The Af CS Protein Chemistry Laboratory Deirdre Brekken

The Af. CS Protein Chemistry Laboratory Deirdre Brekken* Hongjun Shu* Farah El-Mazouni* She Chen Robert Cox Kathleen Lyons* Deepa Sethuraman

Goals of the Af. CS Protein Chemistry Laboratory 1. Expand the signaling protein “parts list”, in Af. CS cells, by identifying ligand-stimulated phosphoproteins 2. Expand the repertoire of phosphoproteins monitored during ligand screening by: a. identifying phosphoproteins b. determining if they are regulated by ligands c. identifying the ligand-regulated phosphorylation sites for generation of new phospho-specific antibodies

Current strategy for identifying ligand-regulated phosphoproteins in Af. CS cells 1. Identify phosphoproteins in Af. CS cells (known and unknown proteins) • 2 -D gels (where needed) • Enrichment with phospecfic antibodies (P-Tyr proteins) • Enrichment by immobilized metal affinity chromatography (P-Ser/P-Thr proteins) • Identify phosphoproteins by mass spectrometry 2. Test whether proteins are phosphorylated in a ligand-dependent manner • Express epitope-tagged proteins in Af. CS cells • Label cells with 32 P, stimulate with ligand pools, immunoprecipitate • Resolve immunoprecipitated proteins by SDS gels and quantitate 32 P incorporation (alternatively stain with Pro. Q diamond) 3. Identify phosphorylation sites in ligand-regulated phosphoproteins • Immunoprecipitate tagged proteins from ligand-stimulated cells • Identify phosphorylation sites by precursor ion scanning and tandem mass spectrometry

