The Biochemical Abstract Machine BIOCHAM Logic programming steps

The Biochemical Abstract Machine BIOCHAM Logic programming steps towards formal biology François Fages, INRIA Rocquencourt http: //contraintes. inria. fr/ Joint work with Nathalie Chabrier-Rivier and Sylvain Soliman ARC CPBIO “Process Calculi and Biology of Molecular Networks” Alexander Bockmayr , LORIA Nancy, Vincent Danos , CNRS Paris PPS, Vincent Schächter , Genoscope Evry , et al. http: //contraintes. inria. fr/cpbio/ François Fages Evry, April 2004

Current Revolution in Systems Biology • Elucidation of high-level biological processes in terms of their biochemical basis at the molecular level. • Mass production of genomic and post-genomic data: ARN expression, protein synthesis, protein-protein interactions, … • Need for a strong parallel effort on the formal representation biological processes: Systems Biology. of • Need formal tools for modeling and reasoning about their global behavior. François Fages Evry, April 2004
![Formalisms for Modeling Biochemical Systems • Diagrammatic notation • Boolean networks [Thomas 73] • Formalisms for Modeling Biochemical Systems • Diagrammatic notation • Boolean networks [Thomas 73] •](http://slidetodoc.com/presentation_image_h2/607819d83880d1abd758ea35f2143c02/image-3.jpg)
Formalisms for Modeling Biochemical Systems • Diagrammatic notation • Boolean networks [Thomas 73] • Milner’s pi–calculus [ Regev -Silverman-Shapiro 99 -01, Nagasali et al. 00] • Concurrent transition systems [Chabrier-Chiaverini-Danos-Fages. Schachter 03] Biochemical abstract machine BIOCHAM [Chabrier-Fages-Soliman 03] Pathway logic [ Eker-Knapp-Laderoute-Lincoln-Meseguer-Sonmez 02] • Bio- ambients [Regev-Panina-Silverman-Cardelli-Shapiro 03] • • Differential equations Hybrid Petri nets [ Hofestadt-Thelen 98, Matsuno et al. 00] Hybrid automata [ Alur et al. 01, Ghosh -Tomlin 01] Hybrid concurrent constraint languages [Bockmayr-Courtois François Fages 01] Evry, April 2004

Our Goal Beyond simulation, provide formal tools for querying, validating and completing biological models. Our proposal: • Use of temporal logic CTL as a query language for models of biological processes; • Use of concurrent transition systems for their modeling; • Use of symbolic and constraint-based model checkers for automatically evaluating CTL queries in qualitative and quantitative models. • Use of inductive logic programming for learning models In course, learn and teach bits of biology with logic programs. François Fages Evry, April 2004

References A wonderful textbook: Molecular Cell Biology. 5 th Edition, 1100 pages+CD , Freeman Publ. Lodish , Berk , Zipursky , Matsudaira , Baltimore, Darnell. Nov. 2003. Genes and signals. Ptashne , Gann. CSHL Press. 2002. Modeling dynamic phenomena in molecular and cellular biology Segel. Cambridge Univ. Press. 1987. . Modeling and querying bio-molecular interaction networks. Chabrier , Chiaverini , Danos , Fages, Schächter. To appear in TCS. 2004. The biochemical abstract machine BIOCHAM. Chabrier , Fages, Soliman. http: //contraintes. inria. fr/BIOCHAM François Fages Evry, April 2004

Plan of the Talk 1. Introduction 2. A simple algebra of cell molecules 3. Concurrent transition systems of biochemical reactions • Example of the mammalian cell cycle control 4. Temporal logic CTL as a query language • Computational results with BIOCHAM 5. Learning reaction rules • An experiment with inductive logic programming 6. Kinetics models • • Simulation with differential equations Hybrid systems 7. Conclusion François Fages Evry, April 2004

