The bioinformatics behind shotgun metagenomic sequencing Roche UG

The bioinformatics behind shotgun metagenomic sequencing Roche UG 2012 Rob Edwards

Outline Metagenomics Annotation Virus: host prediction
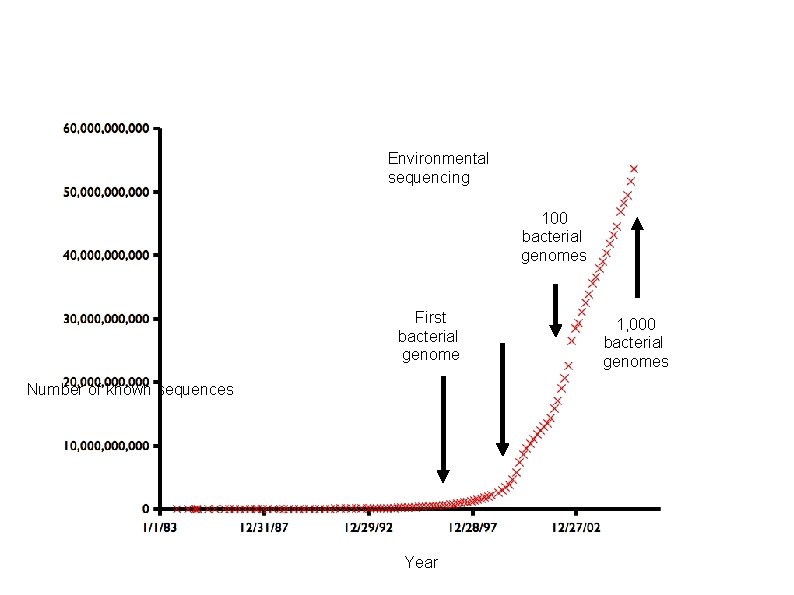
How Much has Been sequenced? Environmental sequencing 100 bacterial genomes First bacterial genome Number of known sequences Year 1, 000 bacterial genomes
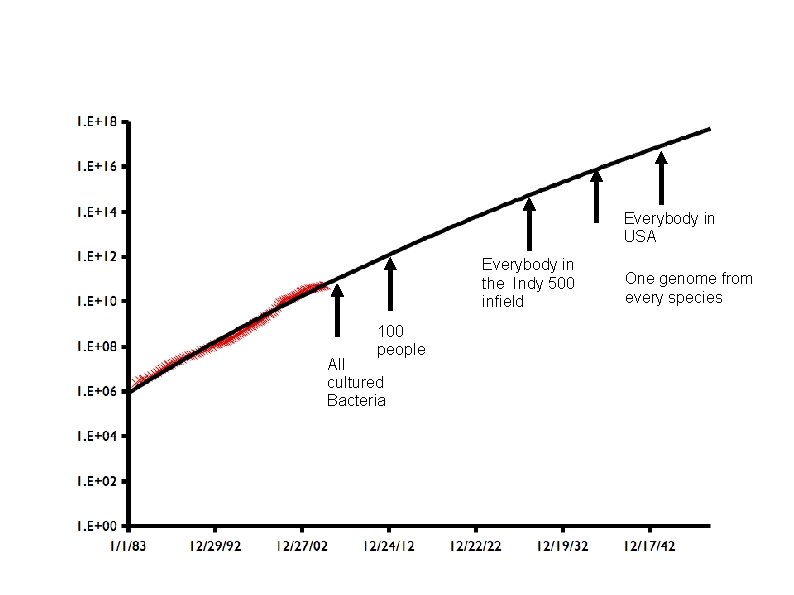
How Much will Be Sequenced? Everybody in USA Everybody in the Indy 500 infield 100 people All cultured Bacteria One genome from every species Most major microbial environments
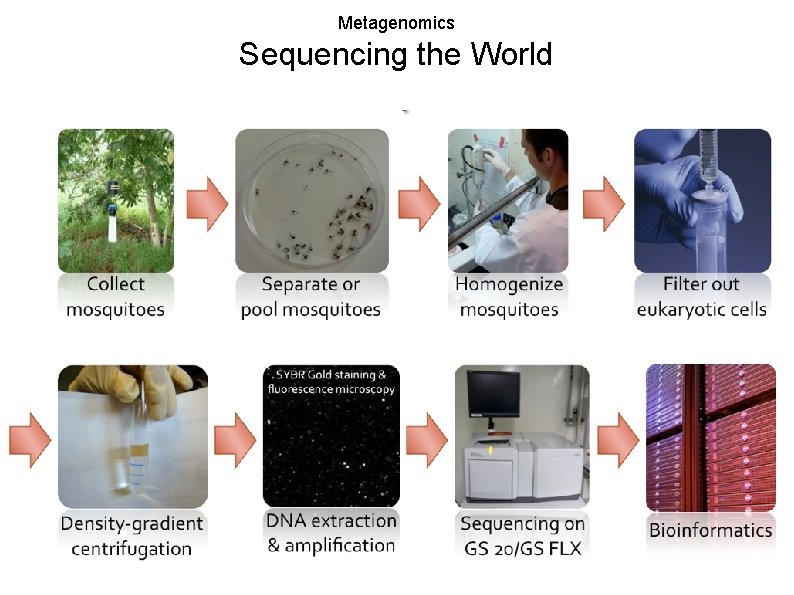
Metagenomics Sequencing the World