Current strategy for identifying ligand-regulated phosphoproteins in Af. CS cells 1. Identify phosphoproteins in Af. CS cells (known and unknown proteins) • 2 -D gels (where needed) • Enrichment with phospecfic antibodies (P-Tyr proteins) • Enrichment by immobilized metal affinity chromatography (P-Ser/P-Thr proteins) • Identify phosphoproteins by mass spectrometry 2. Test whether proteins are phosphorylated in a ligand-dependent manner • Express epitope-tagged proteins in Af. CS cells • Label cells with 32 P, stimulate with ligand pools, immunoprecipitate • Resolve immunoprecipitated proteins by SDS gels and quantitate 32 P incorporation (alternatively stain with Pro. Q diamond) 3. Identify phosphorylation sites in ligand-regulated phosphoproteins • Immunoprecipitate tagged protein from ligand-stimulated cells • Identify phosphorylation sites by precursor ion scanning and tandem mass spectrometry
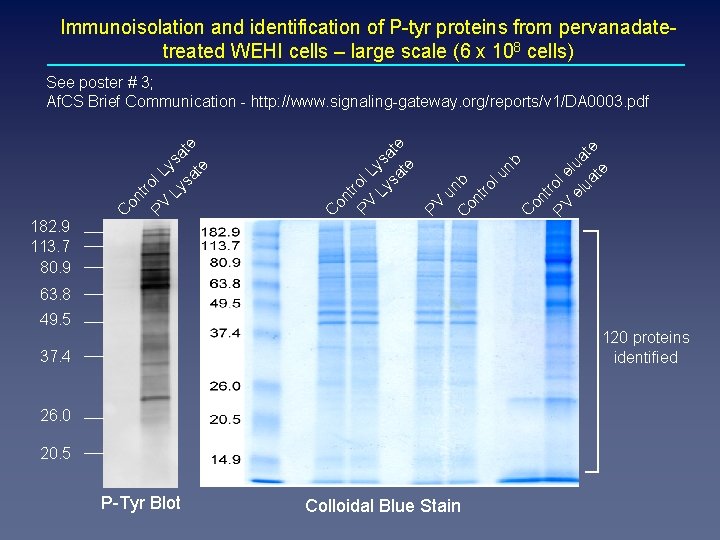
Immunoisolation and identification of P-tyr proteins from pervanadatetreated WEHI cells – large scale (6 x 108 cells) C on t PV rol e el lua ua te te nb PV on t PV rol L Ly ysa sa te te u C nb on tro lu 182. 9 113. 7 80. 9 C C on tr PV ol L Ly ysa sa te te See poster # 3; Af. CS Brief Communication - http: //www. signaling-gateway. org/reports/v 1/DA 0003. pdf 63. 8 49. 5 120 proteins identified 37. 4 26. 0 20. 5 P-Tyr Blot Colloidal Blue Stain
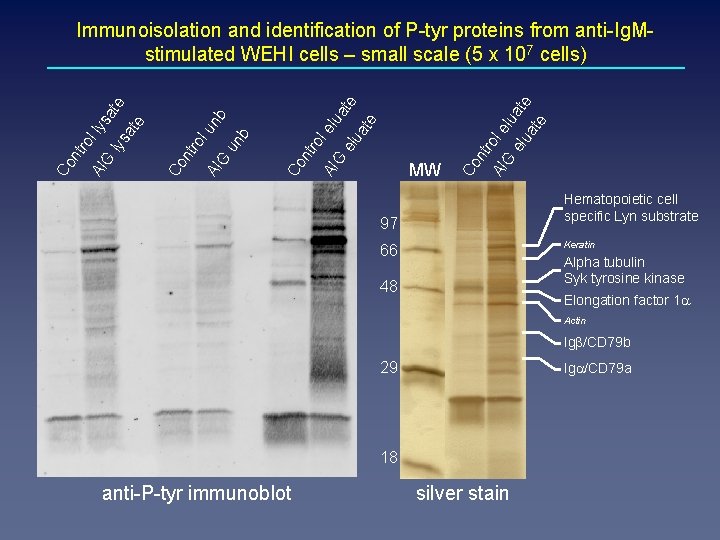
l elu uate at e le at e te G ro AI nt Co ro elu lua G MW AI nt Co AI G un b le nb lu ro nt Co G AI Co nt ro l ly s lys ate at e Immunoisolation and identification of P-tyr proteins from anti-Ig. Mstimulated WEHI cells – small scale (5 x 107 cells) Hematopoietic cell specific Lyn substrate 97 Keratin 66 Alpha tubulin Syk tyrosine kinase 48 Elongation factor 1 a Actin Igb/CD 79 b 29 Iga/CD 79 a 18 anti-P-tyr immunoblot silver stain
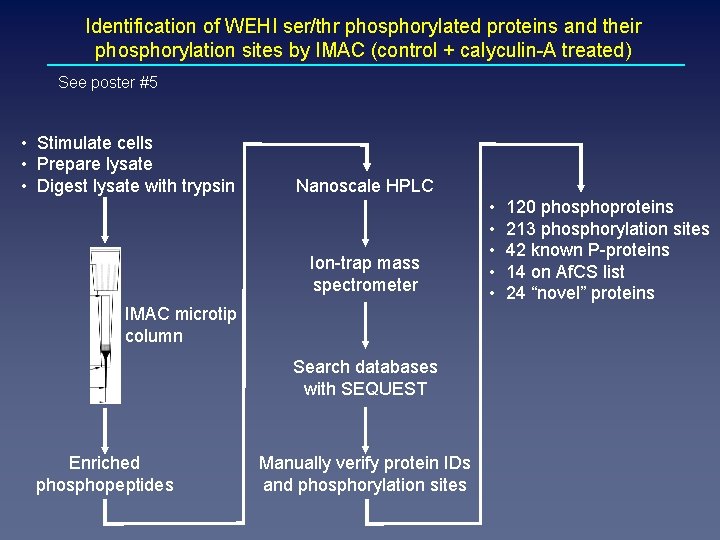
Identification of WEHI ser/thr phosphorylated proteins and their phosphorylation sites by IMAC (control + calyculin-A treated) See poster #5 • Stimulate cells • Prepare lysate • Digest lysate with trypsin Nanoscale HPLC Ion-trap mass spectrometer IMAC microtip column Search databases with SEQUEST Enriched phosphopeptides Manually verify protein IDs and phosphorylation sites • • • 120 phosphoproteins 213 phosphorylation sites 42 known P-proteins 14 on Af. CS list 24 “novel” proteins