2. A Simple Algebra of Cell Molecules Small molecules : covalent bonds (outer electrons shared) 50 -200 kcal/mol • 70% water • 1% ions • 6% amino acids (20), nucleotides (5), fats, sugars, ATP, ADP, … Macromolecules : hydrogen bonds, ionic, hydrophobic, Waals 1 -5 kcal/mol Stability and bindings determined by the number of weak bonds: 3 D shape • 20% proteins (50 -10 4 amino acids) • RNA (10 2 -104 nucleotides AGCU) Evry, April 2004 François Fages • DNA (10 2 -106 nucleotides AGCT)
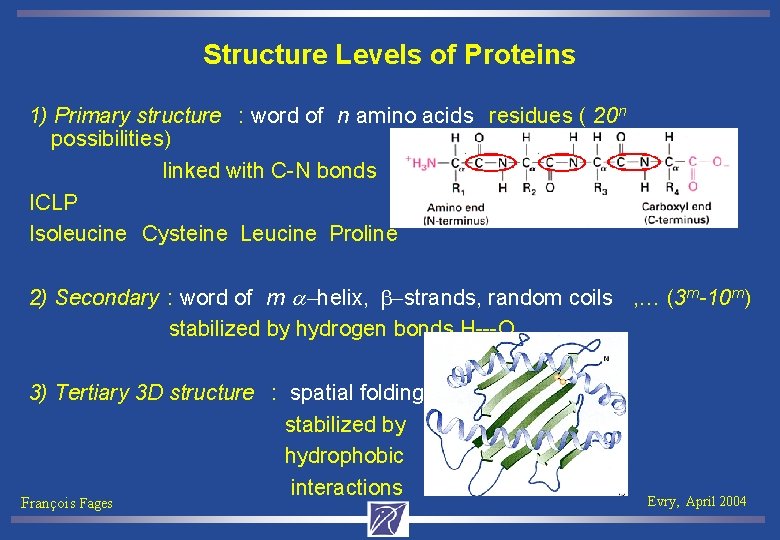
Structure Levels of Proteins 1) Primary structure : word of n amino acids residues ( 20 n possibilities) linked with C-N bonds ICLP Isoleucine Cysteine Leucine Proline 2) Secondary : word of m a-helix, b-strands, random coils , … (3 m -10 m ) stabilized by hydrogen bonds H---O 3) Tertiary 3 D structure : spatial folding stabilized by hydrophobic interactions François Fages Evry, April 2004
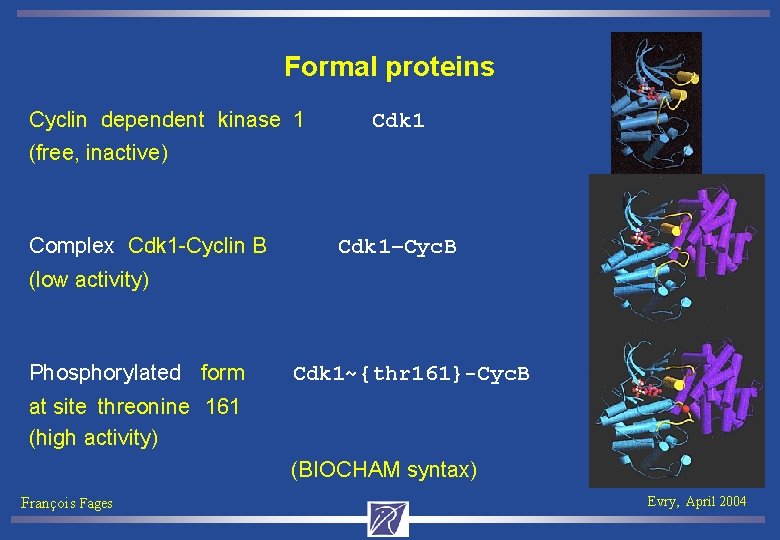
Formal proteins Cyclin dependent kinase 1 Cdk 1 (free, inactive) Complex Cdk 1 -Cyclin B Cdk 1–Cyc. B (low activity) Phosphorylated form Cdk 1~{thr 161}-Cyc. B at site threonine 161 (high activity) (BIOCHAM syntax) François Fages Evry, April 2004

Abstraction of gene expression DNA RNA protein DNA: word over 4 nucleotides Adenine, Guanine, Cytosine, Thymine double helix of pairs A--T and C---G Replication : DNA synthesis Genes: parts of DNA Transcription : RNA copying from a gene #ERCC 1 -(PRB-JUN-CFOS) François Fages Evry, April 2004

BIOCHAM Algebra of Cell Molecules E : : = Name|E-E|E~{E, …, E}|(E) Names: molecules, proteins, @processes… S : : = _|E+S #gene binding sites, abstract - : binding operator for protein complexes, gene binding sites, … Associative and commutative. ~{…}: modification operator for phosphorylated sites, … Set of modified sites (Associative, Commutative, Idempotent). + : solution operator, “soup aspect”, Assoc. Comm. Idempotent, Neutral _ Evry, April 2004 François Fages No membranes, no transport formalized. Bitonal calculi [ Cardelli 03].

Plan of the talk 1. Introduction 2. A simple algebra of cell molecules 3. Concurrent transition systems of biochemical reactions • Example of the mammalian cell cycle control 4. Temporal logic CTL as a query language • Computational results with BIOCHAM 5. Learning reaction rules • An experiment with inductive logic programming 6. Kinetics models • • Simulation with differential equations Hybrid systems 7. Conclusion François Fages Evry, April 2004

3. Concurrent Transition Syst. of Biochemical Reactions Enzymatic reactions: R : : = S=>S | S=[E]=>S | S=[R]=>S | S<=[E]=>S (where A<=>B stands for A=>B B=>A and A=[C]=>B for A+C=>B+C, etc. ) define a concurrent transition system over integers denoting the multiplicity of the molecules ( multiset rewriting). One can associate a finite abstract CTS over boolean state variables denoting the presence/absence of molecules which correctly over-approximates the set of all possible behaviors a reaction A+B=>C+D is translated with 4 rules for possible consumption: A+B+C+D A+B +C+D Evry, April 2004 François Fages A+B A+ B+C+D A+B A+ B+C+D

