Text Mining for Biomedicine Techniques tools Sophia Ananiadou

Text Mining for Biomedicine: Techniques & tools Sophia Ananiadou, Chikashi Nobata, Yutaka Sasaki, Yoshimasa Tsuruoka School of Computer Science National Centre for Text Mining www. nactem. ac. uk Sophia. Ananiadou@manchester. ac. uk

Outline • Challenges / objectives of TM in biomedicine • Terminology processing – Term extraction, term variation, named entity recognition • • • Resources for TM in biomedicine Document classification Information Extraction approaches Levels of Text Mining Processing Biomedical text mining services and systems @ Na. CTe. M – Ter. Mine, Acro. Mine, Smart dictionary look up, Phenetica – Medie, Info. Pub. Med, KLEIO 2

Material • Further background on TM for Biology Ananiadou, S. & Mc. Naught, J. (eds) (2006) Text Mining for Biology and Biomedicine. Boston, MA: Artech House • Numerous papers on line from bibliography • See BLIMP http: //blimp. cs. queensu. ca/ – Biomedical Literature (and text) mining publications 3

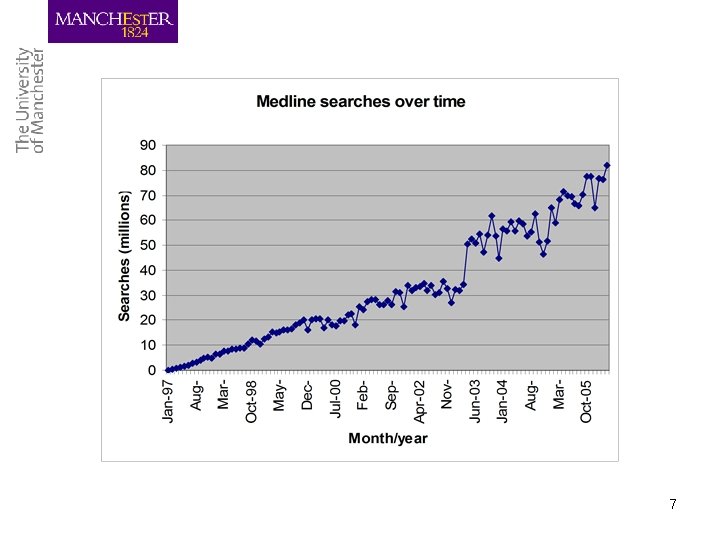
Text Mining in biomedicine • Why biomedicine? – Consider just MEDLINE: 16, 000 references, 40, 000 added per month – Dynamic nature of the domain: new terms (genes, proteins, chemical compounds, drugs) constantly created – Impossible to manage such an information overload 4
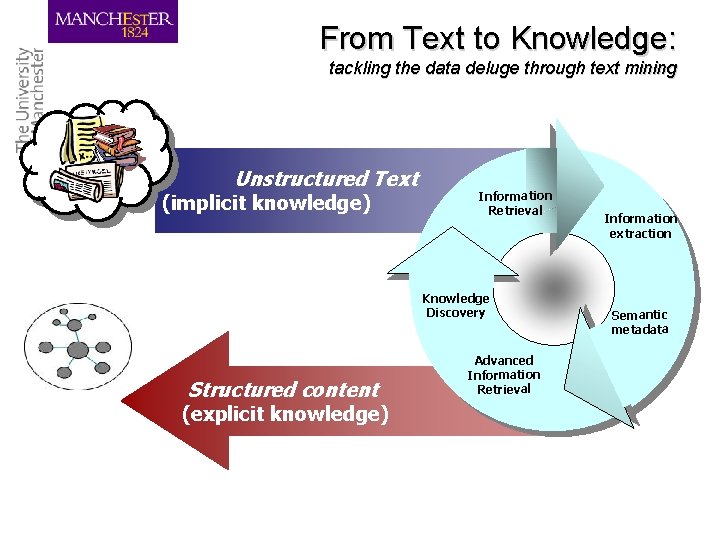
From Text to Knowledge: tackling the data deluge through text mining Unstructured Text (implicit knowledge) Information Retrieval Knowledge Discovery Structured content (explicit knowledge) Advanced Information Retrieval Information extraction Semantic metadata

Information deluge • Bio-databases, controlled vocabularies and bioontologies encode only small fraction of information • Linking text to databases and ontologies – Curators struggling to process scientific literature – Discovery of facts and events crucial for gaining insights in biosciences: need for text mining 6
7

The solution: The UK National Centre for Text Mining www. nactem. ac. uk • Location: Manchester Interdisciplinary Biocentre (MIB) www. mib. ac. uk • First publicly funded text mining centre in the world. . • Focus: biology, medicine, social sciences… 8

We don’t just press a button… • TM involves – Many components (converters, analysers, miners, visualisers, . . . ) – Many resources (grammars, ontologies, lexicons, terminologies, thesauri, CVs) – Many combinations of components and resources for different applications – Many different user requirements and scenarios, training needs • The best solutions are customised 9

People behind Na. CTe. M • Text Mining Team: 14 members • Close collaboration with University of Tokyo, Tsujii Lab http: //www-tsujii. is. s. u-tokyo. ac. jp/ 10

What Na. CTe. M is building: • Resources: ontologies, lexicons, terminologies, thesauri, grammars, annotated corpora – BOOTStrep project http: //www. nactem. ac. uk/bootstrep. php • Tools: tokenisers, taggers, chunkers, parsers, NE recognisers, semantic analysers • Na. CTe. M is also providing services • Our related bio-text mining projects – REFINE http: //dbkgroup. org/refine/ – Representing Evidence For Interacting Network Elements – ONDEX (data integration, workflows, text mining) 11

Individual tools for user data • Splitters, taggers, chunkers, parsers, NER, term extractors • Modes of use Demonstrators: for small-scale online use Batch mode: upload data, get email with link to download site when job done Web Services Integration into Workflows (Taverna) • Some services are compositions of tools 12

Aims • Text mining: discover & extract unstructured knowledge hidden in text – Hearst (1999) • Text mining aids to construct hypotheses from associations derived from text – protein-protein interactions – associations of genes – phenotypes – functional relationships among genes 13

Impact of text mining • Extraction of named entities (genes, proteins, metabolites, etc) • Discovery of concepts allows semantic annotation of documents – Improves information access by going beyond index terms, enabling semantic querying • Construction of concept networks from text – Allows clustering, classification of documents – Visualisation of concept maps 14

Impact of TM • Extraction of relationships (events and facts) for knowledge discovery – Information extraction, more sophisticated annotation of texts (event annotation) – Beyond named entities: facts, events – Enables even more advanced semantic querying 15

Hypothesis generation from literature • Swanson experiments (1986) influenced conceptual biology – rapid ‘mining’ of candidate hypotheses from the literature – migraine and magnesium deficiency (Swanson, 1988) – indomethacin and Alzheimer’s disease (Swanson and Smalheiser 1994), – Curcuma longa and retinal diseases, Crohn's disease and disorders related to the spinal cord (Srinivasan and Libbus 2004). – (Weeber M, Rein et al. 2003) thalidomide for treating a series of diseases such as acute pancreatitis, chronic hepatitis C. 16

Text mining steps • Information Retrieval yields all relevant texts – Gathers, selects, filters documents that may prove useful – Finds what is known • Information Extraction extracts facts & events of interest to user – Finds relevant concepts, facts about concepts – Finds only what we are looking for • Data Mining discovers unsuspected associations – Combines & links facts and events – Discovers new knowledge, finds new associations 17
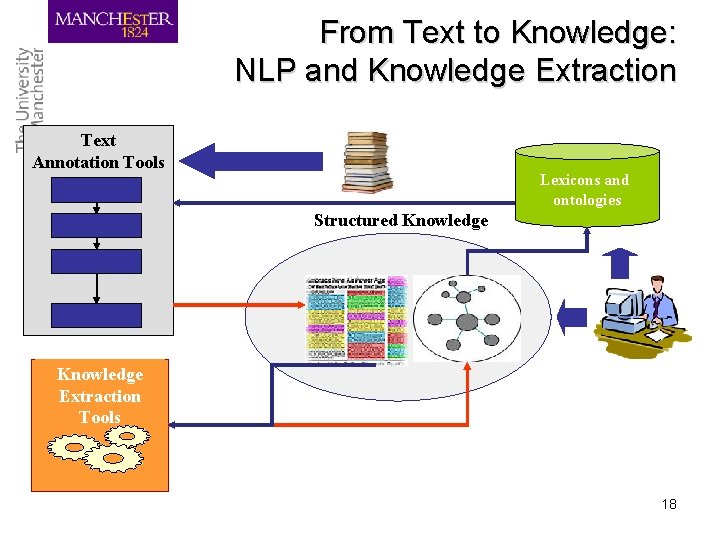
From Text to Knowledge: NLP and Knowledge Extraction Text Annotation Tools Lexicons and ontologies Structured Knowledge Extraction Tools 18

