The Central Dogma of Molecular Biology by E

The Central Dogma of Molecular Biology by E. Börje Lindström This learning object has been funded by the European Commissions FP 6 Bio. Min. E project
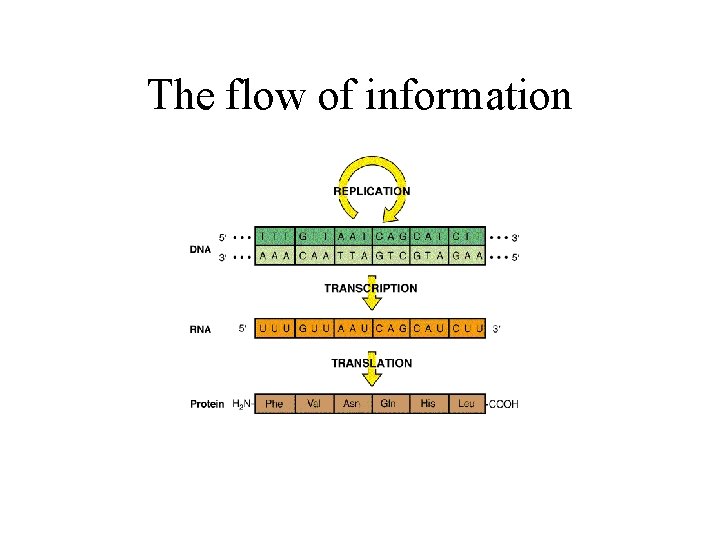
The flow of information

DNA molecule • General structure: - double stranded - complementary - helical - antiparallel • Strands: - backbone of alternating phosphate and deoxyribos units - four different bases; adenine (A), guanine (G), cytosine ( C ), and thymine (T). • Double helix: • Major and Minor groove - due to base pairing: A=T and G C

DNA molecule, cont. • Size: - units: kilobase (kb) or kilobase pairs (kb pairs) - E. coli chromosome 4 700 kb pairs • Form: - closed chromosome molecule (in bacteria) - 1 mm long packing problem in bacteria - solved by supercoiling • DNA binding proteins: Un-specific: - histones Specific: -Repressors - RNA polymerase - restriction enzymes - modification enzymes
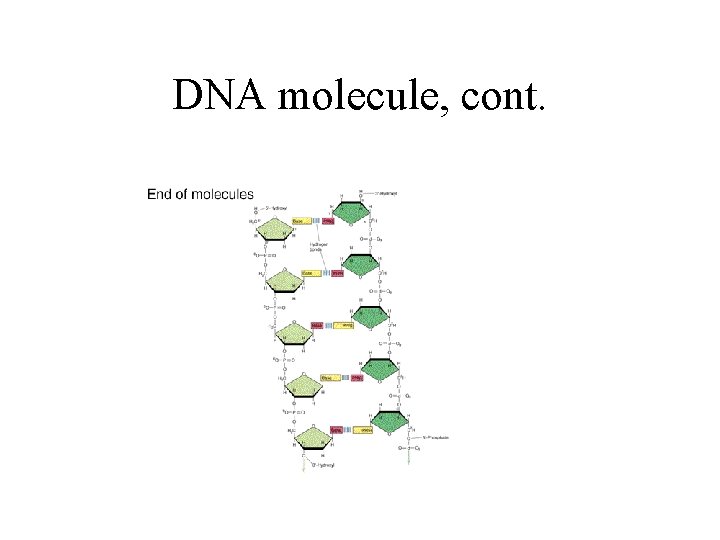
DNA molecule, cont.
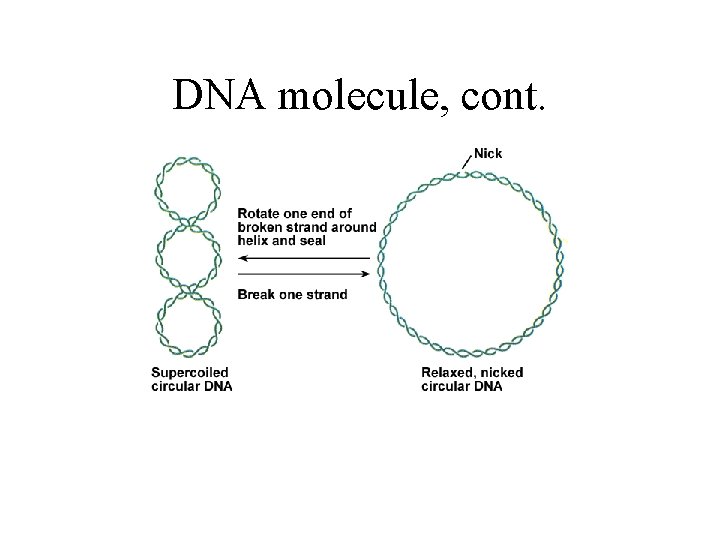
DNA molecule, cont.