Shark Metagenomes

Shark Metagenomes

Metagenomics Analysis Steps 1. clean the data (prinseq) Check the quality of the data Remove bad reads 2. annotate the data (rtmg) What is there? Who is there? 3. analyze the data

Prinseq Clean, dereplicate, check quality, analyze PRINSEQ Results Rob http: //edwards. sdsu. edu/prinseq/

Data Processed May 2012 Datasets processed: Sequences processed: Bases processed: 6, 223 8, 698, 402, 976 802, 472, 118, 0 http: //edwards. sdsu. edu/prinseq/
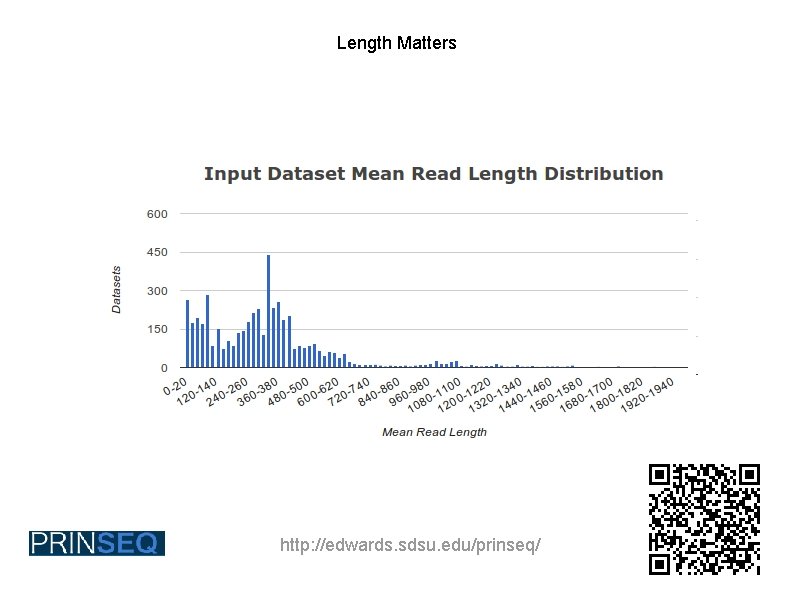
Length Matters http: //edwards. sdsu. edu/prinseq/
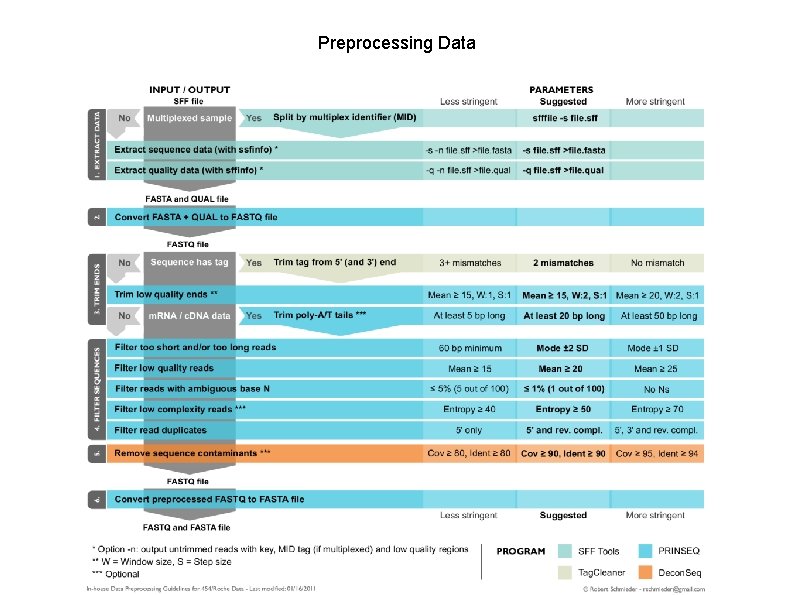
Preprocessing Data

