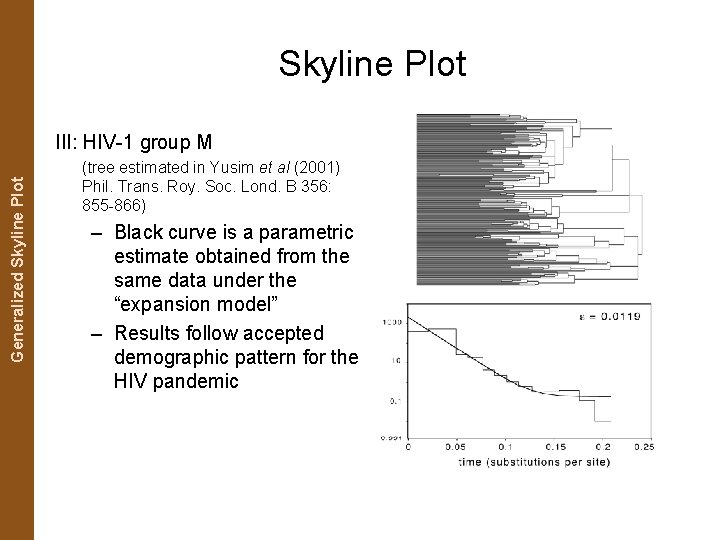
The Coalescent. Generalized and Measurably Evolving Skyline Plot Populations Skyline Plot III: HIV-1 group M (tree estimated in Yusim et al (2001) Phil. Trans. Roy. Soc. Lond. B 356: 855 -866) – Black curve is a parametric estimate obtained from the same data under the “expansion model” – Results follow accepted demographic pattern for the HIV pandemic
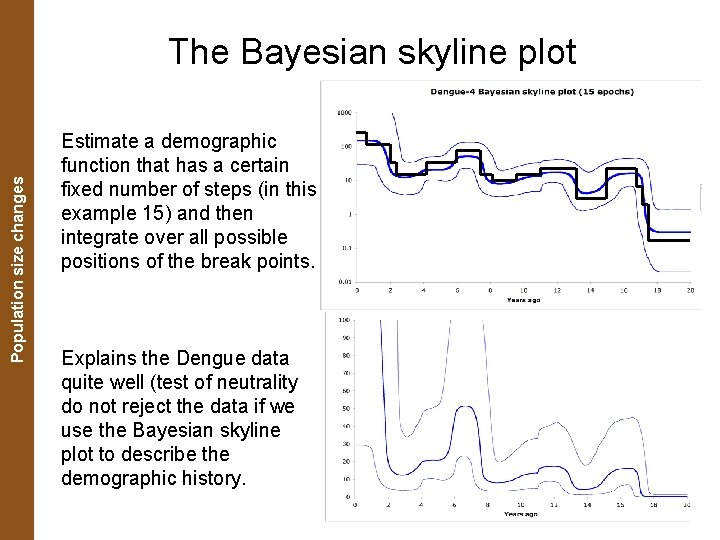
The Coalescent. Population and Measurably Evolving Populations size changes The Bayesian skyline plot Estimate a demographic function that has a certain fixed number of steps (in this example 15) and then integrate over all possible positions of the break points. Explains the Dengue data quite well (test of neutrality do not reject the data if we use the Bayesian skyline plot to describe the demographic history.

