The Accelerated Weighted Ensemble Greatly Improved Protein Folding

The Accelerated Weighted Ensemble Greatly Improved Protein Folding Statistics Using Work. Queue and Condor Jeff Kinnison & Dr. Jesus A. Izaguirre
Studying a New Protein HP 24 stab ● Subdomain of the Villin headpiece ● Two-helical supersecondary structure ● 24 amino acids (406 atoms) ● Discovered in 2015, little kinetic information available

Problems with Traditional MD ● Computationally Expensive ○ Molecular force fields perform expensive operations on all atoms ○ Timescales of interest quickly become intractable with protein size ○ GPU resources to increase efficiency are not always readily available ● Events of Interest are Rare ○ Protein folding occurs on O(ns) to O(ms) scale ○ There is no guarantee that a folding event will occur in a given simulation With these two issues, it is difficult to generate enough data to make statistically significant kinetic approximations.
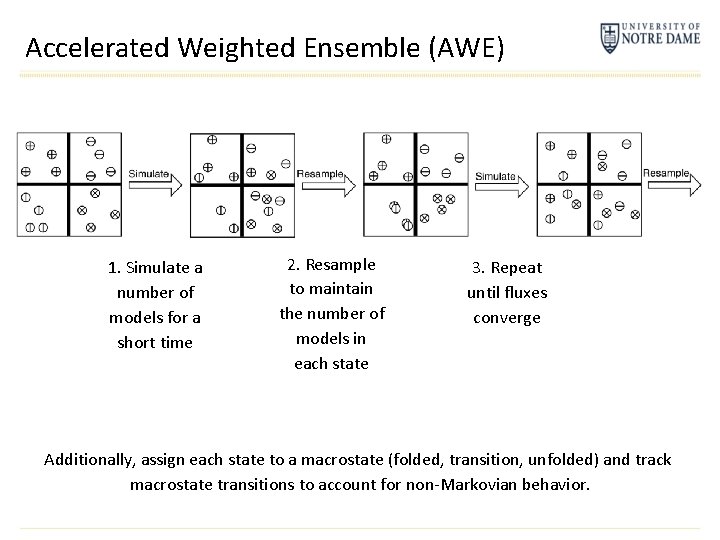
Accelerated Weighted Ensemble (AWE) 1. Simulate a number of models for a short time 2. Resample to maintain the number of models in each state 3. Repeat until fluxes converge Additionally, assign each state to a macrostate (folded, transition, unfolded) and track macrostate transitions to account for non-Markovian behavior.
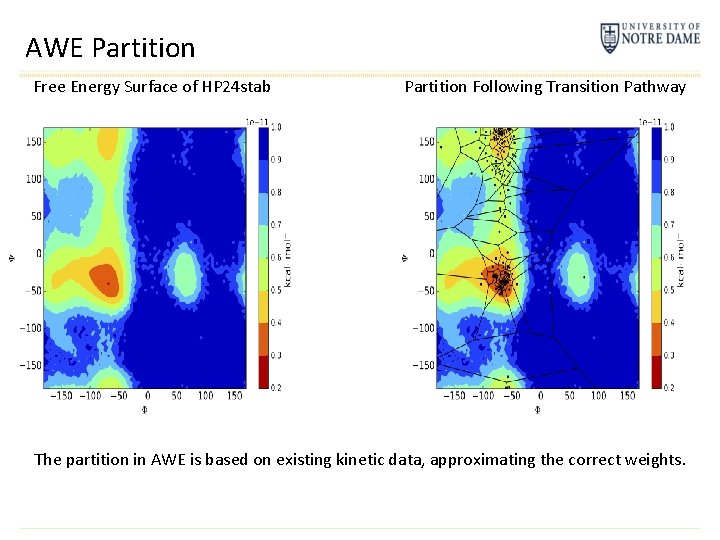
AWE Partition Free Energy Surface of HP 24 stab Partition Following Transition Pathway The partition in AWE is based on existing kinetic data, approximating the correct weights.

Distributing Simulations with Work. Queue • Each simulation is independent, so parallelize simulations to increase efficiency • Work. Queue allows scaling to the number of simulations in a particular AWE run • AWE includes task cloning to overcome bottlenecks caused by slow worker
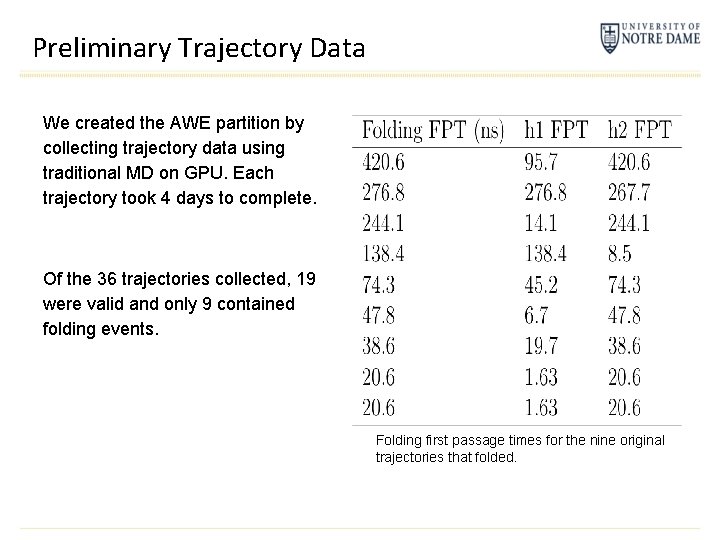
Preliminary Trajectory Data We created the AWE partition by collecting trajectory data using traditional MD on GPU. Each trajectory took 4 days to complete. Of the 36 trajectories collected, 19 were valid and only 9 contained folding events. Folding first passage times for the nine original trajectories that folded.