Challenge: the resource bottleneck • Lack of large-scale, richly annotated corpora – Support training of ML algorithms – Development of computational grammars – Evaluation of text mining components • Lack of knowledge resources: lexica, terminologies, ontologies. 19
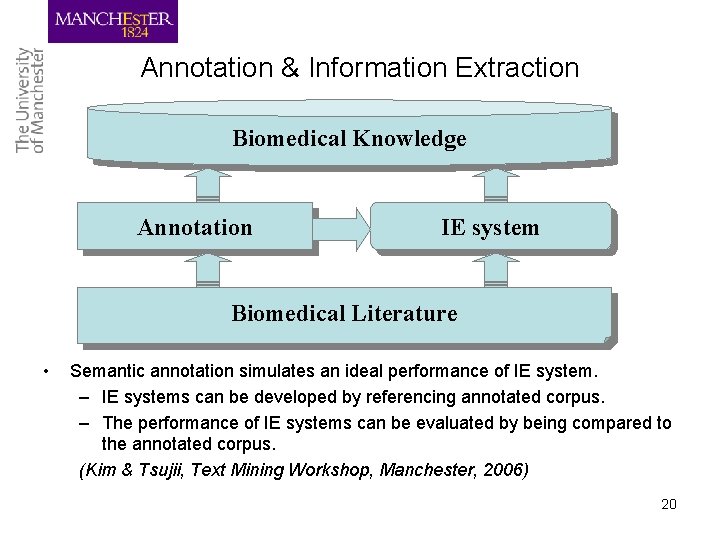
Annotation & Information Extraction Biomedical Knowledge Annotation IE system Biomedical Literature • Semantic annotation simulates an ideal performance of IE system. – IE systems can be developed by referencing annotated corpus. – The performance of IE systems can be evaluated by being compared to the annotated corpus. (Kim & Tsujii, Text Mining Workshop, Manchester, 2006) 20

Text Annotation • • Task-oriented Annotation Task-neutral Annotation – Application annotated text – – User system development – Development of generic tools – Defined by specific tasks – Defined by theories • • Interoperable Tools Specific curation tasks in specific environments Mapping of Protein names to database IDs in specific text types Specific event types such as Protein Interaction Disease-Gene Association of specific diseases GENIA Corpus [U-Tokyo, Na. CTe. M] • • • Linguistics – Tokens – POS – Phrase Structure – Dependency Structure – Deep Syntax (PAS) Biology – Named Entities of various semantic types – Events Linguistics + Biology – Co-references 21

Annotation of GENIA corpus – Term&POS Part-of-speech annotation 2, 000 abstracts Term (entity) annotation 2000+400 abstracts 22

Text semantic annotation • annotation of events and involved named entities – Example: “Regulation of Transcription events” – BOOTSTrep project http: //www. nactem. ac. uk/bootstrep. php • two different types of annotation levels • linguistic annotation levels • biological annotation level, in charge of marking the biological knowledge contained in the text • Linking text with biological knowledge 23

Events and variables • Biological events can be centred on: – verbs, e. g. activate, – nouns with verb-like meanings (nominalised verbs), e. g. transcription • Different parts of sentence correspond to different types of variables in the event e. g. – What caused event • The nar. L gene product activates the nitrate reductase operon – What was affected by event • Analysis of mutants … – Where event took place • These fusions were formed on plasmid cloning vectors
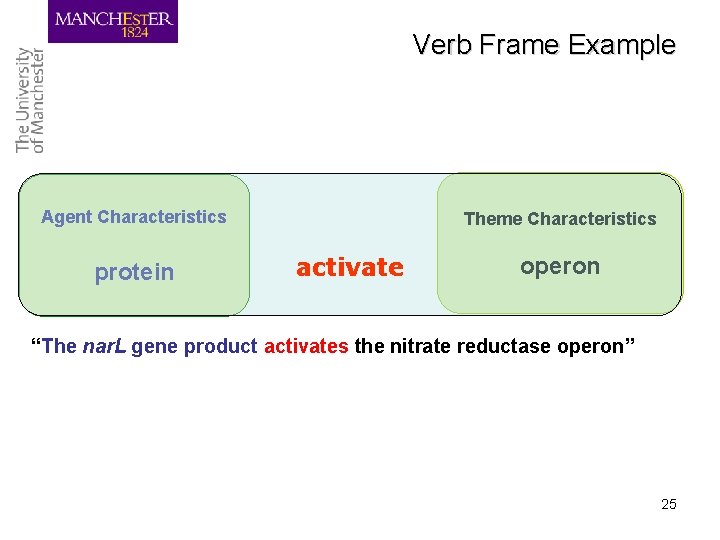
Verb Frame Example Agent Characteristics protein Theme Characteristics activate operon “The nar. L gene product activates the nitrate reductase operon” 25
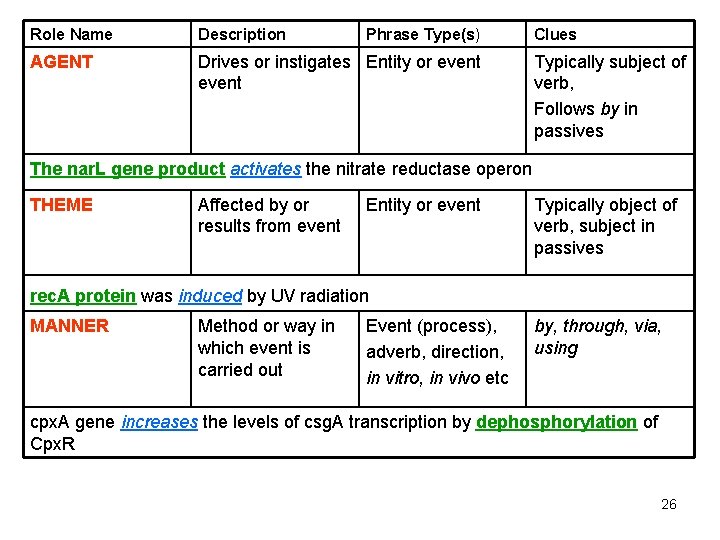
Role Name Description Phrase Type(s) AGENT Drives or instigates Entity or event Clues Typically subject of verb, Follows by in passives The nar. L gene product activates the nitrate reductase operon THEME Affected by or results from event Entity or event Typically object of verb, subject in passives rec. A protein was induced by UV radiation MANNER Method or way in which event is carried out Event (process), adverb, direction, in vitro, in vivo etc by, through, via, using cpx. A gene increases the levels of csg. A transcription by dephosphorylation of Cpx. R 26
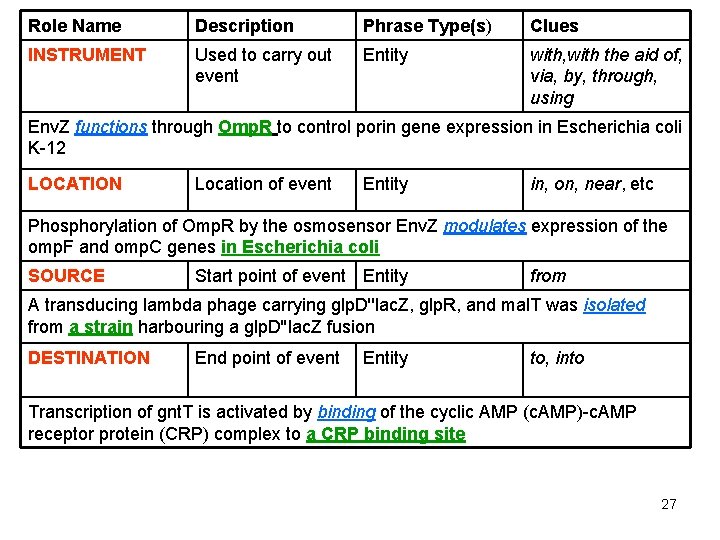
Role Name Description Phrase Type(s) Clues INSTRUMENT Used to carry out event Entity with, with the aid of, via, by, through, using Env. Z functions through Omp. R to control porin gene expression in Escherichia coli K-12 LOCATION Location of event Entity in, on, near, etc Phosphorylation of Omp. R by the osmosensor Env. Z modulates expression of the omp. F and omp. C genes in Escherichia coli SOURCE Start point of event Entity from A transducing lambda phage carrying glp. D''lac. Z, glp. R, and mal. T was isolated from a strain harbouring a glp. D''lac. Z fusion DESTINATION End point of event Entity to, into Transcription of gnt. T is activated by binding of the cyclic AMP (c. AMP)-c. AMP receptor protein (CRP) complex to a CRP binding site 27
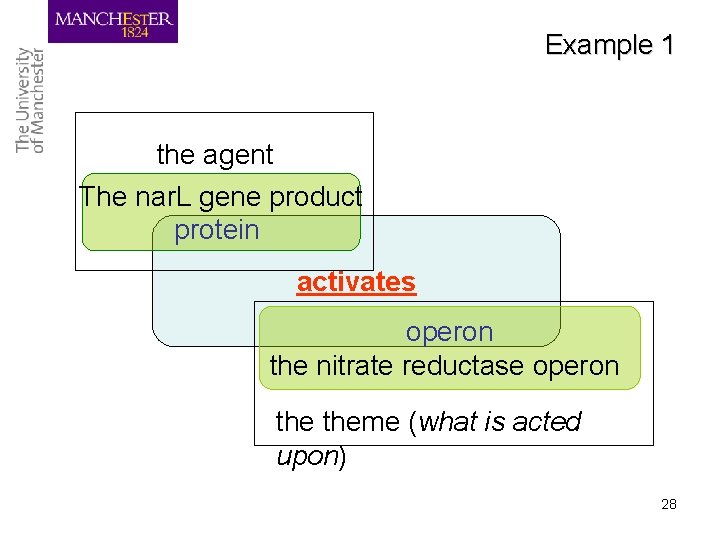
Example 1 the agent The nar. L gene product protein activates operon the nitrate reductase operon theme (what is acted upon) 28

