Tetrads with n genes ABCD x abcd Aa
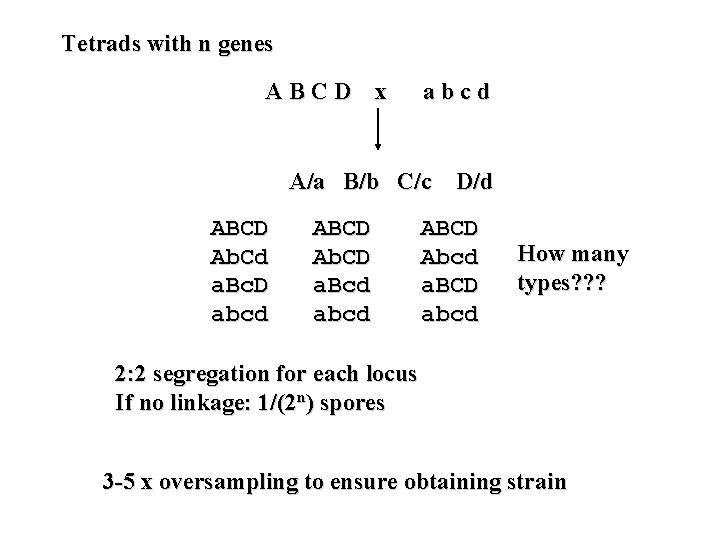
Tetrads with n genes ABCD x abcd A/a B/b C/c D/d ABCD Ab. Cd a. Bc. D abcd ABCD Ab. CD a. Bcd abcd ABCD Abcd a. BCD abcd How many types? ? ? 2: 2 segregation for each locus If no linkage: 1/(2 n) spores 3 -5 x oversampling to ensure obtaining strain

Suppressor screens, examples Suppressor of Multivulva in C. elegans Activated Go-alpha in C. elegans Enhancer screens, examples Rough eye in Drosophila
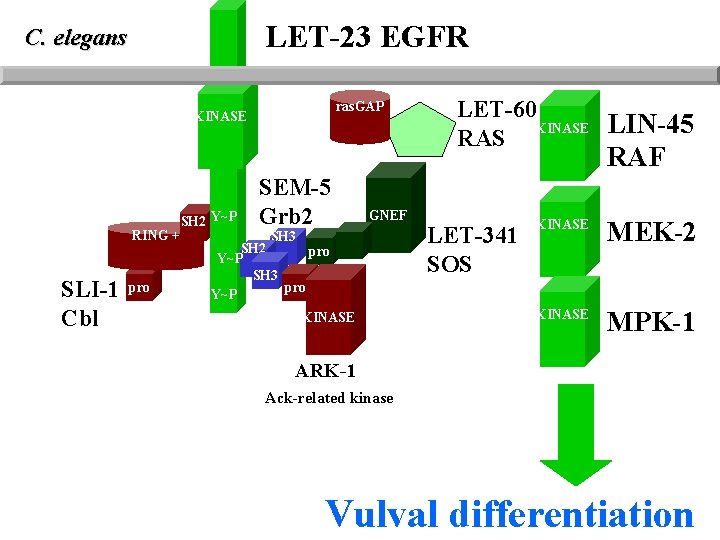
LET-23 EGFR C. elegans ras. GAP KINASE LET-60 KINASE LIN-45 RAS RAF RING + SLI-1 Cbl pro SH 2 Y~P SEM-5 Grb 2 GNEF SH 3 SH 2 pro Y~P SH 3 pro Y~P KINASE LET-341 SOS KINASE MEK-2 KINASE MPK-1 ARK-1 Ack-related kinase Vulval differentiation
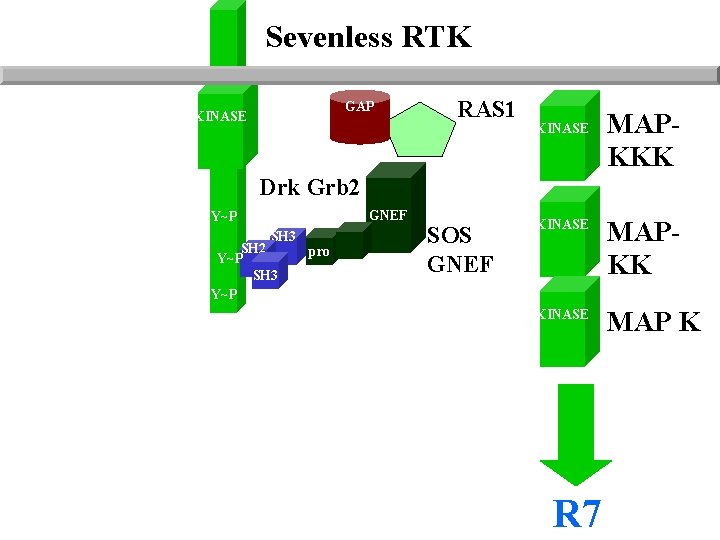
Sevenless RTK GAP KINASE RAS 1 KINASE MAPKKK Drk Grb 2 GNEF Y~P SH 3 SH 2 Y~P SH 3 Y~P pro SOS GNEF KINASE MAPKK KINASE MAP K R 7