Six Rule Schemas Complexation : A + B => A-B Cdk 1+Cyc. B => Cdk 1–Cyc. B Decomplexation A-B => A + B Phosphorylation : A =[C]=> A~{p } Dephosphorylation A~{p } =[C]=> A Cdk 1–Cyc. B =[Myt 1]=> Cdk 1~{thr 161}-Cyc. B Cdk 1~{thr 14, tyr 15}-Cyc. B =[Cdc 25~{Nterm}]=> Cdk 1 -Cyc. B Synthesis : _ =[C]=> A. _ =[#Ge 2 -E 2 f 13 -Dp 12]=> Cyc. A Degradation : A =[C]=> _. Cyc. E =[@Ubi. Pro]=> _ François Fages (not for Cyc. E-Cdk 2 which is stable) Evry, April 2004
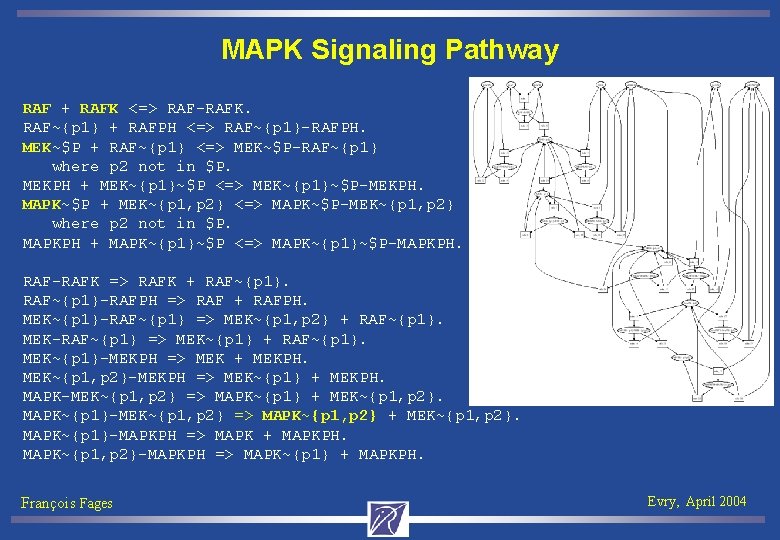
MAPK Signaling Pathway RAF + RAFK <=> RAF-RAFK. RAF~{p 1} + RAFPH <=> RAF~{p 1}-RAFPH. MEK~$P + RAF~{p 1} <=> MEK~$P-RAF~{p 1} where p 2 not in $P. MEKPH + MEK~{p 1}~$P <=> MEK~{p 1}~$P-MEKPH. MAPK~$P + MEK~{p 1, p 2} <=> MAPK~$P-MEK~{p 1, p 2} where p 2 not in $P. MAPKPH + MAPK~{p 1}~$P <=> MAPK~{p 1}~$P-MAPKPH. RAF-RAFK => RAFK + RAF~{p 1}-RAFPH => RAF + RAFPH. MEK~{p 1}-RAF~{p 1} => MEK~{p 1, p 2} + RAF~{p 1}. MEK-RAF~{p 1} => MEK~{p 1} + RAF~{p 1}. MEK~{p 1}-MEKPH => MEK + MEKPH. MEK~{p 1, p 2}-MEKPH => MEK~{p 1} + MEKPH. MAPK-MEK~{p 1, p 2} => MAPK~{p 1} + MEK~{p 1, p 2}. MAPK~{p 1}-MEK~{p 1, p 2} => MAPK~{p 1, p 2} + MEK~{p 1, p 2}. MAPK~{p 1}-MAPKPH => MAPK + MAPKPH. MAPK~{p 1, p 2}-MAPKPH => MAPK~{p 1} + MAPKPH. François Fages Evry, April 2004
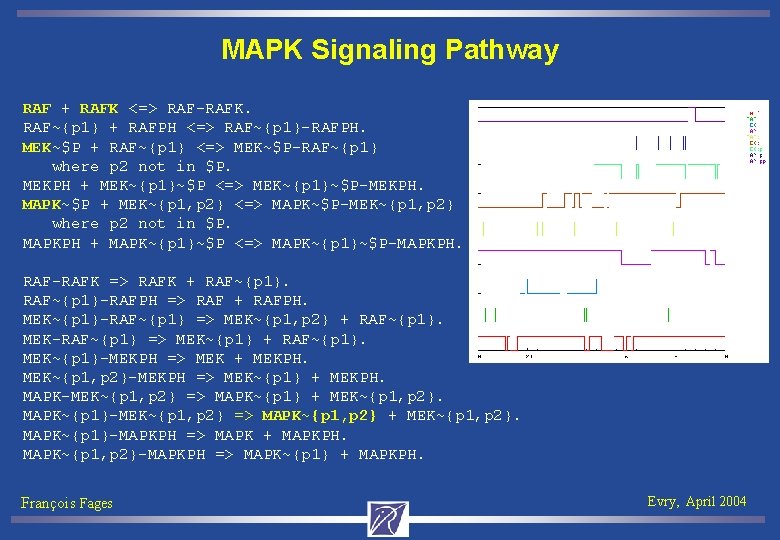
MAPK Signaling Pathway RAF + RAFK <=> RAF-RAFK. RAF~{p 1} + RAFPH <=> RAF~{p 1}-RAFPH. MEK~$P + RAF~{p 1} <=> MEK~$P-RAF~{p 1} where p 2 not in $P. MEKPH + MEK~{p 1}~$P <=> MEK~{p 1}~$P-MEKPH. MAPK~$P + MEK~{p 1, p 2} <=> MAPK~$P-MEK~{p 1, p 2} where p 2 not in $P. MAPKPH + MAPK~{p 1}~$P <=> MAPK~{p 1}~$P-MAPKPH. RAF-RAFK => RAFK + RAF~{p 1}-RAFPH => RAF + RAFPH. MEK~{p 1}-RAF~{p 1} => MEK~{p 1, p 2} + RAF~{p 1}. MEK-RAF~{p 1} => MEK~{p 1} + RAF~{p 1}. MEK~{p 1}-MEKPH => MEK + MEKPH. MEK~{p 1, p 2}-MEKPH => MEK~{p 1} + MEKPH. MAPK-MEK~{p 1, p 2} => MAPK~{p 1} + MEK~{p 1, p 2}. MAPK~{p 1}-MEK~{p 1, p 2} => MAPK~{p 1, p 2} + MEK~{p 1, p 2}. MAPK~{p 1}-MAPKPH => MAPK + MAPKPH. MAPK~{p 1, p 2}-MAPKPH => MAPK~{p 1} + MAPKPH. François Fages Evry, April 2004
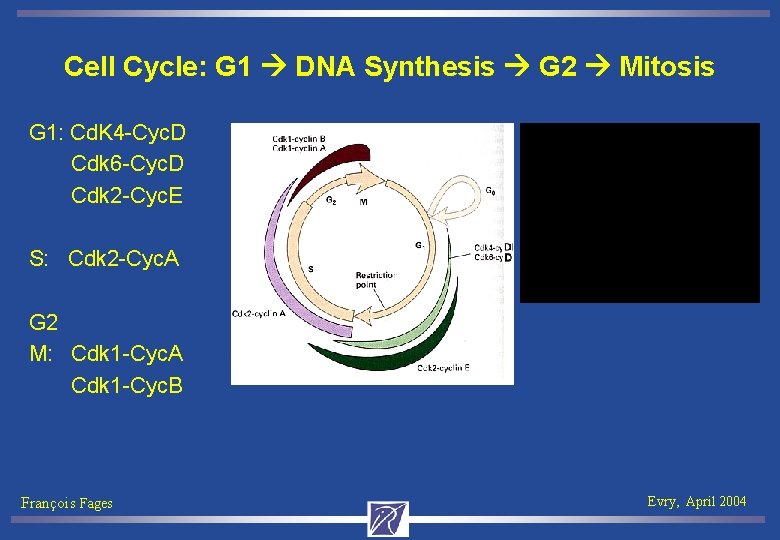
Cell Cycle: G 1 DNA Synthesis G 2 Mitosis G 1: Cd. K 4 -Cyc. D Cdk 6 -Cyc. D Cdk 2 -Cyc. E S: Cdk 2 -Cyc. A G 2 M: Cdk 1 -Cyc. A Cdk 1 -Cyc. B François Fages Evry, April 2004
![Mammalian Cell Cycle Control Map [Kohn 99] François Fages Evry, April 2004 Mammalian Cell Cycle Control Map [Kohn 99] François Fages Evry, April 2004](http://slidetodoc.com/presentation_image_h2/607819d83880d1abd758ea35f2143c02/image-18.jpg)
Mammalian Cell Cycle Control Map [Kohn 99] François Fages Evry, April 2004
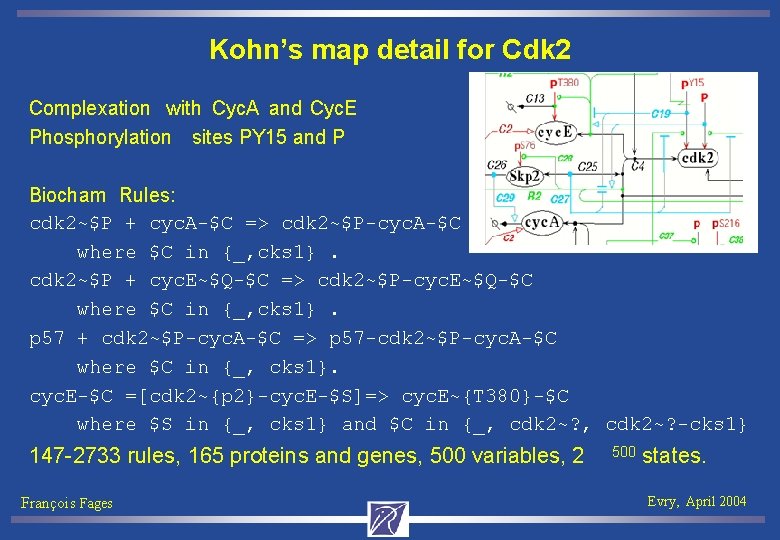
Kohn’s map detail for Cdk 2 Complexation with Cyc. A and Cyc. E Phosphorylation sites PY 15 and P Biocham Rules: cdk 2~$P + cyc. A-$C => cdk 2~$P-cyc. A-$C where $C in {_, cks 1}. cdk 2~$P + cyc. E~$Q-$C => cdk 2~$P-cyc. E~$Q-$C where $C in {_, cks 1}. p 57 + cdk 2~$P-cyc. A-$C => p 57 -cdk 2~$P-cyc. A-$C where $C in {_, cks 1}. cyc. E-$C =[cdk 2~{p 2}-cyc. E-$S]=> cyc. E~{T 380}-$C where $S in {_, cks 1} and $C in {_, cdk 2~? -cks 1} 147 -2733 rules, 165 proteins and genes, 500 variables, 2 François Fages 500 states. Evry, April 2004