The Coalescent. Population and Measurably Evolving Populations size changes Prior/Model: population is autocorrelated through time
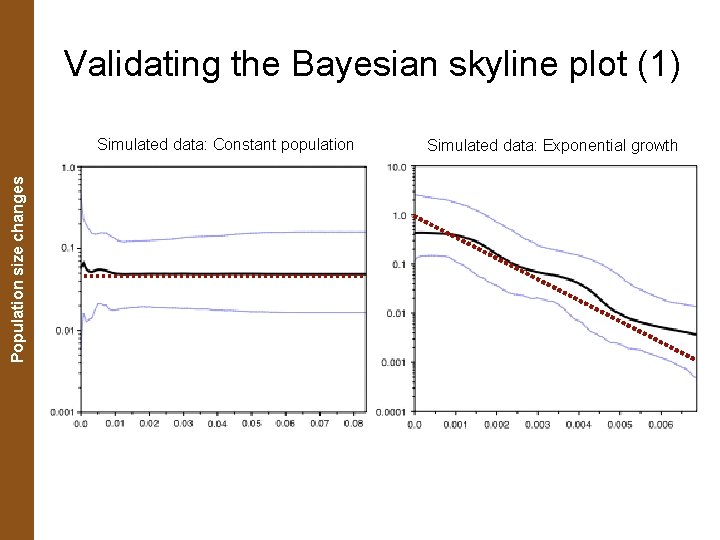
The Coalescent. Population and Measurably Evolving Populations size changes Validating the Bayesian skyline plot (1) Simulated data: Constant population Simulated data: Exponential growth
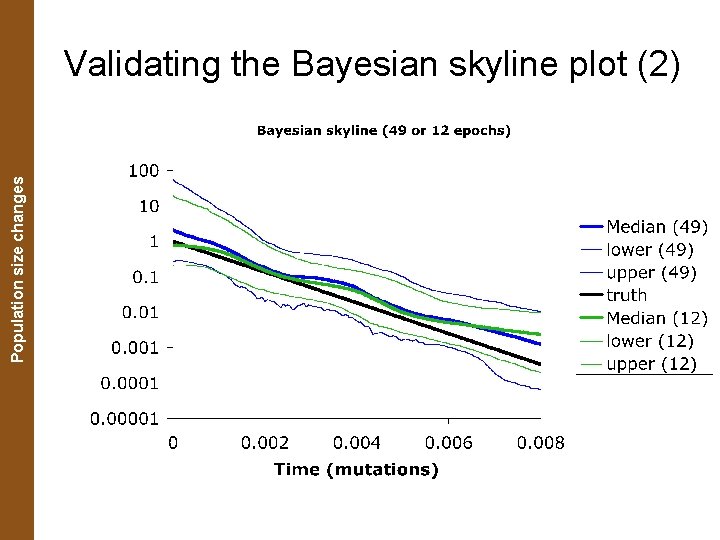
The Coalescent. Population and Measurably Evolving Populations size changes Validating the Bayesian skyline plot (2)
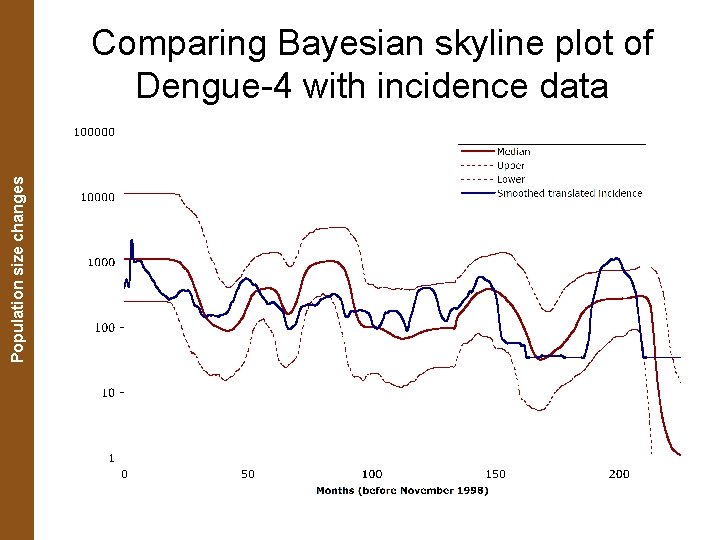
The Coalescent. Population and Measurably Evolving Populations size changes Comparing Bayesian skyline plot of Dengue-4 with incidence data
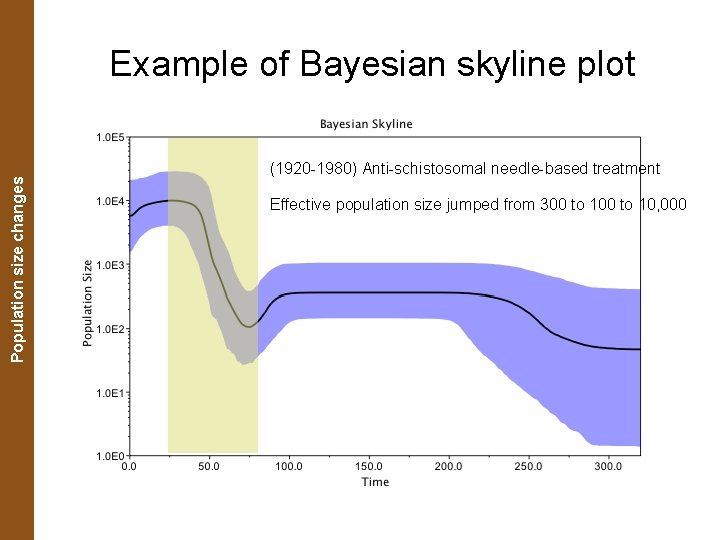
The Coalescent. Population and Measurably Evolving Populations size changes Example of Bayesian skyline plot (1920 -1980) Anti-schistosomal needle-based treatment Effective population size jumped from 300 to 10, 000
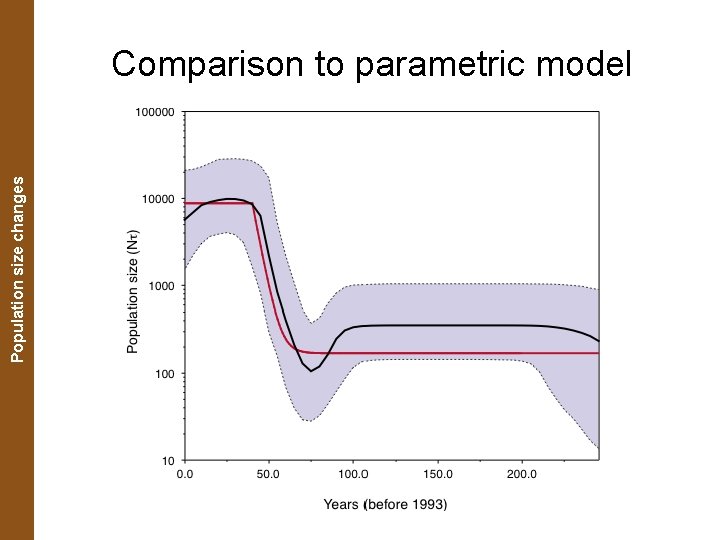