Current strategy for identifying ligand-regulated phosphoproteins in Af. CS cells 1. Identify phosphoproteins in Af. CS cells (known and unknown proteins) • 2 -D gels (where needed) • Enrichment with phospecfic antibodies (P-Tyr proteins) • Enrichment by immobilized metal affinity chromatography (P-Ser/P-Thr proteins) • Identify phosphoproteins by mass spectrometry 2. Test whether proteins are phosphorylated in a ligand-dependent manner • Express epitope-tagged proteins in Af. CS cells • Label cells with 32 P, stimulate with ligand pools, immunoprecipitate • Resolve immunoprecipitated proteins by SDS gels and quantitate 32 P incorporation (alternatively stain with Pro. Q diamond) 3. Identify phosphorylation sites in ligand-regulated phosphoproteins • Immunoprecipitate tagged protein from ligand-stimulated cells • Identify phosphorylation sites by precursor ion scanning and tandem mass spectrometry
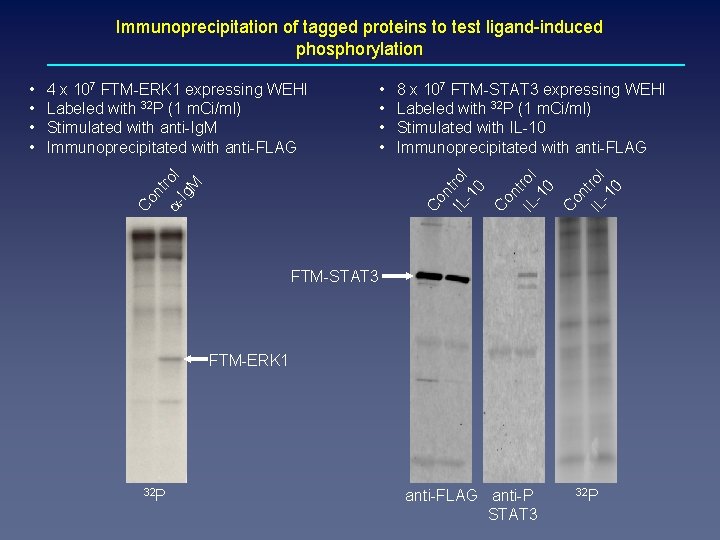
Immunoprecipitation of tagged proteins to test ligand-induced phosphorylation nt a- rol Ig M Co nt on IL trol -1 0 8 x 107 FTM-STAT 3 expressing WEHI Labeled with 32 P (1 m. Ci/ml) Stimulated with IL-10 Immunoprecipitated with anti-FLAG C • • IL rol -1 0 C on t IL rol -1 0 4 x 107 FTM-ERK 1 expressing WEHI Labeled with 32 P (1 m. Ci/ml) Stimulated with anti-Ig. M Immunoprecipitated with anti-FLAG Co • • FTM-STAT 3 FTM-ERK 1 32 P anti-FLAG anti-P STAT 3 32 P
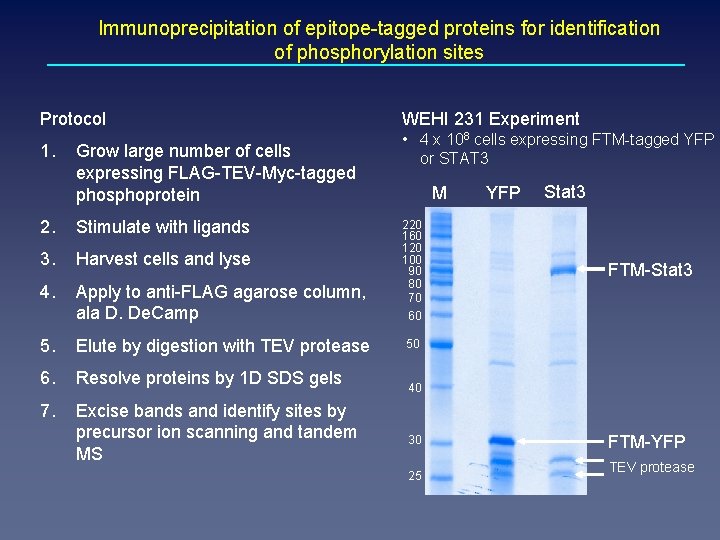
Immunoprecipitation of epitope-tagged proteins for identification of phosphorylation sites Protocol WEHI 231 Experiment 1. • 4 x 108 cells expressing FTM-tagged YFP or STAT 3 Grow large number of cells expressing FLAG-TEV-Myc-tagged phosphoprotein 2. Stimulate with ligands 3. Harvest cells and lyse 4. Apply to anti-FLAG agarose column, ala D. De. Camp 5. Elute by digestion with TEV protease 6. Resolve proteins by 1 D SDS gels 7. Excise bands and identify sites by precursor ion scanning and tandem MS M 220 160 120 100 90 80 70 YFP Stat 3 FTM-Stat 3 60 50 40 30 25 FTM-YFP TEV protease