Drosophila Photoreceptor Development R 8 induces R 7
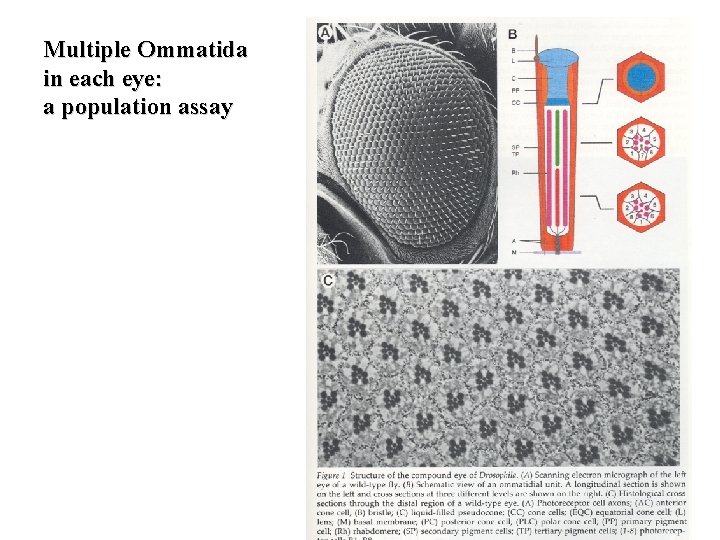
Multiple Ommatida in each eye: a population assay