The Coalescent. Population and Measurably Evolving Populations size changes Comparison to parametric model
The Coalescent and Measurably Evolving Populations http: //evolve. zoo. ox. ac. uk/BEAST
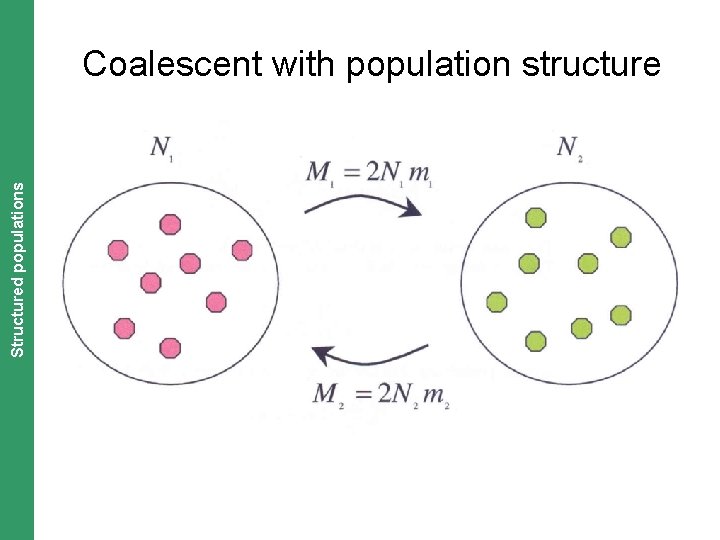
The Coalescent. Structured and Measurably Evolving Populations populations Coalescent with population structure
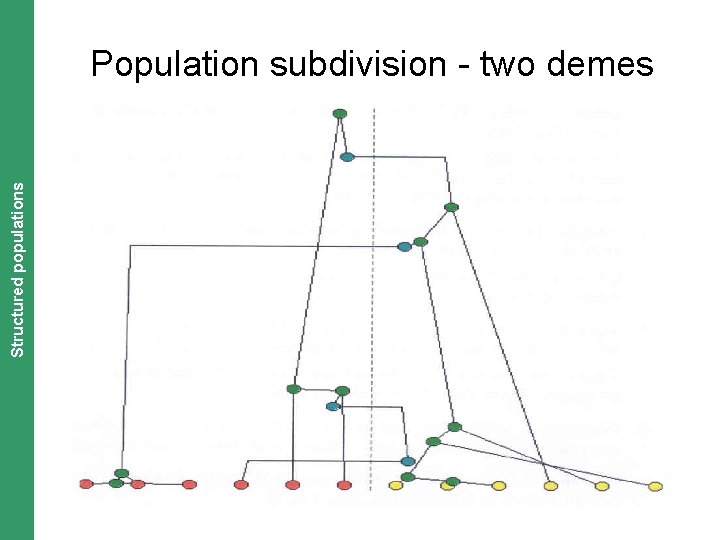
The Coalescent. Structured and Measurably Evolving Populations populations Population subdivision - two demes
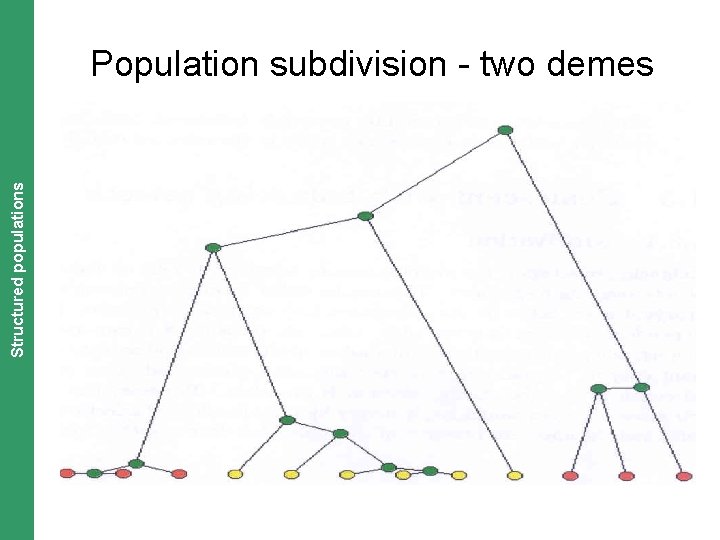
The Coalescent. Structured and Measurably Evolving Populations populations Population subdivision - two demes
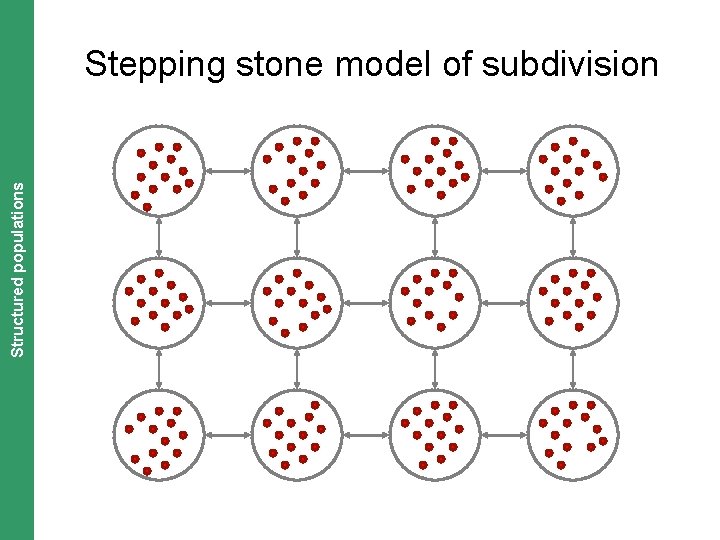
The Coalescent. Structured and Measurably Evolving Populations populations Stepping stone model of subdivision
The Coalescent. Structured and Measurably Evolving Populations populations Human migration From Cavalli-Sforza, 2001
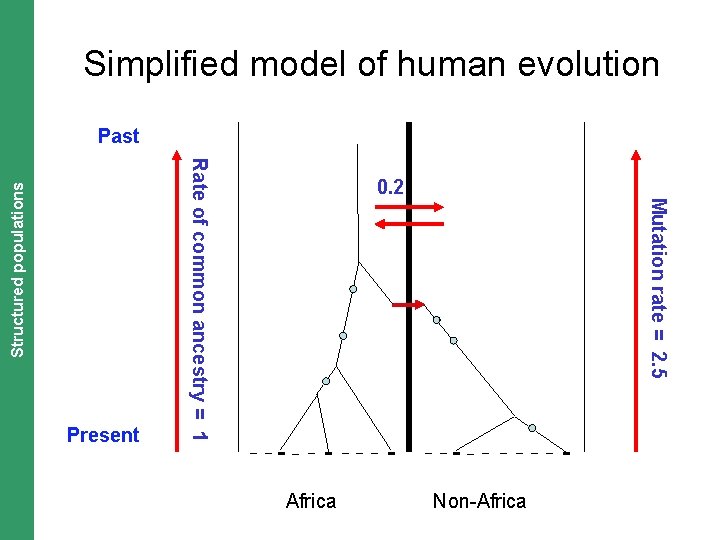