Plan of the talk 1. Introduction 2. A simple algebra of cell molecules 3. Concurrent transition systems of biochemical reactions • Example of the mammalian cell cycle control 4. Temporal logic CTL as a query language • Expressivity and computational results 5. Learning reaction rules • An experiment with inductive logic programming 6. Kinetics models • Simulation with differential equations • Hybrid systems 7. Conclusion François Fages Evry, April 2004
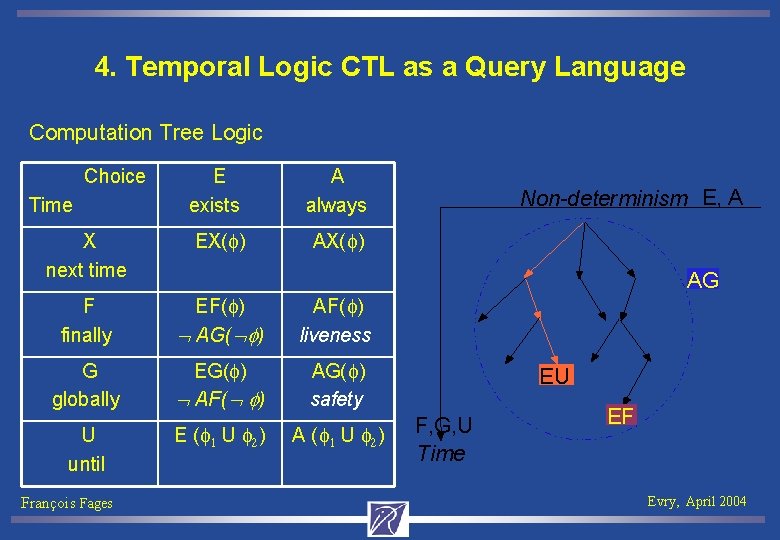
4. Temporal Logic CTL as a Query Language Computation Tree Logic Choice E exists A always X next time EX(f) AX(f) F finally EF(f) AG( f) AF(f) liveness G globally EG(f) AF( f) AG(f) safety Time U until François Fages Non-determinism E, A AG E (f 1 U f 2) A (f 1 U f 2) EU F, G, U Time EF Evry, April 2004