An enhancer screen for essential genes required for R 7 development The fly eye consists of approximately 800 20 -cell repeating units known as ommatidia. Each ommatidium consists of eight photoreceptor neurons (R 1 -R 8), four lens secreting cone cells and eight additional accessory cells. The ommatidia arise from an undifferentiated epithelium by a series of cell interactions. We will only consider an interaction between the R 8 and presumptive R 7 cells that determines the fate of R 7. The R 7 photoreceptor detects light in the UV range. Screens for mutants with ommatidia that lack R 7 cells identified three genes: sevenless (sev), bride of sevenless (Boss) and seven-in-abstentia (sina). Adult flies homozygous for mutations in any of these genes have ommatidia that lack an R 7 cell and contain an additional cone cell. In the absence of R 7 differentiation, the presumptive R 7 cell becomes a cone cell. sev and sina are a receptor tyrosine kinase and a nuclear protein, respectively, and both genes act in R 7 to specify R 7's fate. boss appears to encode the ligand for the Sev receptor tyrosine kinase, and in contrast to sev and sina, acts in R 8 cell to specify R 7's fate. Now consider the problem that many genes functioning downstream of receptor tyrosine kinse receptor activation are likely to be required for other tyrosine kinase signaling pathways that are required for the viability of the organism. How can one use the fly eye to identify such mutations in such genes. Make a partially active mutant version of sev and introduce it into a sev mutant background. These flies have a temperature-sensitive phenotype. A fly carrying one copy of this transgene is wildtype at 22. 7 o. C (R 7 is present). However, at 24. 3 o. C R 7 is absent
![sev/Y; +/+male * X sev/sev; /+; P[sev-ts]/+ sev/sev; +/+; P[sev-ts]/balancer * sev/Y; /+; P[sev-ts]/+ sev/Y; +/+male * X sev/sev; /+; P[sev-ts]/+ sev/sev; +/+; P[sev-ts]/balancer * sev/Y; /+; P[sev-ts]/+](http://slidetodoc.com/presentation_image/1b0fe2965d4fe7e04f091ed70ac0b93e/image-9.jpg)
sev/Y; +/+male * X sev/sev; /+; P[sev-ts]/+ sev/sev; +/+; P[sev-ts]/balancer * sev/Y; /+; P[sev-ts]/+ Screen for absence of R 7 in individual flies. Isolate these chromosomes by balancing. R 7 present R 7 absent sev/sev; +/+; P[sev-ts]/Y at 22. 7 o. C sev/sev; +/+; P[sev-ts]/Y at 24. 3 o. C sev/sev; */+; P[sev-ts]/Y at 22. 7 o. C Look for mutation (*) that confers dominant enhancement of sev phenotype
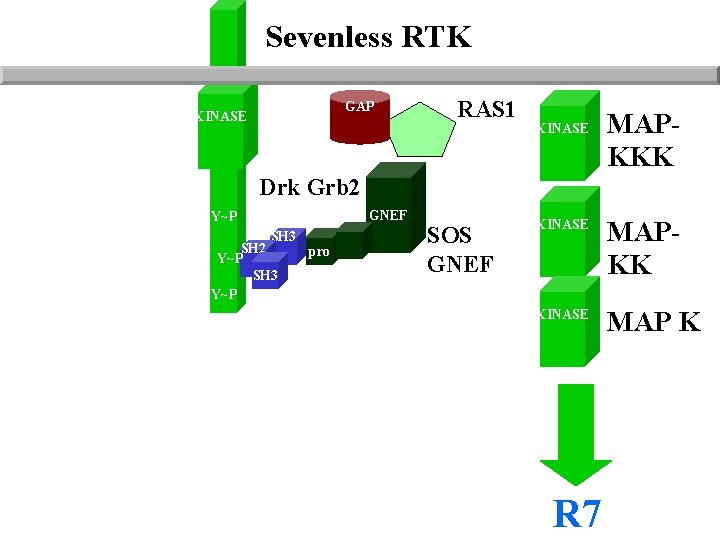
Sevenless RTK GAP KINASE RAS 1 KINASE MAPKKK Drk Grb 2 GNEF Y~P SH 3 SH 2 Y~P SH 3 Y~P pro SOS GNEF KINASE MAPKK KINASE MAP K R 7
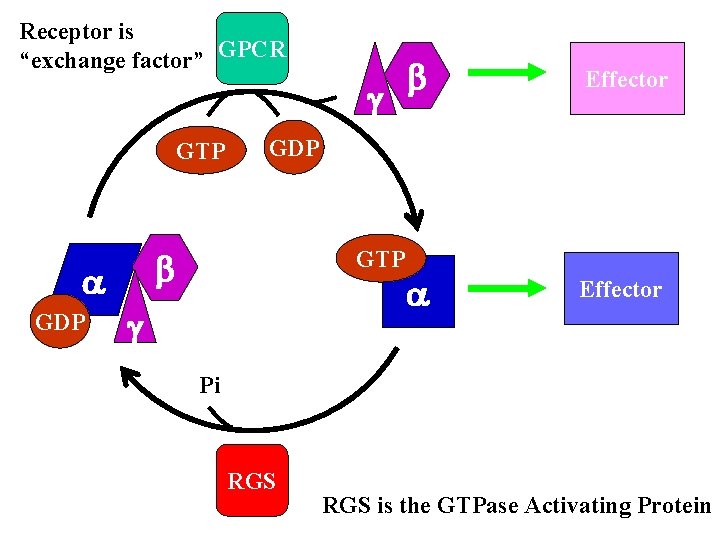
Receptor is “exchange factor” GPCR g GTP GDP Effector GDP b b GTP g Pi RGS is the GTPase Activating Protein
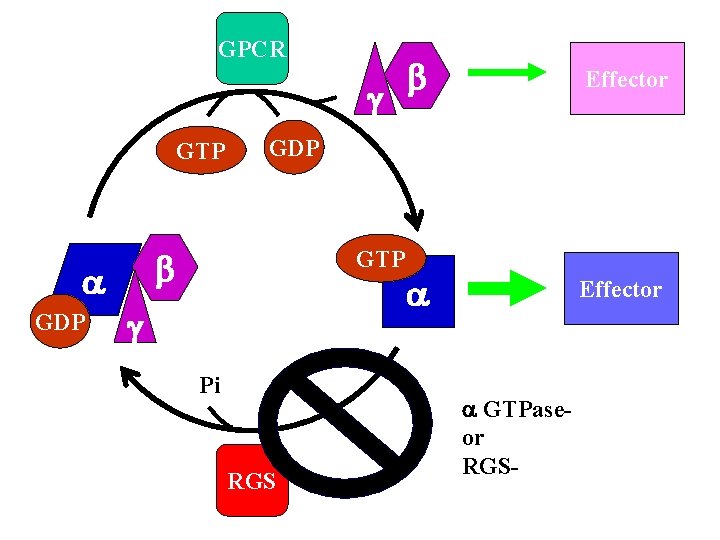
GPCR g GTP GDP Effector GDP b b GTP g Pi RGS GTPaseor RGS-