AWE Setup Two Systems • 1000 -cell • 10 models per state Work. Queue • Maintained a factory requesting between 100 and 1000 workers • All simulations run on 4 -core workers • Used Condor workers only to prevent AWE workers from taking over the cluster MD Parameters • T = 325 K • Langevin Dynamics with implicit solvent (λ =. 91 ps-1) • Amber 03 force field • 250 ps simulation time
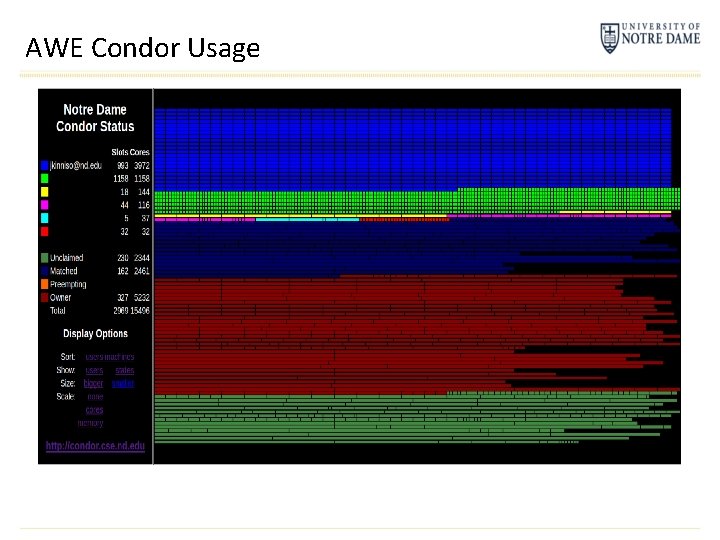
AWE Condor Usage
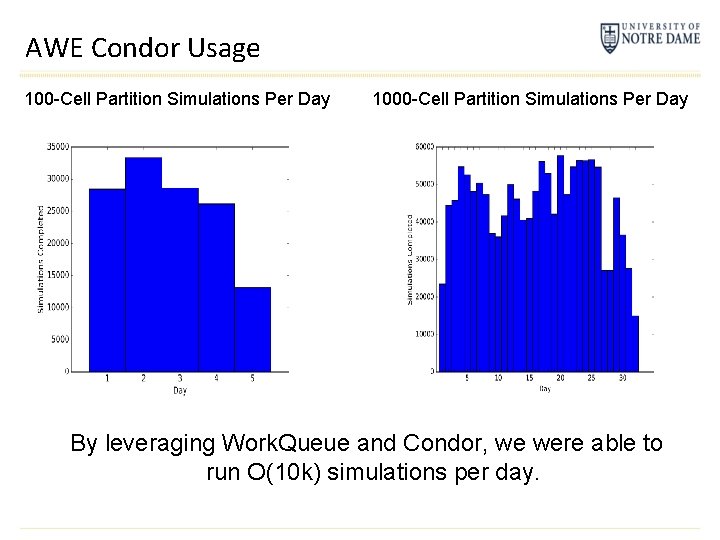
AWE Condor Usage 100 -Cell Partition Simulations Per Day 1000 -Cell Partition Simulations Per Day By leveraging Work. Queue and Condor, we were able to run O(10 k) simulations per day.

AWE Results Started with 19 microseconds of traditional MD trajectory data containing nine folding events computed over one month.

Conclusion Both the coarse and fine partitions converged in one-sixth the time needed to generate the original trajectories and generated several orders of magnitude more folding events. By leveraging Work. Queue and Condor, AWE is able to quickly generate reliable approximations of protein kinetic properties.

Acknowledgements We would like to thank Dr. Douglas Thain and the Cooperative Computing Lab students for making Work. Queue available and helping to integrate it with AWE. All computations were run on compute nodes provided by the Notre Dame Center for Research Computing.

References • Hocking, H. G. ; Häse, F. ; Madl, T. ; Zacharias, M. ; Rief, M. ; Žoldák, G. A Compact Native 24 Residue Supersecondary Structure Derived from the Villin Headpiece Sub- Domain. Biophys. J. 2015, 108, 678– 686. • Huber, G. A. ; Kim, S. Weighted-ensemble Brownian dynamics simulations for protein association reactions. Biophys. J. 1996, 70, 97. • Bhatt, D. ; Zhang, B. W. ; Zuckerman, D. M. Steady-state simulations using weighted ensemble path sampling. J Chem. Phys. 2010, 133, 014110. • Abdul-Wahid, B. ; Yu, L. ; Rajan, D. ; Feng, H. ; Darve, E. ; Thain, D. ; Izaguirre, J. A. Folding Proteins at 500 ns/hour with Work Queue. E-Science (e-Science), 2012 IEEE 8 th International Conference on. 2012; pp 1– 8.
- Slides: 14