Current strategy for identifying ligand-regulated phosphoproteins in Af. CS cells 1. Identify phosphoproteins in Af. CS cells (known and unknown proteins) • 2 -D gels (where needed) • Enrichment with phospecfic antibodies (P-Tyr proteins) • Enrichment by immobilized metal affinity chromatography (P-Ser/P-Thr proteins) • Identify phosphoproteins by mass spectrometry 2. Test whether proteins are phosphorylated in a ligand-dependent manner • Express epitope-tagged proteins in Af. CS cells • Label cells with 32 P, stimulate with ligand pools, immunoprecipitate • Resolve immunoprecipitated proteins by SDS gels and quantitate 32 P incorporation (alternatively stain with Pro. Q diamond) 3. Identify phosphorylation sites in ligand-regulated phosphoproteins • Immunoprecipitate tagged proteins from ligand-stimulated cells • Identify phosphorylation sites by precursor ion scanning and tandem mass spectrometry

Identification of phosphorylation sites by precursor ion scanning and tandem mass spectrometry See poster #4 Af. CS Research Report - http: //www. signaling-gateway. org/reports/v 1/DA 0008. htm Protocol 1. For example: 7 sites identified in LSP 1 Isolate proteins A. Spots resolved by 2 -D gels B. Epitope tagged proteins immunoprecipitated from stimulated cells 2. Digest with proteases 3. Detect phosphopeptides by precursor ion scanning using nanospray mass spectrometer 4. Fragment phosphopeptides and detect fragment ions by nanospray tandem MS 5. Compare fragmentation pattern with sequence 6. Use partial sequence information to identify phosphorylation sites

Current strategy for identifying ligand-regulated phosphoproteins in Af. CS cells 1. Identify phosphoproteins in Af. CS cells (known and unknown proteins) • 2 -D gels (where needed) • Enrichment with phospecfic antibodies (P-Tyr proteins) • Enrichment by immobilized metal affinity chromatography (P-Ser/P-Thr proteins) • Identify phosphoproteins by mass spectrometry 2. Test whether proteins are phosphorylated in a ligand-dependent manner • Express epitope-tagged proteins in Af. CS cells • Label cells with 32 P, stimulate with ligand pools, immunoprecipitate • Resolve immunoprecipitated proteins by SDS gels and quantitate 32 P incorporation (alternatively stain with Pro. Q diamond) 3. Identify phosphorylation sites in ligand-regulated phosphoproteins • Immunoprecipitate tagged proteins from ligand-stimulated cells • Identify phosphorylation sites by precursor ion scanning and tandem mass spectrometry
- Slides: 13