DNA replication General • Semi conservative: -new DNA molecules contain: 1 old strand 1 new strand • use a ’template’: - one of the strand is used • ’primers’: -usually a piece of RNA - DNA-polymerase unable to start replication
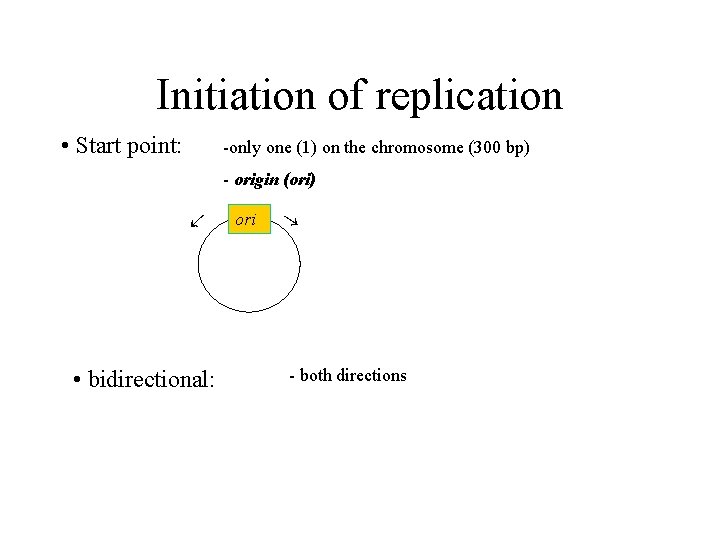
Initiation of replication • Start point: -only one (1) on the chromosome (300 bp) - origin (ori) • bidirectional: ori - both directions

Synthesis of DNA (replication) • several enzymes involved (~ 20 pc) - DNA helicase Unwinding the molecule - DNA gyrase (topoisemerase II) Open up (cut) the strands - DNA-binding enzymes Protect ss-DNA from nucleases - Primase Synthesises the RNA primer - DNA-plymerase III Synthesis in direction 5’ 3’ There are 3 enz. in E. coli; pol I, II and III - DNA-plymerase I Removes the primer Repair any missing bp in DNA - DNA ligase Makes a phospho-di-ester bond (glueing)