Kripke Structures A Kripke structure K is a triple (S; R; L) where S is a set of states, and R Sx. S is a total relation. s |= f if f is true in s, s |= E f if there is a path from s such that |= f, s |= A f if for every path from s, |= f if s |= f where s is the starting state of , |= X f if 1 |= f, |= F f if there exists k >0 such that k |= f, |= G f if for every k >0, k |= f, |= f 1 U f 2 iff there exists k>0 such that k |= f for all j < k j |= f. Following [Emerson 90] we identify a formula f to the set of states which satisfy it f ~ {s S : s |= f }. François Fages Evry, April 2004

Symbolic Model Checking is an algorithm for computing, in a given finite Kripke structure the set of states satisfying a CTL formula: {s S : s |= f }. Basic algorithm : represent K as a graph and iteratively label the nodes with the subformulas of f which are true in that node. Add f to the states satisfying f Add EF f (EX f) to the (immediate) predecessors of states labeled by f Add E(f 1 U f 2 ) to the predecessor states of f 2 while they satisfy f 1 Add EG f to the states for which there exists a path leading to a non trivial strongly connected component of the subgraph of states satisfying f Symbolic François Fages model checking : use OBDDs to represent states and Evry, transitions as boolean formulas (S is finite). April 2004

Biological Queries (1/3) About reachability : • Given an initial state init, can the cell produce some protein P? init EF(P) • Which are the states from which a set of products P 1, . . . , be produced simultaneously? EF(P 1^…^Pn) Pn can About pathways : • Can the cell reach a state s while passing by another state s 2? init EF(s 2^EFs) • Is state s 2 a necessary checkpoint for reaching state s? EF( s 2 U s) • Is it possible to produce P without using nor creating Q? EF( Q U s) • Can the François Fages cell reach a state s without violating some constraints. Evry, c? April 2004 init EF(c U s)

Biological Queries (2/3) About stability : • Is a certain (partially described) state s a stable state? s AG(s) (s denotes both the state and the formula describing it). • Is s a steady state (with possibility of escaping) ? s EG(s) • Can the cell reach a stable state? init EF(AG(s))not a LTL formula. • Must the cell reach a stable state? init AF(AG(s)) • What are the stable states? Not expressible in CTL [Chan 00]. • Can the system exhibit a cyclic behavior w. r. t. the presence of P ? init EG((P EF P) ^ ( P EF P)) François Fages Evry, April 2004

