TextBased Searching Lesson 3 Bioinformatics Laboratory 1 TextBased

Text-Based Searching Lesson 3 Bioinformatics Laboratory 1 Text-Based Searching 12/6/2020

EMBnet European Molecular Biology Network • In 1988 a network was established to link European laboratories that used bio-computing and bioinformatics in molecular biology research. • In each country a national node provides local biocomputing services • INN serves as Israel’s National Node 3 Text-Based Searching 12/6/2020

Israel National Node • INN serves as Israel’s National Node • Authorized by the Ministry of Science in 1990. • INN is located at the Biological Computing Unit, Weizmann Institute of Science. 4 Text-Based Searching 12/6/2020

Bioinformatics Units at Universities • In Israel, in the mid 1990 s Bioinformatics Units arose at Universities to serve local needs • At TAU – http: //www. tau. ac. il/lifesci/bioinfo 5 Text-Based Searching 12/6/2020

Database Interrogation • Two ways to search databases – Database interrogation – searches textual information contained in header sections of database entries – Database searching – searches sequence information with sequence queries 6 Text-Based Searching 12/6/2020

Database Interrogation Problem of EMBnet • No effective way of interrogating all the resources together at a particular site, since formats differ • A research project was undertaken with EMBnet to address problems inherent in interfacing complex environments – resulting in SRS – sequence retrieval system, a network browser for databases in molecular biology 7 Text-Based Searching 12/6/2020

SRS • SRS allows any flat file database to be indexed to any other. • Powerful tool that allows users to formulate queries across a range of different database types via a single interface, without having to worry about underlying data structures, query languages, etc. Sequence Retrieval System 8 Text-Based Searching 12/6/2020
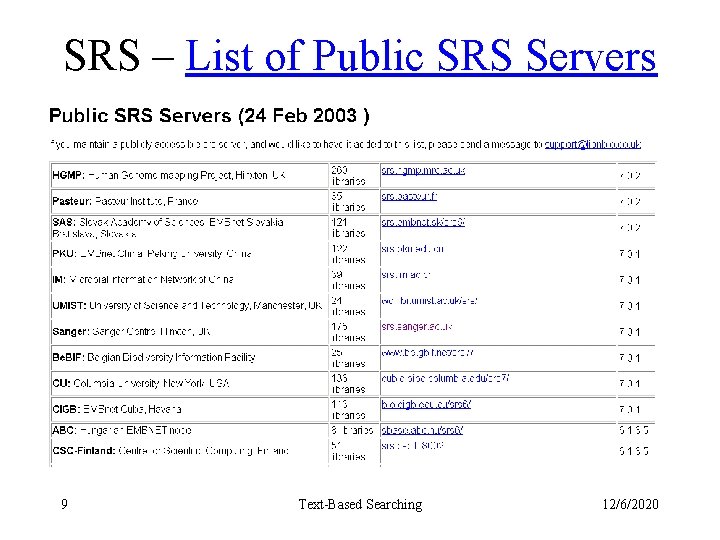
SRS – List of Public SRS Servers 9 Text-Based Searching 12/6/2020
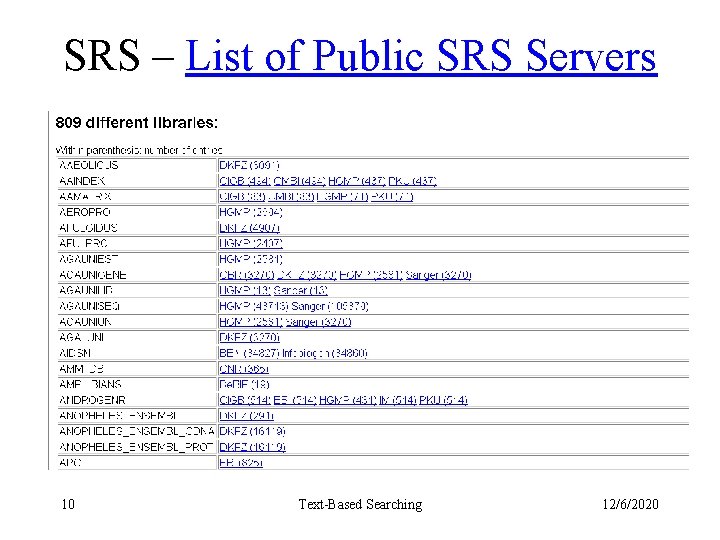
SRS – List of Public SRS Servers 10 Text-Based Searching 12/6/2020