G proteins Gq and Go control movement C. elegans Genotype Phenotype Wild type egl-30(lf) egl-30(gf) goa-1(lf) goa-1(gf) egl-30(lf) goa-1(lf) wild-type paralyzed hyperactive paralyzed lf, loss-of-function; gf, gain-of-function

Mutations that Suppress activated Go sy. Is 17; sag-4(sy 433) Before Heat Shock After Heat Shock Jane Mendel, Yvonne Hajdu-Cronin, Wen Chen

Suppressors of Activated Go (Sag) CLASS I hyperactive • dgk-1/sag-1 (14 alleles) encodes diacylgycerol kinase • eat-16(sy 348) (p. k. a. sag-2) encodes RGS 7 homologue CLASS II wild type • sag-4, 8 sag-4 encodes cyclin L homologue CLASS III Egg-laying defective sag-3 encodes Heat Shock Factor • sag-3, 5 CLASS IV wild type • sag-6 CLASS V Egg-laying defective • sag-7 Yvonne Hajdu-Cronin & Wen Chen
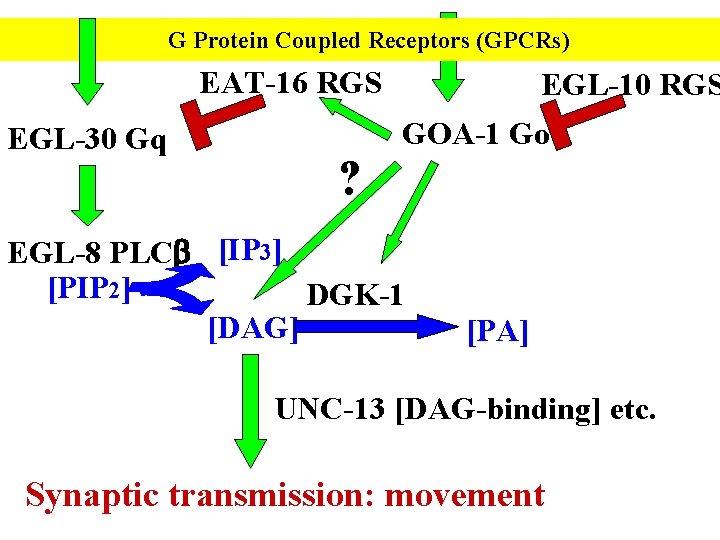
G Protein Coupled Receptors (GPCRs) EAT-16 RGS EGL-30 Gq ? EGL-10 RGS GOA-1 Go EGL-8 PLCb [IP 3] [PIP 2] DGK-1 [DAG] [PA] UNC-13 [DAG-binding] etc. Synaptic transmission: movement

Extragenic suppression • many mechanisms--key issue is the genetic specificity of the suppressor gene-specific allele-nonspecific gene-specific allele-specific gene-nonspecific allele-specific epistasis (bypass suppression) direct interaction? ‘informational’ suppression

suppression by compensatory change in direct interactor? • ‘Lock and Key’ model: binding site is restored • in general a very rare event as target size is 1(or a few) bp-need screens of >106 genomes • RNA-RNA interactions: – restoration of base pairing (nonsense suppression) – splice site suppression e. g. Lesser + Guthrie 1993 Science 262: 1982 • protein-DNA interactions – lac operon: o. C mutations suppressed by mutations in repressor that bind more tightly to operator (Pfahl 1981, J. Mol. Biol. 147: 1 -10) • protein-protein interactions?