Linguistically Annotated Corpora • GENIA – Domain • Mesh term: Human, Blood Cells, and Transcription Factors. – Annotation: POS, named entity, parse tree • Penn Bio. IE – Domain • the molecular genetics of oncology • the inhibition of enzymes of the CYP 450 class. – Annotation: POS, named entity, parse tree • Yapex • GENETAG a corpus of 20 K MEDLINE® sentences for gene/protein NER 29

The GENIA annotation • Linguistic annotation – Reveals linguistic structures behind the text • Part-of-speech annotation – annotates for the syntactic category of each word. • Syntactic Tree annotation – annotates for the syntactic structure of sentences. • Semantic annotation – Reveals knowledge pieces delivered by the text. • Term annotation – annotates domain-specific terms • Event annotation – annotates events on biological entities. Ontology-driven annotation 30

Annotation Tool • Word. Freak http: //wordfreak. sourceforge. net/ • Java-based linguistic annotation tool developed at University of Pennsylvania • Extensible to new tasks and domains • Customised visualisation and annotation specification – Allows annotation process to be made as simple as possible 31

Resources 32

What about existing resources? • Ontologies important for knowledge discovery – They form the link between terms in texts and biological databases – Can be used to add meaning, semantic annotation of texts 33

Link between text and ontologies Adding new knowledge UMLS KEGG Ontological resources GO GENIA text Supporting semantics 34
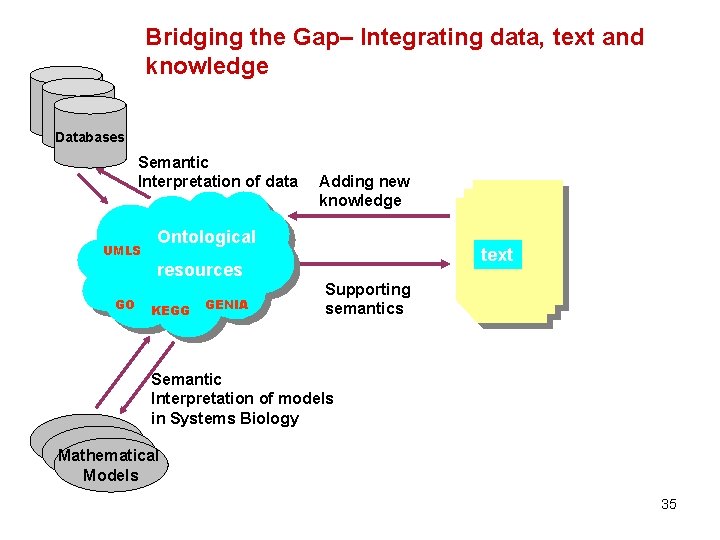
Bridging the Gap– Integrating data, text and knowledge Databases Semantic Interpretation of data UMLS Adding new knowledge Ontological text resources GO KEGG GENIA Supporting semantics Semantic Interpretation of models in Systems Biology Mathematical Models 35

Resources for Bio-Text Mining • Lexical / terminological resources – SPECIALIST lexicon, Metathesaurus (UMLS) – Lists of terms / lexical entries (hierarchical relations) • Ontological resources – Metathesaurus, Semantic Network, GO, SNOMED CT, etc – Encode relations among entities Bodenreider, O. “Lexical, Terminological, and Ontological Resources for Biological Text Mining”, Chapter 3, Text Mining for Biology and Biomedicine, pp. 43 -66 36

SPECIALIST lexicon – UMLS specialist lexicon http: //SPECIALIST. nlm. nih. gov • Each lexical entry contains morphological (e. g. cauterize, cauterizes, cauterized, cauterizing), syntactic (e. g. complementation patterns for verbs, nouns, adjectives), orthographic information (e. g. esophagus – oesophagus) • General language lexicon with many biomedical terms (over 180, 000 records) • Lexical programs include variation (spelling), base form, inflection, acronyms 37

Lexicon record {base=Kaposi's sarcoma spelling_variant=Kaposi sarcoma entry=E 0003576 cat=noun variants=uncount variants=reg variants=glreg} Kaposi’s sarcomas Kaposi’s sarcomata Kaposi sarcomas Kaposi sarcomata The SPECIALIST Lexicon and Lexical Tools Allen C. Browne, Guy Divita, and Chris Lu Ph. D 2002 NLM Associates Presentation, 12/03/2002, Bethesda, MD 38

Normalisation (lexical tools) Hodgkin Disease HODGKIN DISEASE Hodgkin’s Disease Hodgkin’s disease Disease, Hodgkin. . . disease hodgkin normalise 39

Steps of Norm Remove genitive Hodgkin’s Diseases Replace punctuation with spaces Hodgkin Diseases Remove stop words Hodgkin Diseases Lowercase hodgkin diseases Uninflect each word hodgkin disease Word order sort disease hodgkin Lexical tools of the UMLS http: //lexsrv 3. nlm. nih. gov/SPECIALIST/index. html 40

The Gene Ontology (GO) • Controlled vocabulary for the annotation of gene products http: //www. geneontology. org/ 19, 468 terms. 95. 3% with definitions 10391 biological_process 1681 cellular_component 7396 molecular_function 41

Gene Ontology • GOA database (http: //www. ebi. ac. uk/GOA/) assigns gene products to the Gene Ontology • GO terms follow certain conventions of creation, have synonyms such as: – ornithine cycle is an exact synonym of urea cycle – cell division is a broad synonym of cytokinesis – cytochrome bc 1 complex is a related synonym of ubiquinol-cytochrome-c reductase activity 42

GO terms, definitions and ontologies in OBO id: GO: 0000002 name: mitochondrial genome maintenance namespace: biological_process def: "The maintenance of the structure and integrity of the mitochondrial genome. “ [GOC: ai] is_a: GO: 0007005 ! mitochondrion organization and biogenesis 43

Metathesaurus • organised by concept – 5 M names, 1 M concepts, 16 M relations • built from 134 electronic versions of many different thesauri, classifications, code sets, and lists of controlled terms • "source vocabularies“ • common representation 44

Are the existing knowledge resources sufficient for TM? No! Why? Limited lexical & terminological coverage of biological sub-domains Resources focused on human specialists GO, UMLS, Uni. Prot ontology concept names frequently confused with terms 45

Naming conventions 3. Update and curation of resources – Fly. Base gene name coverage 31% (abstracts) to 84% (full texts) 4. Naming conventions and representation in heterogeneous resources – Term formation guidelines from formal bodies e. g. HUGO, IPI not uniformly used – Problems with integration of resources dystrophin used for 18 gene products “Dystrophin (muscular dystrophy, Duchenne and Becker types), included DXS 143, DXS 164, DXS 206, …” HUGO 46

Term variation 5. Terminological variation and complexity of names – High correlation between degree of term variation and dynamic nature of biomedicine – Variation occurs in controlled vocabularies and texts but discrepancy between the two – Exact match methods fail to associate term occurrences in texts with databases 47

What’s in a name? Terms, named entities in biology 48

What’s in a name? • • • Breast cancer 1 (BRCA 1) p 53 Ribosomal protein S 27 Heat shock protein 110 Mitogen activated protein kinase 15 Mitogen activated protein kinase 5 From K. Cohen, NAACL 2007 49