Annotating Metagenomes Identify functions // organisms present in the sample � BLAST is very slow � Immediate processing of data � RTMG http: //edwards. sdsu. edu/rtmg/

A Better Way To Annotate Metagenomes Real time metagenomics RTMg RTMG http: //edwards. sdsu. edu/rtmg/
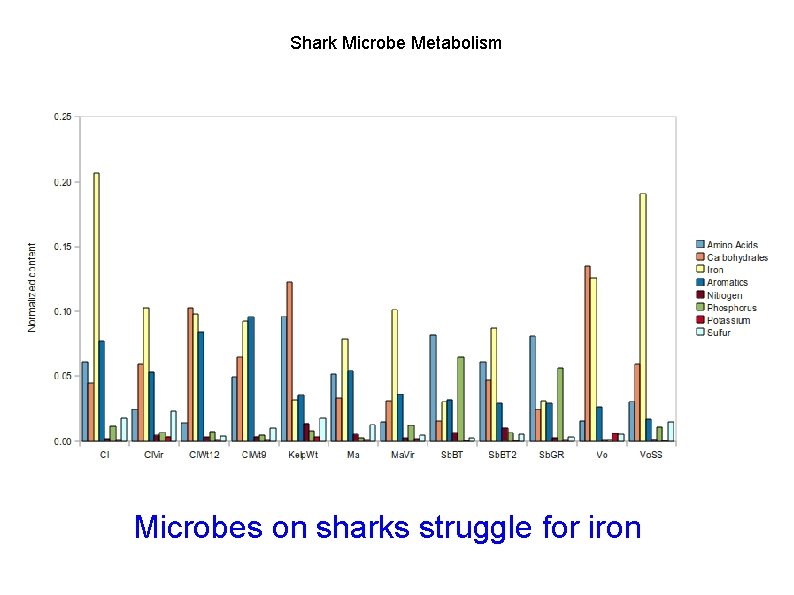
Shark Microbe Metabolism Microbes on sharks struggle for iron

Host – Virus Interactions Can we predict the host a virus infects just from the sequence? Bas Michiyo

How To Predict Hosts Count all 2 -, 3 -, 4 -, 5 -, 6 -, 7 - bp sequences � AA, AG, AC, AT, GA, GG, GC, GT … � AAA, AAG, AAC, AAT, AGA, AGG, AGC, AGT … � AAAA, AAAG, AAAC, AAAT, AAGA, AAGG, … � Count in host and virus � Use machine learning (Random Forest) to identify which hosts and which viruses match �
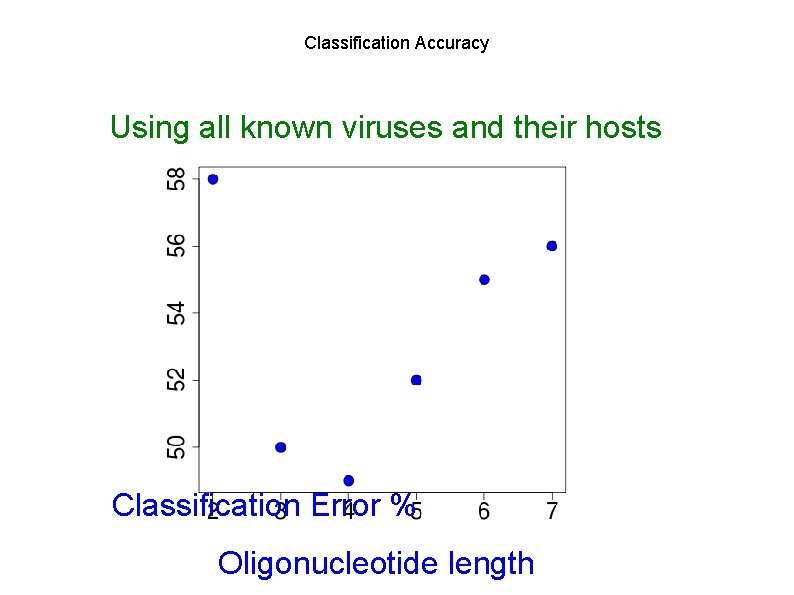
Classification Accuracy Using all known viruses and their hosts Classification Error % Oligonucleotide length