allele-specific suppression • null mutants are not suppressed, so not bypass suppressor • stabilization or altered processing of mutant gene product
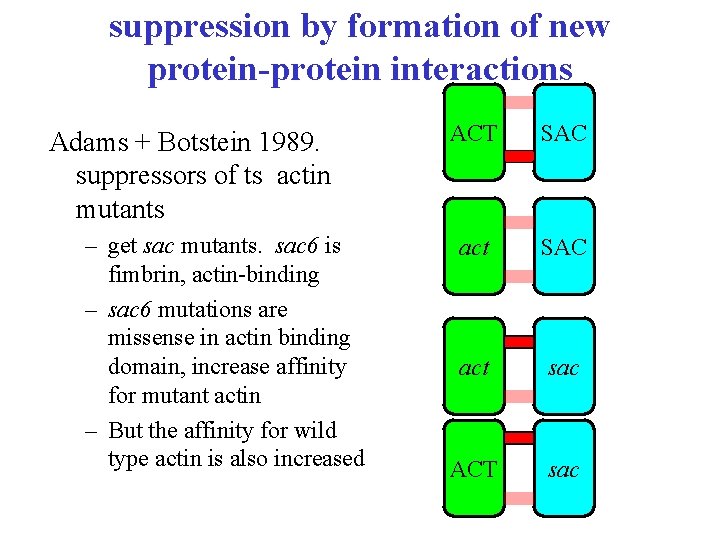
suppression by formation of new protein-protein interactions Adams + Botstein 1989. suppressors of ts actin mutants – get sac mutants. sac 6 is fimbrin, actin-binding – sac 6 mutations are missense in actin binding domain, increase affinity for mutant actin – But the affinity for wild type actin is also increased ACT SAC act sac ACT sac

gene non-specific, allele specific • 1. 2. 3. 4. 5. suppression at level of gene expression: ‘informational’ Nonsense suppression Frameshift suppression Splicing machinery stabilization of unstable m. RNA or protein suppression of transposon insertion alleles

nonsense suppression • conditional ‘amber’ mutations in many T 4 genes (Epstein et al) – grow on one E coli strain (CR 63) but not on B – cause premature termination – suppression due to mutant t. RNA that can recognize amber codon UAG and insert amino acid (usually Trp; codon is UGG) – amber suppressor strains are a bit sick because of readthrough

frameshift suppression • extragenic suppression of frameshifts by two mechanisms – limitation of Trp-t. RNA – other t. RNAs loosely bind to codon (mismatch) and allow frameshifting – also mutant t. RNA with 4 -base anticodon now ‘reads’ frameshift as a 3 -base codon…