Worst gene names • sema domain, seven thrombospondin repeats (type 1 and type 1 -like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5 A K. Cohen NAACL 2007 50

Worst gene names • sema domain, seven thrombospondin repeats (type 1 and type 1 -like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5 A K. Cohen NAACL 2007 51

Worst gene names • sema domain, seven thrombospondin repeats (type 1 and type 1 -like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5 A • SEMA 5 A K. Cohen NAACL 2007 52

Worst gene names • sema domain, seven thrombospondin repeats (type 1 and type 1 -like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5 A • SEMA 5 A • Tyrosine kinase with immunoglobulin and epidermal growth factor homology domains • tie K. Cohen NAACL 2007 53
![Term ambiguity Neurofibromatosis 2 [disease] NF 2 Neurofibromin 2 [protein] Neurofibromatosis 2 gene [gene] Term ambiguity Neurofibromatosis 2 [disease] NF 2 Neurofibromin 2 [protein] Neurofibromatosis 2 gene [gene]](http://slidetodoc.com/presentation_image_h/988e5bda448f6ad5971b315f2bf4b955/image-54.jpg)
Term ambiguity Neurofibromatosis 2 [disease] NF 2 Neurofibromin 2 [protein] Neurofibromatosis 2 gene [gene] O. Bodenreider, MIE 2005 tutorial http: //www. nactem. ac. uk/ 54

Term ambiguity – Gene terms may be also common English words • BAD human gene encoding BCL-2 family of proteins (bad news, bad prediction) – Gene names are often used to denote gene products (proteins) • suppressor of sable is used ambiguously to refer to either genes and proteins – Existing resources lack information that can support term disambiguation – Difficult to establish equivalences between termforms and concepts 55

Homologues • Cycline-dependent kinase inhibitor first introduced to represent a protein family p 27 – But it is used interchangeably with p 27 or p 27 kip 1, as the name of the individual protein and not as the name of the protein family (Morgan 2003). • NFKB 2 denotes the name of a family of 2 individual proteins with separate IDs in Swiss. Prot. – These proteins are homologues belonging to different species, homo sapiens & chicken. 56

Terms – Term: linguistic realisation of specialised concepts, e. g. genes, proteins, diseases – Terminology: collection of terms structured (hierarchy) denoting relationships among concepts, part-whole, is -a, specific, generic, etc. – Terms link text and ontologies – Mapping is not trivial (main challenge) 57

Term variation and ambiguity Term variation Term 1 Term 2 Term 3 TEXT Term ambiguity Concept 1 concept 3 concept 2 ONTOLOGY 58
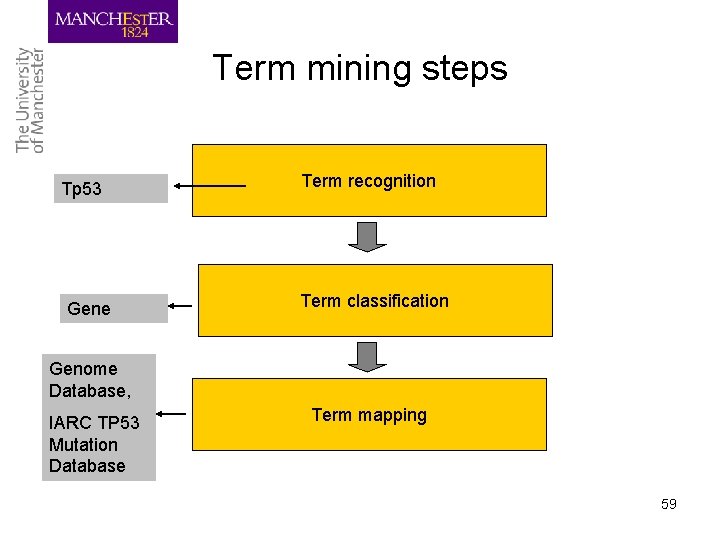
Term mining steps Tp 53 Gene Term recognition Term classification Genome Database, IARC TP 53 Mutation Database Term mapping 59

Term recognition techniques • ATR extracts terms (variants) from a collection of document • Distinguishes terms vs non-terms • In NER the steps of recognition and classification are merged, a classified terminological instance is a named entity • The tasks of ATR and NER share techniques but their ultimate goals are different – ATR for resource building, lexica & ontologies – NER first step of IE, text mining 60

Overview papers 1. S. Ananiadou & G. Nenadic (2006) Automatic Terminology Management in Biomedicine, Text Mining for Biology and Biomedicine, pp. 67 - 97. 2. M. Krauthammer & G. Nenadic (2004) Term identification in the biomedical literature, JBI 37 (2004) 512 -526 3. J. C. Park & J. Kim (2006) Named Entity Recognition, Text Mining for Biology and Biomedicine, pp. 121 -142 Detailed bibliography in Bio-Text Mining 1. BLIMPhttp: //blimp. cs. queensu. ca/ 2. http: //www. ccs. neu. edu/home/futrelle/bionlp/ Book on Bio. Text Mining 1. S. Ananiadou & J. Mc. Naught (eds) (2006) Text Mining for Biology and Biomedicine, Artech House. Other Bio-Text Mining tutorials Kevin Cohen (NAACL 2007 tutorial) U. Colorado 61

Main ATR approaches ATR Dictionary based Rule based Machine learning 62

Dictionary NER (1) • Use terminological resources to locate term occurrences in text – NCBI http: //www. ncbi. nlm. nih. gov/ – EBI http: //www. ebi. ac. uk/ – neologisms, variations, ambiguity problematic for simple dictionary look-up – Ambiguous words e. g. an, for, can … – spelling variants, punctuation, word order variations • estrogen oestrogen • NF kappa B / NF k. B 63

Dictionary NER (2) – Hirschman (2002) used Fly. Base for gene name recognition, results disappointing due to homonymy, spelling variations • Precision, 7% abstracts, 2% full papers • Recall, 31% -- 84% – Tuason (2004) reports term variation as main problem of mismatch • bmp-4 bmp 4 • syt 4 syt iv • integrin alpha 4 integrin 64

Dictionary NER (3) – Tsuruoka & Tsujii (2003) suggest a probabilistic generator of spelling variants, edit distance operations (delete, substitute, insert) • Terms with ED ≤ 1 considered spelling variants • Used a dictionary of protein terms – Support query expansion – Augment dictionaries with variation 65
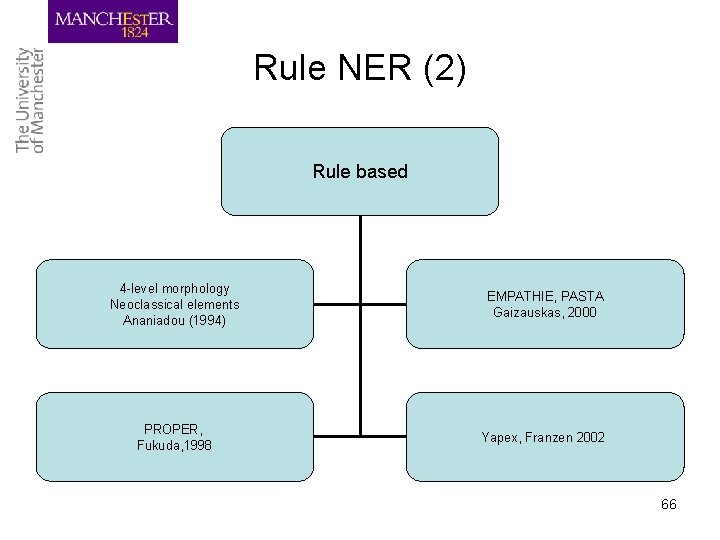
Rule NER (2) Rule based 4 -level morphology Neoclassical elements Ananiadou (1994) EMPATHIE, PASTA Gaizauskas, 2000 PROPER, Fukuda, 1998 Yapex, Franzen 2002 66

Rule based (1) • Use orthographic, morpho-syntactic features of terms – Rules that make use of internal term formation patterns (tagging, morphological analysers) e. g. affixes, combining forms – Do not take into account contextual features – Dictionaries of constituents e. g. affixes, neoclassical forms included • Portability to different domains? 67

Rule based (2) • Ananiadou, S. (1994) recognised single-word terms based on morphological analysis of term formation patterns (internal term make up) • based on analysis of neoclassical and hybrid elements ‘alphafetoprotein’ ‘immunoosmoelectrophoresis’ ‘radioimmunoassay’ • some elements are used for creating terms term word + term_suffix term + word_suffix • neoclassical combining forms (electro- adeno-), • prefixes (auto-, hypo-) • suffixes ( -osis, -itis) 68