Searching SRS 11 Text-Based Searching 12/6/2020
SRS Tutorial 12 Text-Based Searching 12/6/2020
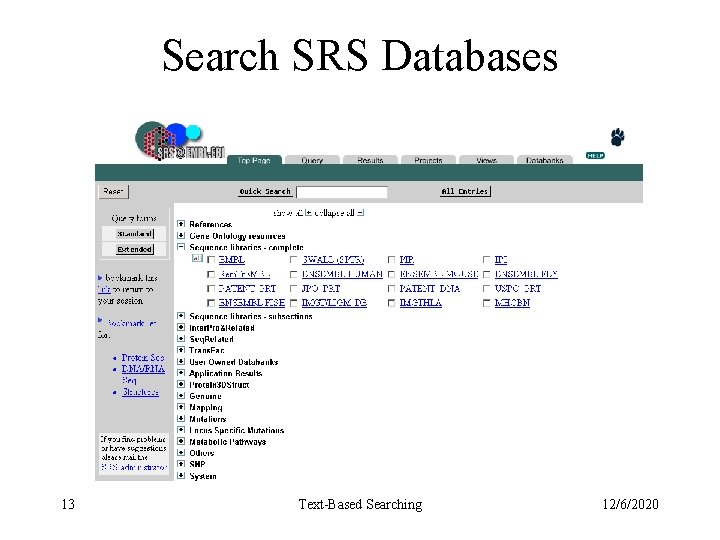
Search SRS Databases 13 Text-Based Searching 12/6/2020
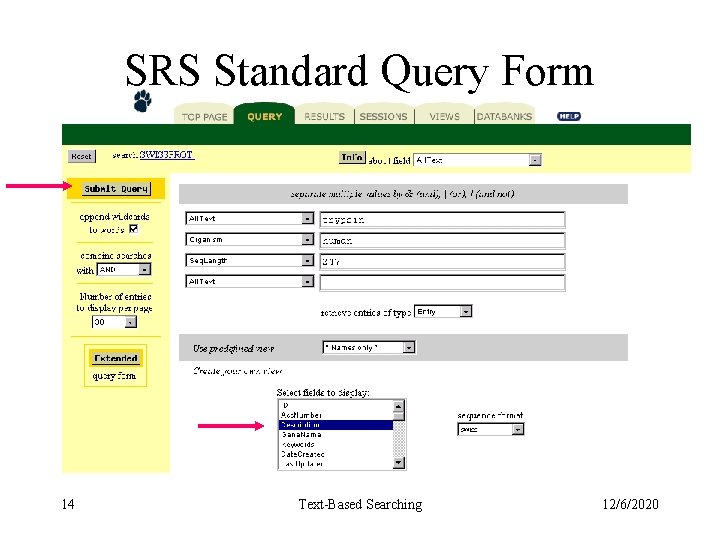
SRS Standard Query Form 14 Text-Based Searching 12/6/2020

SRS Standard Query Form 15 Text-Based Searching 12/6/2020
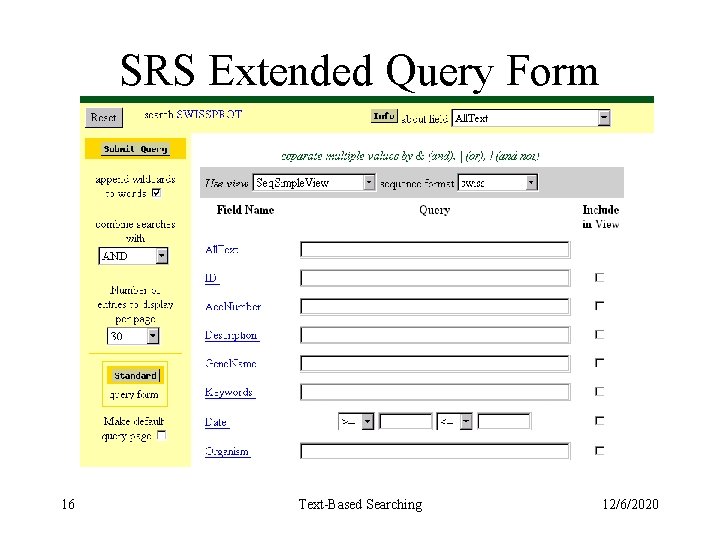
SRS Extended Query Form 16 Text-Based Searching 12/6/2020