Biological Queries (3/3) About the correctness of the model : • Can one see the inaccuracies of the model and correct them? Exhibit a counterexample pathway or a witness. Suggest refinements of the model or biological experiments to validate/invalidate the property of the model. About durations : • How long does it take for a molecule to become activated? • In a given time, how many Cyclins A can be accumulated? • What is the duration of a given cell cycle’s phase? CTL operators abstract from durations. Time intervals can be modeled in FO by adding numerical arguments for start times and durations. François Fages Evry, April 2004
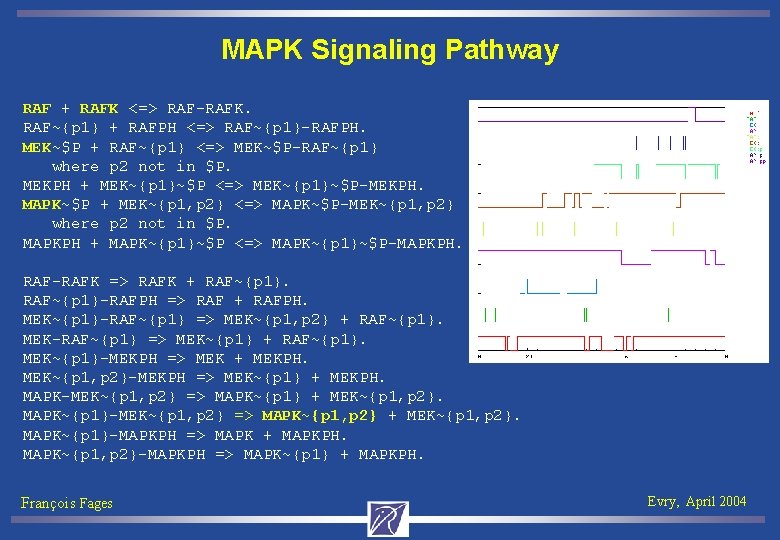
MAPK Signaling Pathway RAF + RAFK <=> RAF-RAFK. RAF~{p 1} + RAFPH <=> RAF~{p 1}-RAFPH. MEK~$P + RAF~{p 1} <=> MEK~$P-RAF~{p 1} where p 2 not in $P. MEKPH + MEK~{p 1}~$P <=> MEK~{p 1}~$P-MEKPH. MAPK~$P + MEK~{p 1, p 2} <=> MAPK~$P-MEK~{p 1, p 2} where p 2 not in $P. MAPKPH + MAPK~{p 1}~$P <=> MAPK~{p 1}~$P-MAPKPH. RAF-RAFK => RAFK + RAF~{p 1}-RAFPH => RAF + RAFPH. MEK~{p 1}-RAF~{p 1} => MEK~{p 1, p 2} + RAF~{p 1}. MEK-RAF~{p 1} => MEK~{p 1} + RAF~{p 1}. MEK~{p 1}-MEKPH => MEK + MEKPH. MEK~{p 1, p 2}-MEKPH => MEK~{p 1} + MEKPH. MAPK-MEK~{p 1, p 2} => MAPK~{p 1} + MEK~{p 1, p 2}. MAPK~{p 1}-MEK~{p 1, p 2} => MAPK~{p 1, p 2} + MEK~{p 1, p 2}. MAPK~{p 1}-MAPKPH => MAPK + MAPKPH. MAPK~{p 1, p 2}-MAPKPH => MAPK~{p 1} + MAPKPH. François Fages Evry, April 2004

MAPK Signaling Pathway MEK~{p 1} is a checkpoint for producing MAPK~{p 1, p 2} biocham: !E(!MEK~{p 1} U MAPK~{p 1, p 2}) True The PH complexes are not compulsory for the cascade biocham: !E(!MEK~{p 1}-MEKPH U MAPK~{p 1, p 2}) false Step 1 rule 15 Step 2 rule 1 RAF-RAFK present Step 3 rule 21 RAF~{p 1} present Step 4 rule 5 MEK-RAF~{p 1} present Step 5 rule 24 MEK~{p 1} present Step 6 rule 7 MEK~{p 1}-RAF~{p 1} present Step 7 rule 23 MEK~{p 1, p 2} present Step 8 rule 13 MAPK-MEK~{p 1, p 2} present Step 9 rule 27 MAPK~{p 1} present Step 10 rule 15 MAPK~{p 1}-MEK~{p 1, p 2} present Step 11 rule 28 MAPK~{p 1, p 2} present François Fages Evry, April 2004
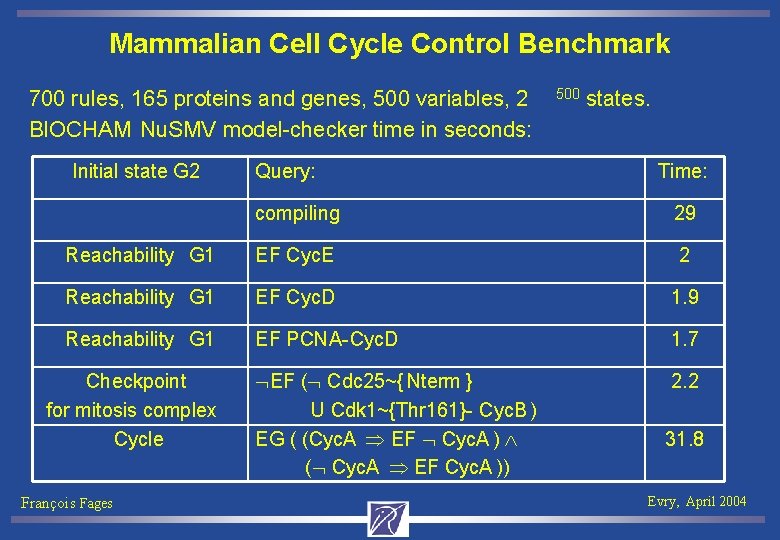
Mammalian Cell Cycle Control Benchmark 700 rules, 165 proteins and genes, 500 variables, 2 BIOCHAM Nu. SMV model-checker time in seconds: Initial state G 2 Query: 500 states. Time: compiling 29 Reachability G 1 EF Cyc. E 2 Reachability G 1 EF Cyc. D 1. 9 Reachability G 1 EF PCNA-Cyc. D 1. 7 EF ( Cdc 25~{ Nterm } U Cdk 1~{Thr 161}- Cyc. B ) EG ( (Cyc. A EF Cyc. A ) ( Cyc. A EF Cyc. A )) 2. 2 Checkpoint for mitosis complex Cycle François Fages 31. 8 Evry, April 2004