Rule-based (3) • Fukuda (1998) used lexical, orthographic features for protein name recognition e. g. upper case character, numerals etc. • PROPER: core and feature elements – Core: meaning bearing elements – Feature: function elements core SAP kinase feature Core elements extended to feature based on concatenation rules (based on POS tags) 69

Rule-based (4) • Gaizauskas (2000) CFG for protein name recognition (PASTA, EMPATHIE) • Based on morphological and lexical characteristics of terms • biochemical suffixes (-ase enzyme name) • dictionary look-up (protein names, chemical compounds, etc) • deduction of term grammar rules from Protein Data Bank Protein -> protein_modifier, protein_head, numeral 70

Rule-based (5) • Inspired by PROPER, Yapex uses Swiss-Prot to add core term elements http: //www. sics. se/humle/projects/prothalt/yapex. cgi • Hou (2003) used Yapex with context information (collocations) appearing with protein names • Rule based approaches construct rule and patterns manually or automatically • Difficult to tune to different domains 71

Machine learning systems • Learn features from training data for term recognition and classification • Most ML systems combine recognition and classification Challenges – Feature selection and optimisation – Availability of training data – detection of term boundaries 72
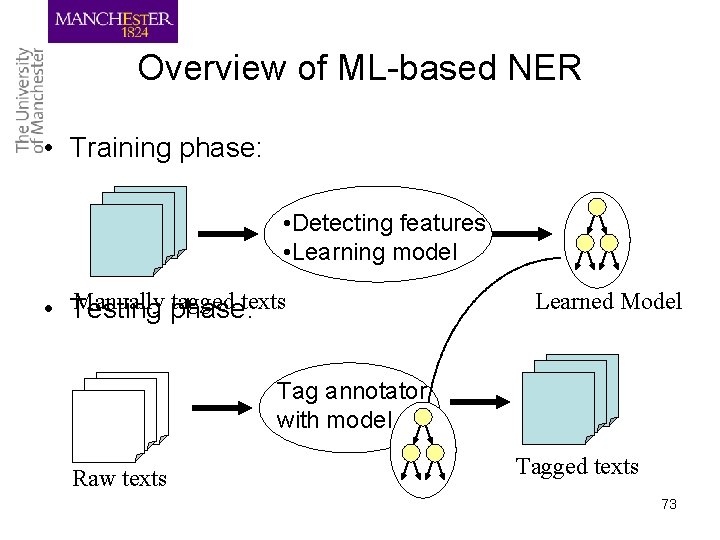
Overview of ML-based NER • Training phase: • Detecting features • Learning model Manually phase: tagged texts • Testing Learned Model Tag annotator with model Raw texts Tagged texts 73

ML (1) • Nobata et al. (1999) used Decision Tree for NER • Decision tree: one of the methods to classify a case using training data – Node: specifies some condition with a subtree – Leaf: indicates a class • Features: – Part-of-speech information – Orthographic information – Term lists 74

Example of a decision tree Each node has one condition: Is the current word in the Protein term list? No Yes Does the previous word What is the have figures? next word’s POS? No Yes Noun Verb … Each leaf has one class: Unknown PROTEIN DNA RNA …… 75

ML (2) • Collier (2000) used HMM, orthographic features for term recognition – HMM looks for most likely sequence of classes corresponding to a word sequence e. g. interleukin-2 protein/DNA – To find similarities between known words (training set) and unknown words, use character features Feature Examples Digit. Number [2]protein[3]DNA Greek. Letter [alpha]protein Two. Caps [Rel. B]protein[TAR]RNA 76

ML (2) • Use of GENIA resources as training data – Results depend on training data • Morgan (2004) used Fly. Base to construct automatically training corpus – Pattern matching for gene name recognition, noisy corpus annotated – HMM was trained on that corpus for gene name recognition 77

Support Vector Machines (1) • Kazama trained multi-class SVMs on Genia corpus • Corpus annotated with B-I-O tags – – B tags denote words at beginning of term I tags inside term O tags outside term B-protein-tag : word in the beginning of a protein name 78

SVMs for NER (2) • Yamamoto used a combination of features for protein name recognition: – Morphological, lexical, boundary, syntactic (head noun), domain specific (if term exists in biomedical database). • Lee use different features for recognition and classification. • orthographic, prefix, suffix • Contextual information 79
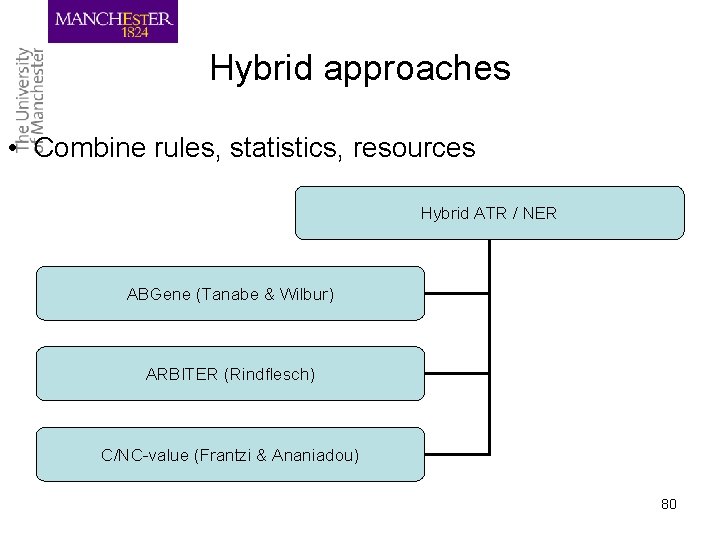
Hybrid approaches • Combine rules, statistics, resources Hybrid ATR / NER ABGene (Tanabe & Wilbur) ARBITER (Rindflesch) C/NC-value (Frantzi & Ananiadou) 80

Hybrid (1) • ABGene: protein and gene name tagger – Combines ML, transformation rules, dictionaries with statistics – Protein tagger trained on MEDLINE abstracts by adapting Brill’s tagger – Transformation rules for recognition of gene, protein names – Used GO, Locus. Link list of genes, proteins for false negative tags 81

Hybrid (2) – ARBITER (Access and Retrieve Binding Terms) uses • UMLS Metathesaurus and Gen. Bank to map NPs (binding terms) • morphological features • lexical information (head noun) – EDGAR recognises gene, cell, drug names using co-occurrences of cell, clone, expression 82

Hybrid (3) • C/NC value (Frantzi & Ananiadou, 1999) • C-value • Linguistic filters • total frequency of occurrence of string in corpus • frequency of string as part of longer candidate terms (nested terms) • number of these longer candidate terms • length of string – Output: automatically ranked terms (Ter. Mine) 83
![C-value • C- value measure extracts multi-word, nested terms [adenoid [cystic [basal [cell carcinoma]]]] C-value • C- value measure extracts multi-word, nested terms [adenoid [cystic [basal [cell carcinoma]]]]](http://slidetodoc.com/presentation_image_h/988e5bda448f6ad5971b315f2bf4b955/image-84.jpg)
C-value • C- value measure extracts multi-word, nested terms [adenoid [cystic [basal [cell carcinoma]]]] cystic basal cell carcinoma ulcerated basal cell carcinoma recurrent basal cell carcinoma 84

Term variation • variation recognition as part of ATR (Nenadic, Ananiadou) • recognise term forms and link them into equivalence classes • important if ATR is based on statistics (e. g. frequency of occurrence) – corpus-based measures are distributed across different variants – conflation of various surface representations of a given term should improve ATR 85

Simple variation • orthographic – – hyphens, slashes (amino acid and amino-acid) lower/upper cases (NF-KB and NF-kb) spelling variations (tumour and tumor) transliterations (oestrogen and estrogen) • morphological – inflectional phenomena (plural, possessives) • lexical – genuine synonyms (carcinoma and cancer) 86

Complex variation • Structural – Possessive usage of nouns using prepositions (clones of human and human clones) – Prepositional variants (cell in blood, cell from blood) – Term coordinations (adrenal glands and gonads) 87