suppression by stabilization of message • m. RNAs with ‘premature’ stop codons are recognized and degraded – nonsense mediated decay/ ‘m. RNA surveillance’ – Upf pathway (yeast), SMG pathway (worms) – get rid of aberrant m. RNAs before they get to ribosome • some nonsense mutations can be suppressed if partially functional protein can be made
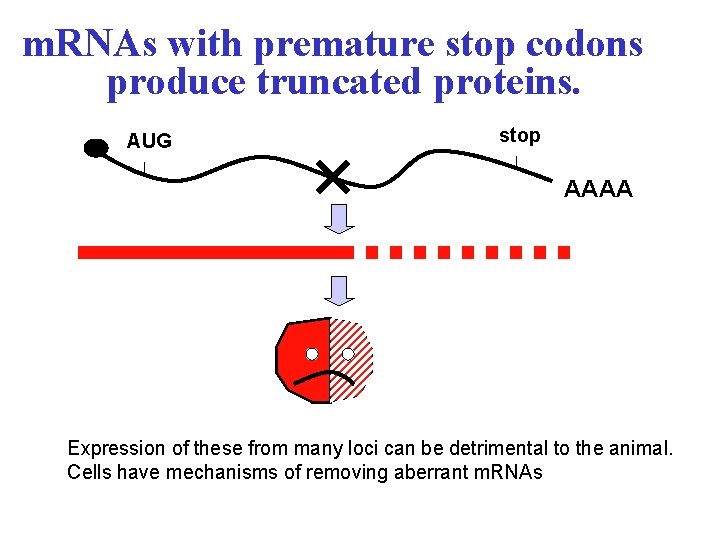
m. RNAs with premature stop codons produce truncated proteins. AUG stop AAAA Expression of these from many loci can be detrimental to the animal. Cells have mechanisms of removing aberrant m. RNAs
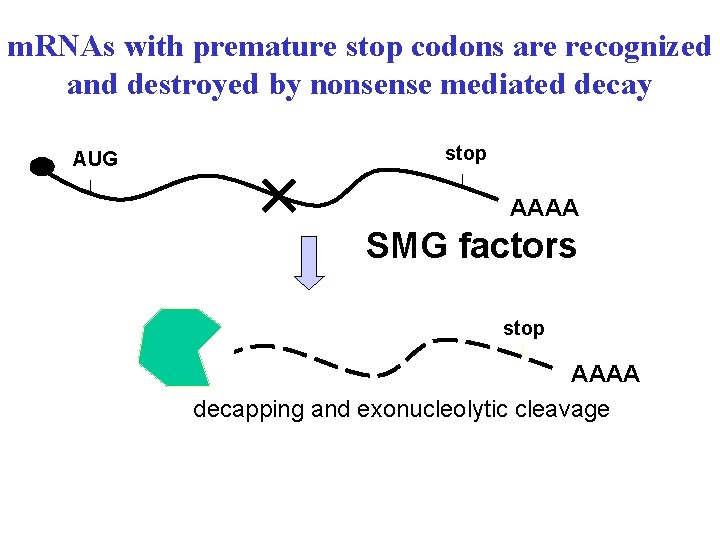
m. RNAs with premature stop codons are recognized and destroyed by nonsense mediated decay AUG stop AAAA SMG factors stop AAAA decapping and exonucleolytic cleavage

Screens for suppressors of nonsense mutations revealed smg genes • • smg-1 phosphatidylinositol-3 kinase homolog smg-2 Upf 1 helicase homolog, phosphoprotein smg-3 Upf 2 homolog smg-4 Upf 3 homolog smg-5 novel, binds SMG-7 smg-6 -smg-7 novel, binds SMG-5 Mutations in the proteins required for nonsense mediated decay suppress nonsense mutations by allowing stabilizing m. RNAs with premature stop codons. Functional proteins are made since low levels of readthrough make some normal protein or because expression of the truncated protein can suppress the phenotype Hodgkin J, Papp A, Pulak R, Ambros V, Anderson P. A new kind of informational suppression in the nematode Caenorhabditis elegans. Genetics. 1989 Oct; 123(2): 301 -13.
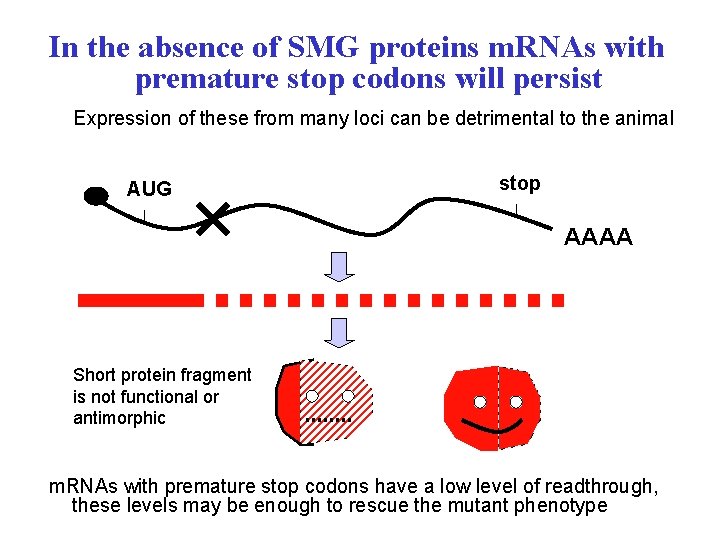
In the absence of SMG proteins m. RNAs with premature stop codons will persist Expression of these from many loci can be detrimental to the animal AUG stop AAAA Short protein fragment is not functional or antimorphic m. RNAs with premature stop codons have a low level of readthrough, these levels may be enough to rescue the mutant phenotype

suppression by stabilization of protein • E. coli lon protease degrades aberrant proteins • mutations in lon suppress thermolabile mutations in many genes (RNA polymerase etc)
- Slides: 29