Synthesis of DNA, cont. • ’leading’ and ’lagging’ strands: - leading: continous synthesis - lagging: dis-continous synthesis • proof-reading: - checking if any mitakes has been made - pol. III removes the wrong nucleotides (3’ 5’)
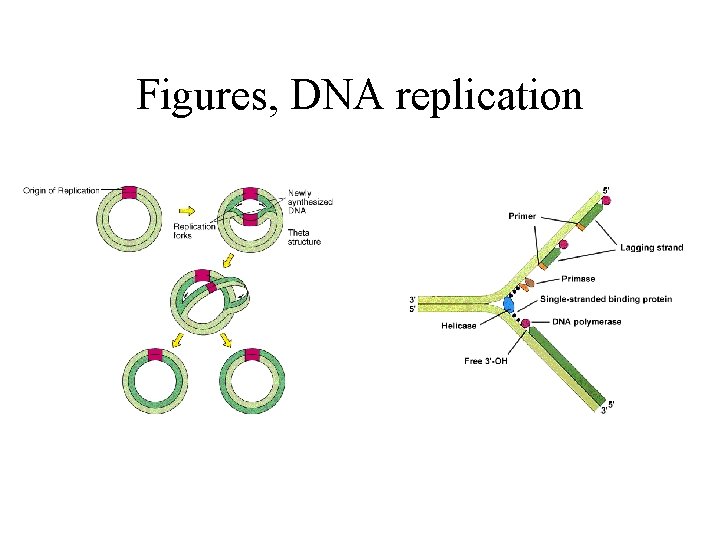
Figures, DNA replication

RNA transcription Three types of RNA: • m. RNA (genetical) • t. RNA (aa-carrier) • r. RNA (structural) Structure: -ss-stranded (internal ds secundary structures) - ribose - four different bases; adenine (A), guanine (G), cytosine ( C ), and uracile (U).

Synthesis of RNA • ds DNA is the template: • RNA polymerase: - only one of the strands - consists of four different subunits - a 2 bb’ = core enzyme - s recognises the start site • Direction of synthesis: - 3’ 5’

Start and stop of RNA synthesis • Where is the start ? - Note! No primers necessary! - The polymerase binds to the promoter - s recognises and attaches to the promoter region - ds-DNA opens up and the synthesis starts - s is detached and the core enzyme continues • Where does the synthesis stop? -termination at special DNA-sequenses, terminators - inverted repeates in DNA ’stem-loop’-structures in RNA

Promoters A sequence in DNA upstreams a structural gene: P -35 bp • -10 sequence SG -10 bp Pribnow box • Strong promoters bind s effective

m. RNA • Short half-time • Polycistronic (in bacteria) - information from several structural genes • Definitions: - operator (O): a gene that can be effected by a repressor protein - operon: structural genes with the same repressor P O SG 1 SG 2 SG 3

Translation Necessary substances: • m. RNA • ribosomes • t. RNA + aa t. RNAaa (attached aa) • different factors • enzymes • energy

t. RNA • DNA-genes: - Linear t. RNA form (primary) - cloverleaf structure (secundary) • Two peoperties: - binds aa (enzymatic) - binds to m. RNA (codon) with its anti-codon
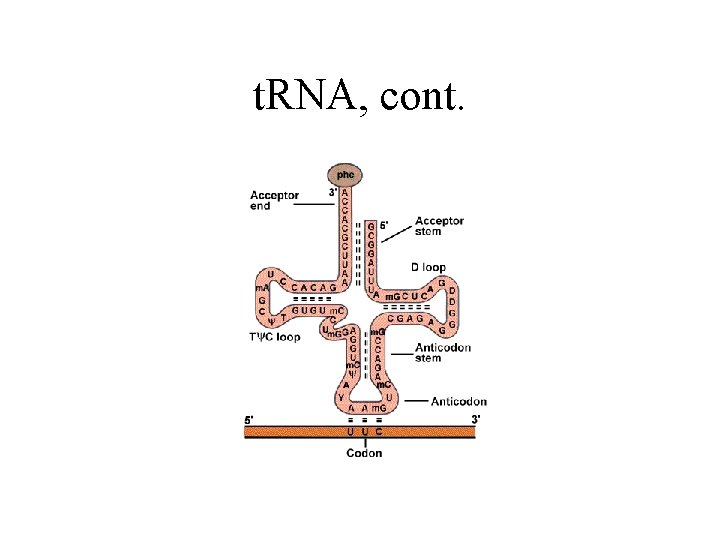
t. RNA, cont.