Coordinated term variants • Structure is ambiguous – • Head coordination or term conjunction? Head or argument coordination? (N|A)+ CC (N|A)* N+ • cell differentiation and proliferation • chicken and mouse receptors 88
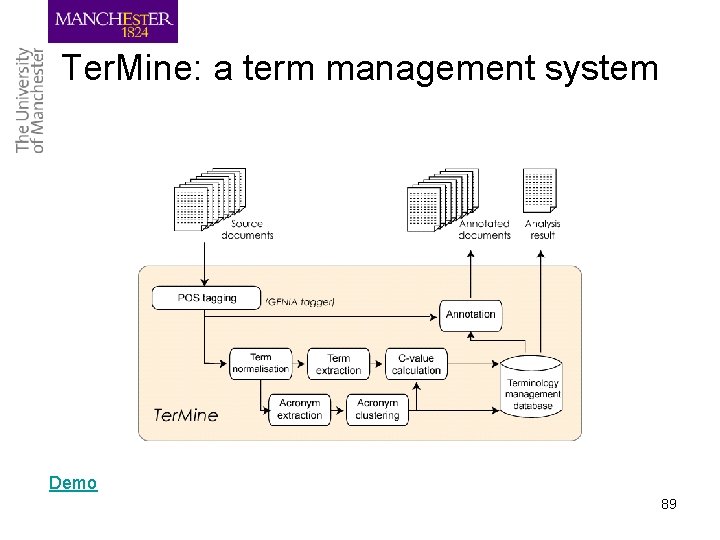
Ter. Mine: a term management system Demo 89
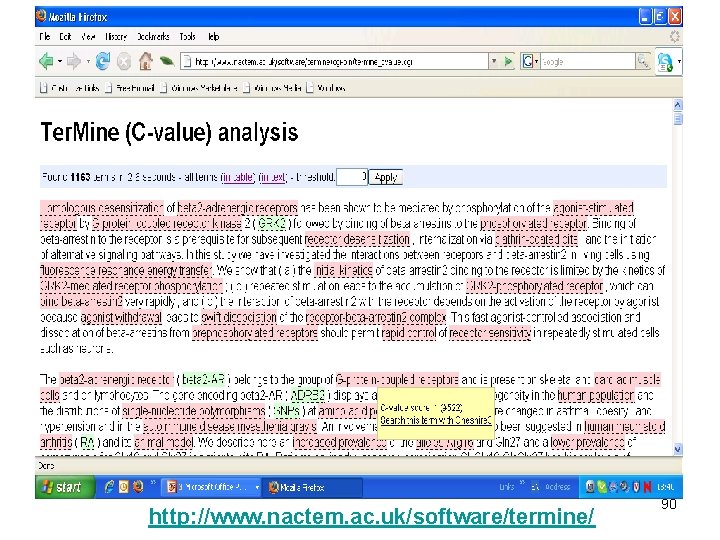
http: //www. nactem. ac. uk/software/termine/ 90

Marrying IR and terminology • IR engine plus Ter. Mine • Discover associated terms ranked according to relevance • Allow user to link term with IR for document discovery • NB compound terms • NB technical terms, not classic index terms • NB terms familiar to user, found in documents 91
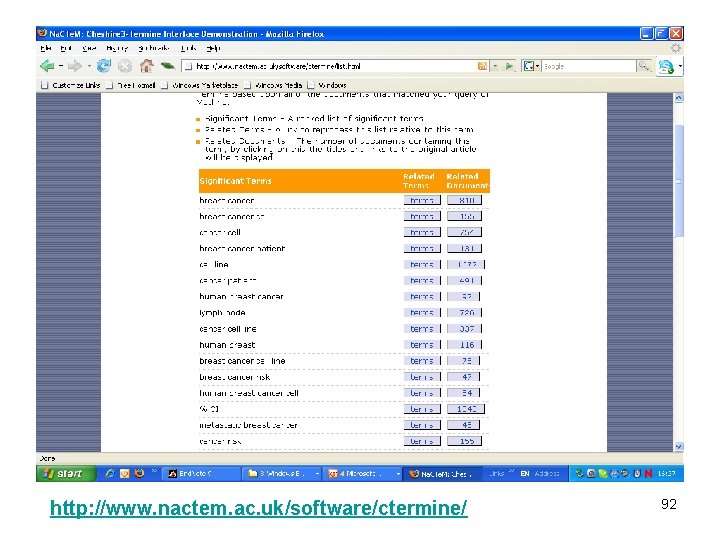
http: //www. nactem. ac. uk/software/ctermine/ 92

Biomedical IE/IR Systems • i. HOP – http: //www. ihop-net. org/Uni. Pub/i. HOP/ • EBIMed – http: //www. ebi. ac. uk/Rebholz-srv/ebimed/index. jsp • Go. Pub. Med – http: //www. gopubmed. org/ • Pub. Finder – http: //www. glycosciences. de/tools/Pub. Finder • Textpresso – http: //www. textpresso. org/ 93

Acronyms • Very productive type of term variation • Acronym variation (synonymy) – NF kappa B/ NF k. B / nuclear factor kappa B • Acronym ambiguity (polysemy) even in controlled vocabularies GR glucocorticoid receptor glutathione reductase 94

Acronym recognition • Swartz, A. & Hearst, M. (2003) A simple algorithm for identifying abbreviation definitions in biomedical text, PSB 2003, 8, 451 -462 • Adar, E. (2004) Sa. RAD: a simple and robust abbreviation dictionary, Bioinformatics, 20(4) 527 -533 • Chang, J. T. & Schutze, H. (2006) Abbreviations in biomedical text, Text Mining for Biology and Biomedicine, pp. 99 -119, Artech • Tsuruoka, Y. , Ananiadou, S. & Tsujii, J. (2005) A Machine learning approach to automatic acronym generation, ISMB, Bio. Link SIG, 25 -31 • Okazaki, N. & S. Ananiadou (2006) Acronym recognition based on term identification, Bioinformatics 95

The importance of acronym recognition • Acronyms are among the most productive type of term variation – 64, 242 new acronyms are introduced in 2004 [Chang and Schütze 06] • Acronyms are used more frequently than full terms – 5, 477 documents could be retrieved by using the acronym JNK while only 3, 773 documents could be retrieved by using its full term, c-jun N-terminal kinase [Wren et al. 05] • No rules or exact patterns for the creation of acronyms from their full form 96

Recognition • Extracting pairs of short and long forms <acronym, long form> – Distinguishing acronyms from parenthetical expressions – Search for parentheses in text; single or more words; e. g. Ab (antibody) – Limit context around ( ); limit number of words according to number of letters in acronym 97

Recognition (heuristics) – Heuristics: match letters of acronym with letters of long form using rules, patterns • letters from beginning of words • combining forms carboxifluorescein diacetate (CFDA) • Acronym normalisation to allow orthographic, structural and lexical variations • morphological information, positional info • Penalise words in long form that do not match acronym • Accidental matching argininosuccitate synthetase (AS) A S 98

Letter matching – Alignment: find all matches between letters of acronyms and their long forms and calculate likelihood (Chang & Schütze) • Solves problem of acronyms containing letters not occurring in LF • Choose best alignment based on features, e. g. position of letter etc. • Finding optimal weight for each feature challenge http: //abbreviation. stanford. edu/ 99

Acronym Recognition Okazaki, N. , Ananiadou, S. (2006) Building an abbreviation dictionary using a term recognition approach. Bioinformatics. S. Ananiadou Na. CTe. M 100

A simple algorithm – Schwartz and Hearst (2003) • Uses parenthetical expressions as a marker of a short form … long-form ‘(‘short-form ‘)’ … • All letters and digits in a short form must appear in the corresponding long form in the same order – We used hidden markov model (HMM) to … – Early repolarization (ER) is an enigma. 101

Problems of letter-matching approach • Highly dependent on the expressions in the target text – o acquired immuno deficiency syndrome (AIDS) – x acquired syndrome (AIDS) – x a patient with human immunodeficiency syndrome (AIDS) – ? magnetic resonance imaging unit (MRI) – ! beta 2 adrenergic receptor (ADRB 2) – ! gamma interferon (IFN-GAMMA) (These examples are obtained from actual MEDLINE abstracts) • Naive with respect to term variations 102

Acro. Mine’s approach • Extract a word or word sequence: – Co-occurring frequently with an acronym (e. g. , TTF-1) • 1, factor 1, transcription factor 1, thyroid transcription factor 1 – Does not co-occur with other surrounding words • thyroid transcription factor 1 • Not necessarily based on letter-matching – Note that this is a difficult case for the letter-matching algorithm • Prune unlikely candidates – Nested candidates: transcription factor 1 – Expansions: expression of thyroid transcription factor 1 – Insertions: thyroid specific transcription factor 1 103