The Coalescent. Structured and Measurably Evolving Populations populations Simplified model of human evolution Past Rate of common ancestry = 1 Present Africa Mutation rate = 2. 5 0. 2 Non-Africa

The Coalescent and Measurably Evolving Populations Why Bayesian? • Probabilistic model-based inference – Can make simple statements about the probability of alternative hypotheses given the data • Markov chain Monte Carlo – Convenient computational technique – Allows for complex models: “if you can simulate you can sample” • Incorporates prior probabilities – P( |D) P(D| )P( ) – Convenient means of assessing alternative sets of assumptions – Allows incorporation of independent sources of information • Easy to include sources of uncertainty – Don’t need to assume perfect knowledge of tree (for example) – Can treat the tree and a nuisance parameter and focus on parameters of interest (strength of selection, mutation rate, growth rate, etc)

The Coalescent and Measurably Evolving Populations Conclusions & cautionary remarks • Bayesian MCMC has advantages – a useful tool for exploring prior hypotheses – Good for assessing levels of uncertainty – Complex models can be investigated on practical datasets • Bayesian MCMC has disadvantages – Diagnostics are difficult, and it is essentially impossible to guarantee correctness – Model comparison can be difficult – Requires large programs that are difficult to optimize and debug.

The Coalescent and Measurably Evolving Populations Conclusions & cautionary remarks (2) • Population genetics has advantages – provides a framework for objective analysis of genetic data – Allows interpretation of genetic data in terms of biological properties of virus – Can be extended to include selection, recombination et cetera • Population genetics has disadvantages – Models are still too simple – Assumptions are too strong – Extending to complex models that include changing selection pressures and recombination are possible in MCMC but still very difficult!
- Slides: 87