Synthesis of proteins A four (4) step process: • Initiation • Elongation • Termination-release • Peptide folding • Initiation: -a complex of - 30 S subunit, - f-meth-t. RNA, (start codon AUG in m. RNA) - m. RNA and - initiation factors are formed • Shine-Delgarno sequence -3 -9 bases in m. RNA - complementary to 16 S r. RNA - addition of 50 S subunit

Synthesis of proteins, cont. • Elongation: -several elongation factors are needed - Next aa-t. RNA is added to the A-site (ribosome) - a peptide bond is created - the peptide is moved to the A-site - translocation to the P-site during - movement of the ribosome forward - a free A-site is created … -Etc. • polysomes: - m. RNA with several ribosomes

Synthesis of proteins, cont. • Termination: -stop codes in m. RNA - UAA, UAG and UGA; nonsence codes - no t. RNA for these codes exist - release factors RF 1 -3 release the protein - the ribosomes disintegrate • The genetic code: - in m. RNA • 3 bases - 1 aa • 43 = combinations -but only 24 aa - degenerated code - the aa has several codes
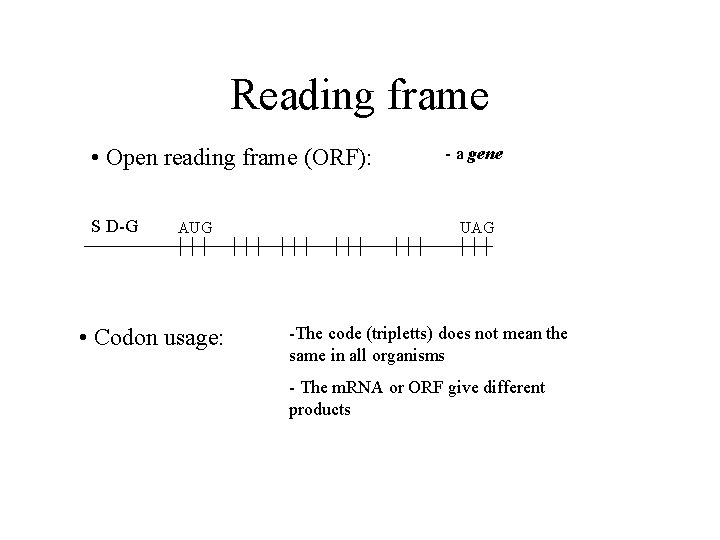
Reading frame • Open reading frame (ORF): S D-G AUG • Codon usage: - a gene UAG -The code (tripletts) does not mean the same in all organisms - The m. RNA or ORF give different products
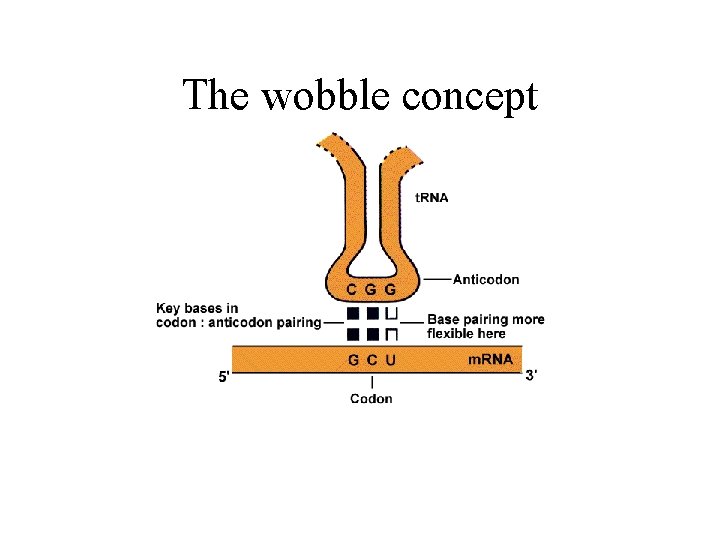
The wobble concept
- Slides: 24