Short-form mining • Enumerate all short forms in a target text – Using parentheses as a clue: … ‘(‘short-form ‘)’ … – Validation rules for identifying acronyms [Schwartz and Hearst 03] • It consists of at most two words • Its length is between two to ten characters • It contains at least an alphabetic letter • The first character is alphanumeric The contextual sentence of HMM and ASR. The present system consists of a hidden Markov model (HMM) based automatic speech recognizer (ASR), with a keyword spotting system to capture the machine sensitive words (registered in a dictionary) from the running utterances. 104
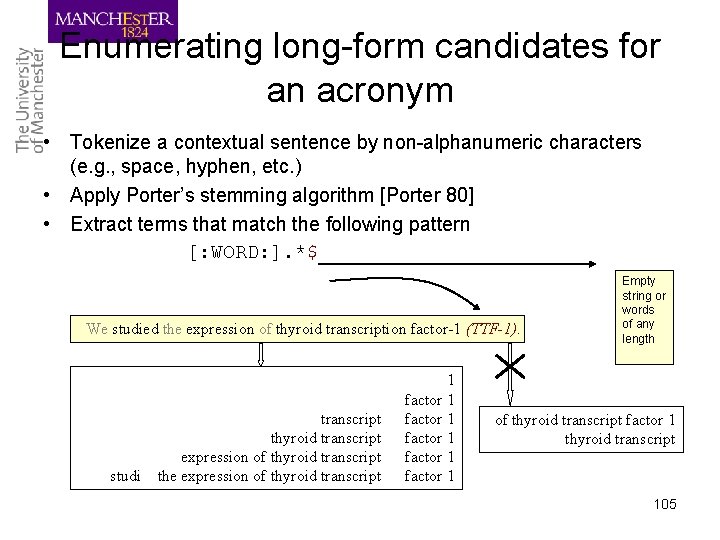
Enumerating long-form candidates for an acronym • Tokenize a contextual sentence by non-alphanumeric characters (e. g. , space, hyphen, etc. ) • Apply Porter’s stemming algorithm [Porter 80] • Extract terms that match the following pattern [: WORD: ]. *$ We studied the expression of thyroid transcription factor-1 (TTF-1). studi transcript thyroid transcript expression of thyroid transcript the expression of thyroid transcript 1 factor 1 factor 1 Empty string or words of any length of thyroid transcript factor 1 thyroid transcript 105
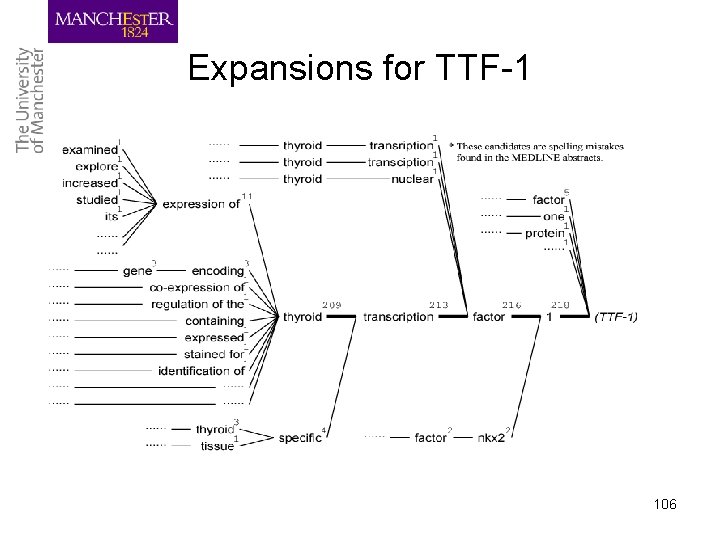
Expansions for TTF-1 106
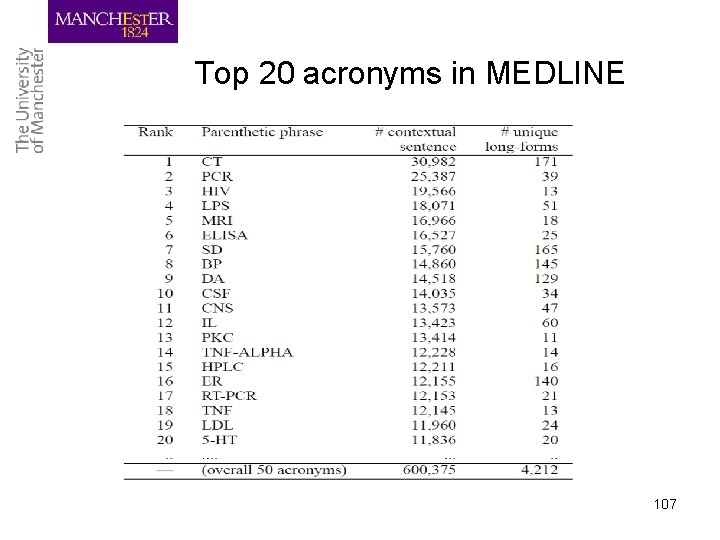
Top 20 acronyms in MEDLINE 107
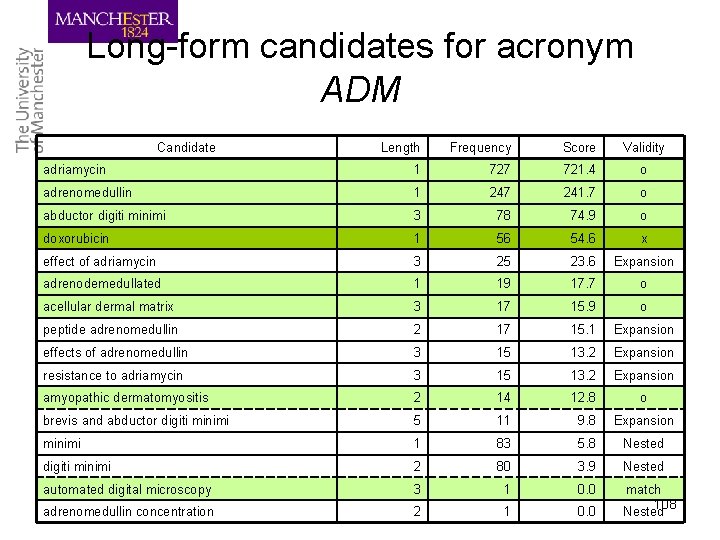
Long-form candidates for acronym ADM Candidate Length Frequency Score Validity adriamycin 1 727 721. 4 o adrenomedullin 1 247 241. 7 o abductor digiti minimi 3 78 74. 9 o doxorubicin 1 56 54. 6 x effect of adriamycin 3 25 23. 6 Expansion adrenodemedullated 1 19 17. 7 o acellular dermal matrix 3 17 15. 9 o peptide adrenomedullin 2 17 15. 1 Expansion effects of adrenomedullin 3 15 13. 2 Expansion resistance to adriamycin 3 15 13. 2 Expansion amyopathic dermatomyositis 2 14 12. 8 o brevis and abductor digiti minimi 5 11 9. 8 Expansion minimi 1 83 5. 8 Nested digiti minimi 2 80 3. 9 Nested automated digital microscopy 3 1 0. 0 match adrenomedullin concentration 2 1 0. 0 Nested 108

Long-form extraction • Long-form candidates are sorted with their scores in a descending order • A long-form candidate is considered valid if: – It has a score greater than 2. 0 – The words in the long form can be rearranged so that all alphanumeric letters appear in the same order as the short form – It is not nested or expansion of the previously chosen long forms 109
http: //www. nactem. ac. uk/software/acromine/ 110

Acronym disambiguation • Local acronyms – Accompany their expanded forms in documents • Global acronyms – Appear in documents without the expanded forms stated – Need to be their correct expanded forms identified • Immunomodulatory effects of CT were investigated in a rat model, and the effects of CT on rat renal allograft (from Lewis rat to WKAH rat) were also examined. • Immunomodulatory effects of cholera toxin (CT) were investigated in a rat model, and the effects of cholera toxin (CT) on rat renal allograft (from Lewis rat to Wistar-King-Aptekman-Hokudai (WKAH) rat) were also examined. 111
Acronym disambiguation Sample text: Considerations in the identification of functional RNA structural elements in genomic alignments (Tomas Babak et al) 112 http: //www. biomedcentral. com/1471 -2105/8/33

Term structuring 113

Term structuring • term clustering (linking semantically similar terms) and term classification (assigning terms to classes from a pre -defined classification scheme) • Hypothesis: similar terms tend to appear in similar contexts (patterns) • combining various sources of similarity: – – lexical syntactic contextual Ontological (using external resources) 114

Term structuring • Based on term similarities – choice of features: – domain specific – linguistic ontology text • ontology-based similarity • textual similarity – internal features – contextual features 115

Using ontologies • two terms should match if they are: – identified as variants – siblings in the is-a hierarchy – in the is-a or part-whole relation • the distance between the corresponding nodes in the ontology should be transformed into the matching score ► I. Spasic presentation MIE Tutorial http: //www. nactem. ac. uk/ 116

Using text • number of neologisms: terms are not in the ontologies • Use of text based techniques to calculate similarities • edit distance (ED) – the minimal number (or cost) of changes needed to transform one string into the other • edit operations: insertion deletion replacement transposition. . . a-c. . . abc. . . a-c. . . adc. . . acb. . . • use of dynamic programming 117