Length Matters! Using all known viruses and their hosts 50 40 200 bp reads 30 Using known samples as control: 20 ~5% of reads classified 10 0 Prediction percent Correct predictions Sequencing error little effect Wrong predictions
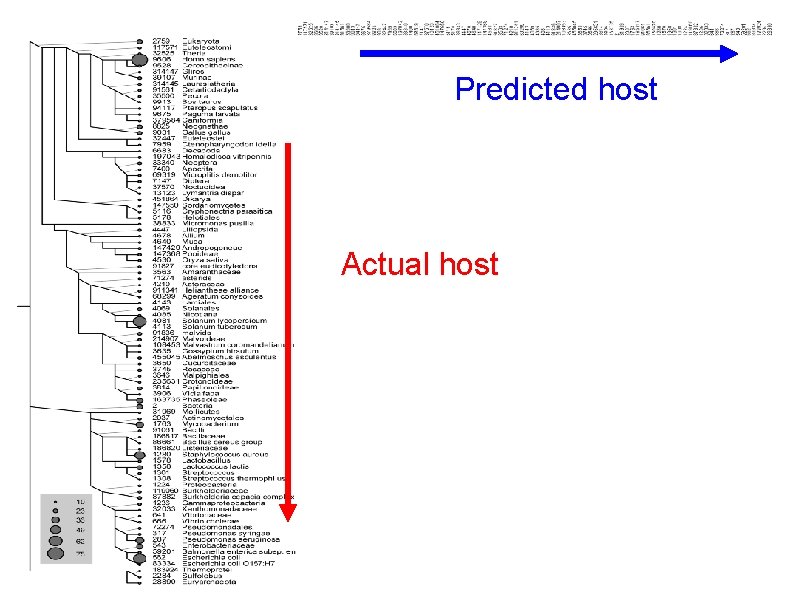
Predicted host Actual host

Predicted 89 % of misclassifications are near-neighbors 11% are outside the near neighbors Actual
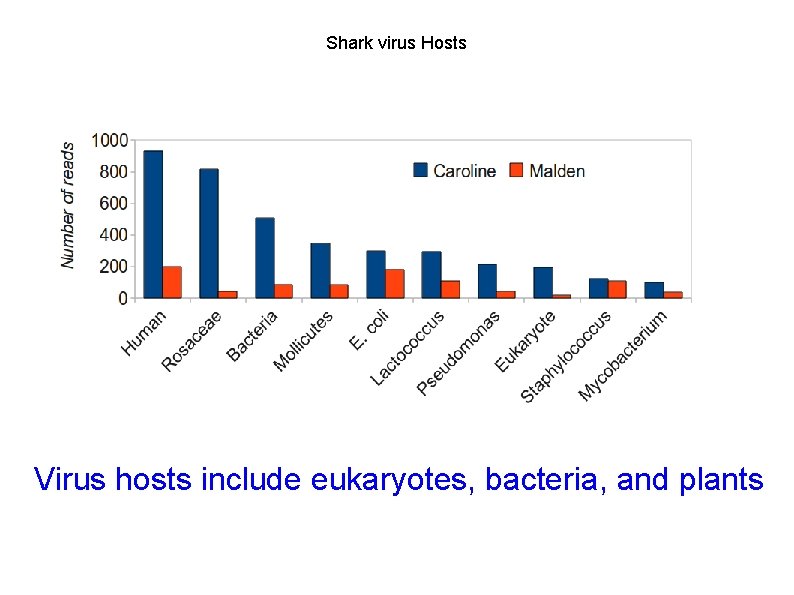
Shark virus Hosts Virus hosts include eukaryotes, bacteria, and plants

Thanks Liz Forest Stuart Katie Alan

Take Home Points Check your data (e. g. prinseq) Annotate the data (e. g. RTMg) Analyze your data

The Lab Ramy � Jeremy � Bas � Dave � Sajia � Rob � Kate � Joakim � Brad � Steve � Sheridan � Stephanie � Adam � Carny � Rima � Josh � Daniel � Michiyo � Vasken � Matt S � Matt H � Bianca � Andrés � Nick C � Nick T � Brian � Geni � Jimmy � Amanda �

Funding Ph. An. To. Me TUES Viral Dark Matter Brazil-US Marine Sciences Consortium Coral Reef Image Analysis
- Slides: 26