Plan of the talk 1. Introduction 2. A simple algebra of cell molecules 3. Concurrent transition systems of biochemical reactions • Example of the mammalian cell cycle control 4. Temporal logic CTL as a query language • Computational results with BIOCHAM 5. Learning reaction rules • An experiment with inductive logic programming 6. Kinetics models • Simulation with differential equations • Hybrid systems 7. Conclusion François Fages Evry, April 2004

5. Learning Reaction Weights and Rules Idea 1: learning reaction weights from temporal properties reaction weights restricts the non-determinism (Markov models) Idea 2: learn reaction rules from temporal properties of the system. Learning of cell cycle reaction rules from reachability properties and counterexamples with Progol [Muggleton 00]. reaction([m_CP, m_Y], [m_p. M]). reaction([m_CP], [m_C 2]). % reaction([m_p. M], [m_M]). reaction([m_M], [m_C 2, m_YP]). reaction([m_C 2], [m_CP]). reaction([m_YP], []). reaction([], [m_Y]). pathway(S 1, S 2) : - same(S 1, S 2). pathway(S 1, S 2) : - reaction(L 1, L 2), transition(S 1, L 1, S 3, L 2), Evry, François Fages pathway(S 3, S 2). April 2004
![Inductive Logic Programming pathway([m_CP, m_Y], [m_M]). pathway([m_CP, m_Y], [m_M, m_p. M]). pathway([m_CP, m_Y], [m_M, Inductive Logic Programming pathway([m_CP, m_Y], [m_M]). pathway([m_CP, m_Y], [m_M, m_p. M]). pathway([m_CP, m_Y], [m_M,](http://slidetodoc.com/presentation_image_h2/607819d83880d1abd758ea35f2143c02/image-32.jpg)
Inductive Logic Programming pathway([m_CP, m_Y], [m_M]). pathway([m_CP, m_Y], [m_M, m_p. M]). pathway([m_CP, m_Y], [m_M, m_Y, m_p. M]). pathway([m_CP, m_Y], [m_M, m_CP, m_Y]). pathway([m_CP, m_Y], [m_M, m_CP, m_p. M]). pathway([m_CP, m_Y], [m_M, m_CP, m_Y, m_p. M ]). pathway([m_p. M], [m_C 2, m_YP]). pathway([m_p. M], [m_M, m_C 2, m_YP]). pathway([m_p. M], [m_p. M, m_C 2, m_YP]). pathway([m_p. M], [m_M, m_p. M, m_C 2, m_YP]). reaction([m_p. M], [m_M]) : -pathway([], [m_C 2]). : -pathway([], [m_CP]). : -pathway([], [m_C 2, m_CP]). : -pathway([], [m_M]). : -pathway([], [m_YP, m_Y]). : -pathway([], [m_Y, m_p. M]). : -pathway([], [m_CP, m_p. M]). : -pathway([], [m_Y, m_M]). : -pathway([m_CP, m_C 2], [m_YP]). : -pathway([m_CP], [m_YP]). : -pathway([m_C 2], [m_YP]). : -pathway([m_Y], []). learned… 6 th PCRD APRIL 2 “Applications of Probabilistic Inductive Logic Progr. ” Luc de Raedt , Univ. Freiburg , Stephen Muggleton , Imperial College London. François Fages Evry, April 2004

Plan of the talk 1. Introduction 2. A simple algebra of cell molecules 3. Concurrent transition systems of biochemical reactions • Example of the mammalian cell cycle control 4. Temporal logic CTL as a query language • Computational results with BIOCHAM 5. Learning reaction rules • An experiment with inductive logic programming 6. Kinetics models • Simulation with differential equations • Hybrid system 7. Conclusion François Fages Evry, April 2004