Term similarities – lexical similarity: based on sharing term head and/or modifier(s) --hyponymy nuclear receptor orphan nuclear receptor – Sharing heads progesterone receptor oestrogen receptor • Specific types of associations – mainly general is_a and part_of – some domain-specific, e. g. binding: CREP binding protein 118

Contextual similarities • Features from context – – syntactic category terminological status position relative to the term syntactic relation between a context element and the term – semantic properties – semantic relation between a context element and the term ……. 119

Lexical & syntactic patterns • a lexico-syntactic pattern: . . . Term (, Term)* [, ] and other Term. . . • the leading Terms hyponyms of the head Term. . . antiandrogens, hydroxyflutamide, bicalutamide, cyproterone acetate, RU 58841, and other compounds. . . • candidate instances of the hyponymy relation: hyponym( antiandrogens, compound ) hyponym( hydroxyflutamide, compound ) hyponym( bicalutamide, compound ) hyponym( cyproterone acetate, compound ) hyponym( RU 58841, compound ) 120

Contextual information • automatic pattern mining for most important context patterns – find most important contexts in which a term appears … receptor is bound to these DNA sequences … … proteins bound to the DNA … … estrogen receptor bound to DNA … … steroid receptor coactivator-1 when bound to DNA … … progesterone receptor complexes bound to DNA … … RXRs bound to respective DNA elements in vitro … … glucocorticoid receptor to bind DNA … pattern: <TERM> V: bind <TERM: DNA> 121

Stumbling blocks • Lexical similarities affected by many neologisms and ad hoc names – only 5% of most frequent terms in GENIA belonging to same biomedical class have some lexical links • how much context to use? (sentence, phrase, abstract, …) • Attempts at using co-occurrence: many report up to 40% of co-occurrence based relationships biologically meaningless 122

Term similarities • SOLD = Syntactic, Ontology-driven & Lexical Distance (Spasic, I. & Ananiadou, S. 2005, Bioinformatics) • hybrid approach to comparing term contexts, which relies on: – linguistic information (acquired through tagging and parsing) – domain-specific knowledge (obtained from the ontology) • based on the approximate pattern matching • combines ontology-based similarity with corpus-based similarity using both internal and contextual features 123

Challenges of biomedical terminology • Linking termforms in text with existing resources • Term clustering, classification and linking to databases, ontologies • Selection of most representative terms (concepts) in documents (important for improved IR, database curation, annotation tasks) • Efficient term management important for updating terminological and ontological resources, text mining applications e. g. IE, Q/A, summarisation, linking heterogeneous resources, IR etc 124

Information Extraction in Biology • Results appear depressed compared to general language – Dependent of earlier stages of processing (tokenisers, taggers, results from NER, etc) – MUC data 80% F-score template relations, 60% events – Challenge for bio-text mining is to achieve similar results • Evaluation see Hirschman, L. (Text mining book) Bio. Cre. ATive 2004 125

I Information Extraction 126

IE in Biology Ø Pattern-matching Ø Context-free grammar approaches Ø Full parsing approaches Ø Sublanguage driven IE Ø Ontology-driven IE Mc. Naught, J. & Black, W. (2006) Information Extraction, Text Mining for Biology & Biomedicine, Artech house, pp. 143 -177 127

Pattern-matching IE – Usual limitations with non inclusion of semantic processing – Large amount of surface grammatical structures = too many patterns (Zipf’s law) – Cannot explore syntactic generalisations (active, passive voice) – Systems extract phrases or entire sentences with matched patterns; restricted usefulness for subsequent mining 128

Pattern-matching systems (1) Ø Bio. IE uses patterns to extract sentences, protein families, structures, functions. . Ø Presents user with relevant information, improvement from classic IR Ø Bio. RAT uses “deeper” analysis, tagging, apply RE over POS tags, stemming, gazetter categories etc Ø Templates apply to extract matching phrases, primitive filters (verbs are not proteins, etc) 129

Pattern matching systems (2) Ø RLIMS-P (Hu) protein phosphorylation by looking for enzymes, substrates, sites assigned to agent, theme, site roles of phosphorylation relations Ø Pos tagger, trained on newswire, chunking, semantic typing of chunks, identification of relations using pattern-matching rules Ø Semantic typing of NPs: using combination of clue words, suffixes, acronyms etc Ø Semantically typed sentences matched with rules Ø Patterns target sentences containing phosphorylate 130

Full parsing approaches • Link Grammar applied for protein-protein interactions; general English grammar adapted to bio-text • Link Grammar finds all possible linkages according to its grammar • Number of analyses reduced by random sampling, heuristics, processing constraints relaxed – 10, 000 results permitted per sentence – 60% of protein interactions extracted – Problems: missing possessive markers & determiners, coordination of compound noun modifiers 131

Full parsing IE (2) • Not all parsing strategies suitable for bio-text mining • Text type, abstracts, “ungrammaticality” related with sublanguage characteristics? • Ambiguity and full parsing; fragmentary phrases (titles, headings, text in table cells, etc) • CADERIGE project used Link grammar but on shallow parsing mode • Kim & Park (Bio. IE) use combinatorial categorial grammar, annotated with GO concepts, extract general biological interactions • 1, 300 patterns applied to find instances of patterns with keywords 132

Full parsing (3) • Keywords indicate basic biological interactions • Patterns find potential arguments of the interaction keywords (verbs or nominalisations) – Validated arguments mapped into GO concepts – Difficult to generalise interaction keyword patterns • Bio. IE’s syntactic parsing performance improved after adding subcategorisation frames on verbal interaction keywords 133

Full parsing (4) – 1. 2. 3. 4. 5. Daraselia(2004) use full parsing and domain specific filter to extract protein interactions All syntactic analyses discovered using CFG and variant of LFG Each alternative parse mapped to its corresponding semantic representation Output= set of semantic trees, lexemes linked by relations indicating thematic or attributive roles Apply custom-built, frame based ontology to filter representations of each sentence Preference mechanism controls construction of frame tree, high precision, low recall (21%) 134

Sublanguage-driven IE (1) • Language of a special community (e. g. biology) • Particular set of constraints re GL • Constraints operate at all linguistic levels – – Special vocabulary (terms) Specialised term formation rules Sublanguage syntactic patterns Sublanguage semantics • These constraints give rise to the informational structure of the domain (Z. Harris) • See JBI 35(4) Special Issue on Sublanguage 135

GENIES system • Employs SL approach to extract biomolecular interactions • Uses hybrid syntactic-semantic rules – Syntactic and semantic constraints referred to in one rule • Able to cope with complex sentences • Frame-based representation – Embedded frames • Domain specific ontology covers both entities and events 136

GENIES system • Default strategy: full parsing – Robust due to sublanguage constraints – Much ambiguity excluded • If full parse fails, partial parsing invoked – Maintains good level of recall • Precision: 96%, Recall: 63% 137

Ontology-driven IE • Until recently most rule based IE have used neither linguistic lexica nor ontologies – Reliance on gazetteers – Small number of semantic categories • Gazetteer approach not well suited in bio. IE • Ontology based vs ontology driven – Passive use of ontologies, map discovered entity to concept – Active use, ontology guides and constrains analysis, fewer rules • Examples: PASTA, Gen. IE not SL • GENIES, SL and ontology driven 138

Summary: simple pattern matching Ø Over text strings Ø Many patterns required, no generalisation possible Ø Over POS Ø Some generalisation but ignore sentence structure Ø POS tagging, chunking, semantic p-m, typing Ø Limited generalisation, some account taken of structure, limited consideration of SL patterns 139

Summary: full parsing Ø Full parsing on its own, parsing done in combination with chunking, partial parsing, heuristics) to reduce ambiguity, filter out implausible readings Ø Ø GL theories not appropriate Difficult to specialise for biotext Many analyses per sentence Missing information due to sublanguage meaning 140

Summary: sublanguage approach Ø Ø Exploits a rich SL lexicon Describes SL verbs in detail Syntactic-semantic grammar Current systems would benefit from adopting ontologydriven approach 141

Ontology-driven Ø Uses event concept frames to guide processing Ø Integration of extracted information Ø Current systems would benefit from adopting also SL approach 142

Applications 143

How do we apply TM to Systems Biology? REFINE project • Adapting TM tools to evaluate the basis in the literature for the structure of biochemical and signalling models in systems biology • Integrating TM with visualisation for better understanding of the evidence for biochemical and signalling pathways • Enriching models encoded in SBML with information derived from TM Kell, Ananiadou, Tsujii 144

Applications • Semantic annotation not only based on concepts but also on facts, events extracted by IE • Enables semantic querying • Facilitates curation • Hypothesis generation for scientific discovery 145

Applications • Other text mining applications – Summarisation – Question answering • Integration of IR with TM – Terms / concepts as index terms – Topic detection – Document clustering and classification 146
- Slides: 146