NCBI • National Center for Biotechnology • Established in 1988 and located at the campus of NIH as a subdivision of NLM (National Library of Medicine) • Since 1992 one of NCBI’s major tasks has been maintenance of Gen. Bank 17 Text-Based Searching 12/6/2020
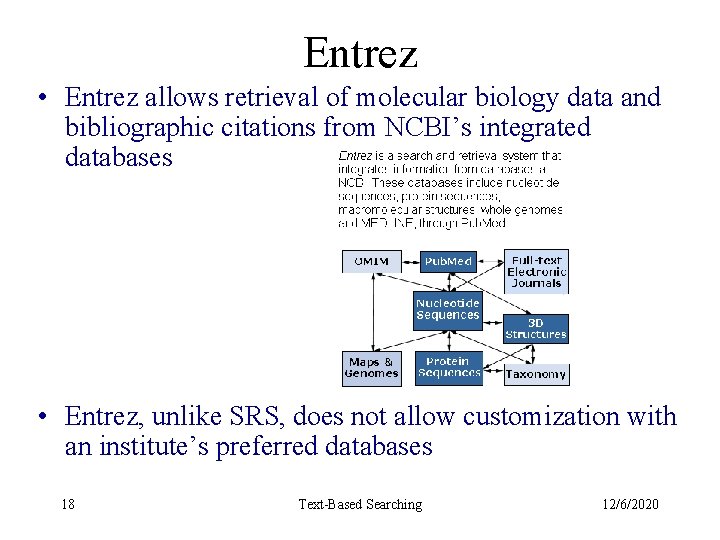
Entrez • Entrez allows retrieval of molecular biology data and bibliographic citations from NCBI’s integrated databases • Entrez, unlike SRS, does not allow customization with an institute’s preferred databases 18 Text-Based Searching 12/6/2020

Entrez • Most records are linked to other records, within a given database and between databases • Sequence databases are linked to the Medline databases so that one can move from paper to sequence and vice versa seamlessly • “Neighboring” allows related papers in Medline, with similar subjects, and sequence entries, found through blast searches, to be grouped together 19 Text-Based Searching 12/6/2020
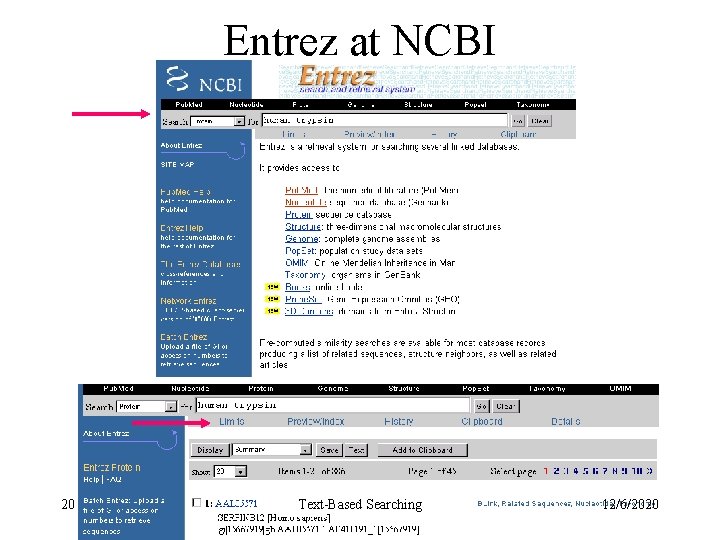
Entrez at NCBI 20 Text-Based Searching 12/6/2020

Entrez at NCBI 21 Text-Based Searching 12/6/2020

Entrez at NCBI 22 Text-Based Searching 12/6/2020

Entrez Pros and Cons • Pros – Integrates reference database with sequence database seamlessly • Cons – Very dependent on the network link as databases being searched are in the US 23 Text-Based Searching 12/6/2020

GCG Software Package • • • Similar syntax to Unix commands Write GCG in every new window to start the program Same principles for all programs: 1. Write command arguments 2. Choose Parameters (default parameters) 3. Receive an output (screen and file) 24 Text-Based Searching 12/6/2020

Searching with GCG • Stringsearch: a simple text-search through local databases. • Searching through definitions or through full annotations. • The definitions contain a minimal amount of the information for each entry: accession, organism name, gene name, sequence length, date. 25 Text-Based Searching 12/6/2020

Searching with GCG • The annotations contain the complete documentation for each entry in the sequence database, including journal and author names, sequence features, comments, etc. Annotations take much longer to search through 26 Text-Based Searching 12/6/2020

Getting a sequence • Fetch: Get a sequence file to your account using the accession number or the id code. Example: fetch hum_hbb • Fetches all the files with the given accession number. Can be limited to a certain data base using database code: • Example: fetch embl: u 01613 27 Text-Based Searching 12/6/2020

Sequence formats • Different applications use different sequence format. • GCG • FASTA/Pearson 28 Text-Based Searching 12/6/2020

Changing file formats • Two GCG commands are used to convert file format. • tofasta • formfasta • Similar commands (fromembl, topir etc) 29 Text-Based Searching 12/6/2020
- Slides: 28