6. Kinetics Models Enzymatic reactions with rates k 1 k 2 k 3 E+S k 1 C k 2 E+P E+S k 3 C can be compiled by the law of mass action into a system of Michaelis-Menten Ordinary Differential Equations (non-linear) d. E/dt d. S/dt d. C/dt d. P/dt = -k 1 ES+(k 2+k 3)C = -k 1 ES+k 3 C = k 1 ES-(k 2+k 3)C = k 2 C François Fages Evry, April 2004

MAPK kinetics model François Fages Evry, April 2004
![Gene Interaction Networks Gene interaction example [Bockmayr-Courtois 01] Hybrid Concurrent Constraint Programming HCC [Saraswat Gene Interaction Networks Gene interaction example [Bockmayr-Courtois 01] Hybrid Concurrent Constraint Programming HCC [Saraswat](http://slidetodoc.com/presentation_image_h2/607819d83880d1abd758ea35f2143c02/image-36.jpg)
Gene Interaction Networks Gene interaction example [Bockmayr-Courtois 01] Hybrid Concurrent Constraint Programming HCC [Saraswat et al. ] 2 genes x and y. Hybrid linear approximation dx/dt = 0. 01 – 0. 02*x if y < 0. 8 dx/dt = – 0. 02*x if y ≥ 0. 8 dy/dt = 0. 01*x François Fages Evry, April 2004

Concurrent Transition System Time discretization using Euler’s method: y < 0. 8 x’ = x + dt *(0. 01 -0. 02*x) , y’ = y + dt *0. 01*x y ≥ 0. 8 x’ = x + dt *(0. 01 -0. 02*x) , y’ = y + dt *0. 01*x Initial condition: x=0, y=0. Translation into a CLP(R) program ( dt =1) Init : - X=0, Y=0, p(X, Y): -X>=0, Y<0. 8, X 1=X-0. 02*X+0. 01, Y 1=Y+0. 01*X, p(X 1, Y 1). p(X, Y): -X>=0, Y>=0. 8, X 1=X-0. 02*X, Y 1=Y+0. 01*X, p(X 1, Y 1). François Fages Evry, April 2004
![Proving CTL properties by computing fixpoints of Constraint Logic Programs Theorem [Delzanno Podelski 99] Proving CTL properties by computing fixpoints of Constraint Logic Programs Theorem [Delzanno Podelski 99]](http://slidetodoc.com/presentation_image_h2/607819d83880d1abd758ea35f2143c02/image-38.jpg)
Proving CTL properties by computing fixpoints of Constraint Logic Programs Theorem [Delzanno Podelski 99] EF(f)=lfp (TP {p(x): - f} ), EG(f)=gfp (TP f ). Safety property AG( f) iff EF(f) iff init lfp (TP {f} ) Liveness property AG(f 1 AF(f 2)) iff init lfp (TP f 1 gfp(T P {f 2} ) ) Implementation in Sicstus-Prolog CLP(R, B) [ Delzanno 00] François Fages Evry, April 2004

Deductive Model Checker DMC: Gene Interaction r(init, p(s_s, A, B), {A=0, B=0}). r(p(s_s, A, B), p(s_s, C, D), {A>=0, B>=0. 8, C=A-0. 02*A, D=B+0. 01*A}). r(p(s_s, A, B), p(s_s, C, D), {A>=0, B<0. 8, C=A-0. 02*A+0. 01, D=B+0. 01*A}). | ? - prop(P, S). P = unsafe, S =p: s*(x>=0. 6) | ? - ti. Property satisfied. Execution time 0. 0 | ? - ls. s(0, p(s_s, A, _), {A>=0. 6}, 1, (0, 0)). François Fages Evry, April 2004

Demonstration DMC (continued) | ? - prop(P, S). P = unsafe, S =p: s*(x>=0. 2) ? | ? - ti. Property NOT satisfied. Execution time 1. 5 | ? - ls. s(0, p(s_s, A, _), {A>=0. 2}, 1, (0, 0)). s(1, p(s_s, A, B), {B<0. 8, B>=-0. 0, A>=0. 19387755102040816}, 2, (2, 1)). … s(26, p(s_s, A, B), {B>=0. 0, A>=0. 0, B+0. 1982676351105516*A<0. 7741338175552753}, 27, (2, 26)). s(27, init, {}, 28, (1, 27)). François Fages Evry, April 2004

Conclusion The biochemical abstract machine BIOCHAM provides: • A first-order-rule-based language for modeling biochemical systems • A powerful query language based on temporal logic CTL • Models of complex biochemical processes, intracellular and extracellular signaling, cell-cycle control, … a repository of models http: //contraintes. inria. fr/CMBSlib • Implementation in Prolog + model-checkers Nu. SMV and DMC Learning techniques investigated in APr. IL 2 • PILP-based learning of reaction weights from temporal properties • PILP-based learning of reaction rules from temporal properties François Fages Evry, April 2004

Perspectives Collaboration with biologists on BIOCHAM models of the cell-cycle control • Colon cancer therapies, Domenjoud , UHP Nancy • Chronotherapies , Clairambault , INSERM Hybrid concurrent constraint logic programming [Bockmayr Courtois 01, Saraswat 04] Multi-scale molecular-electro-physiological models http: //www-rocq. inria. fr/ http: // www. sci. sdsu. edu /movies François Fages [Sorine et al. 03] sosso/icema 2 Evry, April 2004
- Slides: 42