Templating Effect of Lipid Membranes on Alzheimers Amyloid

Templating Effect of Lipid Membranes on Alzheimer’s Amyloid Beta (Ab) Peptide Canay Ege canayege@uchicago. edu Departments of Chemistry, the James Frank Institute and the Institute for Biophysical Dynamics The University of Chicago 5735 S Ellis Ave Chicago IL 60637

Projects worked on Amyloid Beta – lipid membrane interactions X-ray scattering Results: Ege C, Majewski J, Wu G, Kjaer K, Lee KYC (in preparation) Templating: Ege C, Majewski J, Wu G, Kjaer K, Lee KYC, CHEMPHYSCHEM (2004) (accepted) Isotherm and FM: Ege C, Lee KYC, BIOPHYSICAL JOURNAL: 87 1732 (2004) Membrane Sealing with tri-block copolymers Wu G, Majewski J, Ege C, Kjaer K, Weygand MJ, Lee KYC, BIOPHYSICAL JOURNAL (submitted) Wu G, Majewski J, Ege C, Kjaer K, Weygand MJ, Lee KYC PHYSICAL REVIEW LETTERS 93, Art. No. 028101 (2004) Cholesterol complexes (lipid rafts) Ege C, Ratajczak M, Majewski J, Kjaer K, Lee KYC (in preparation) Domain boundary instability Diamant H, Witten TA, Ege C, Gopal A, Lee KYC, PHYSICAL REVIEW E 63: Art. No. 061602 (2001) 2

OUTLINE • Alzheimer’s Disease - background • Motivation • Experimental Techniques, part 1 • Langmuir Isotherms • Fluorescence Microscopy • Results – part 1 • Experimental Techniques, part 2 • X-ray and Neutron Reflectivity • Grazing Incidence X-ray Diffraction • Results – part 2 • Oligomeric vs. monomeric Ab is the culprit ? (part 3) • Conclusion • Future Directions
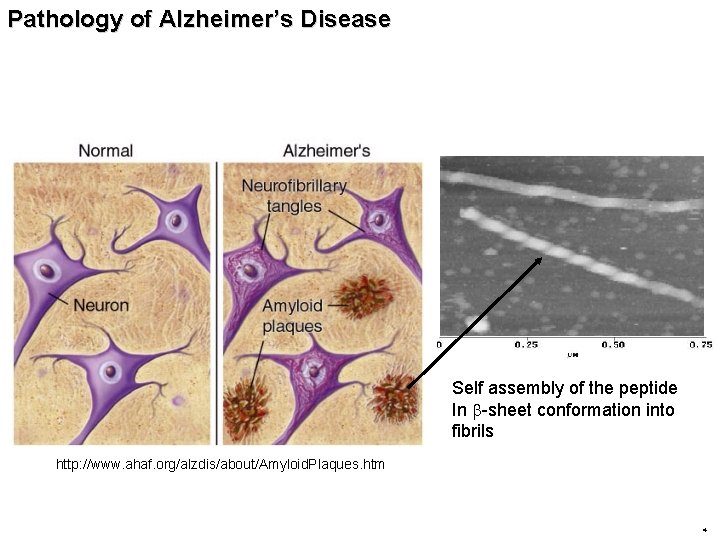
Pathology of Alzheimer’s Disease Self assembly of the peptide In b-sheet conformation into fibrils http: //www. ahaf. org/alzdis/about/Amyloid. Plaques. htm 4
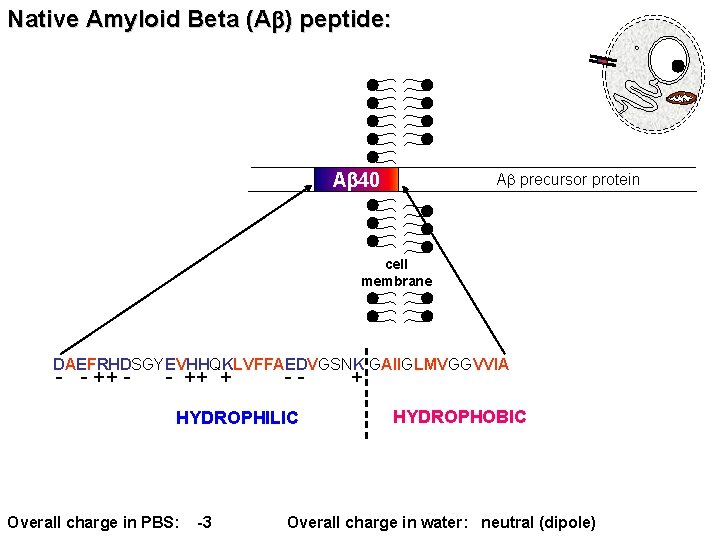
Native Amyloid Beta (Ab) peptide: Ab 40 Ab precursor protein cell membrane DAEFRHDSGYEVHHQKLVFFAEDVGSNK GAIIGLMVGGVVIA - - ++ + -- HYDROPHILIC Overall charge in PBS: -3 + HYDROPHOBIC Overall charge in water: neutral (dipole)
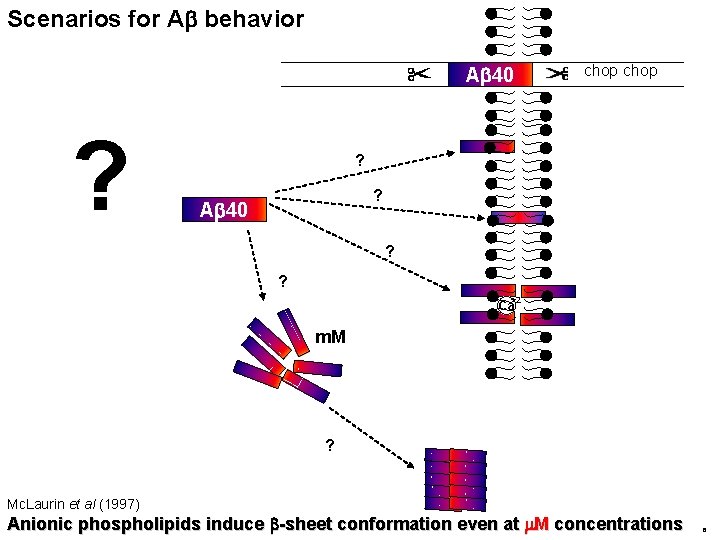
Scenarios for Ab behavior Ab 40 ? chop ? ? Ab 40 ? ? +2 Ca m. M ? Mc. Laurin et al (1997) Anionic phospholipids induce b-sheet conformation even at m. M concentrations 6
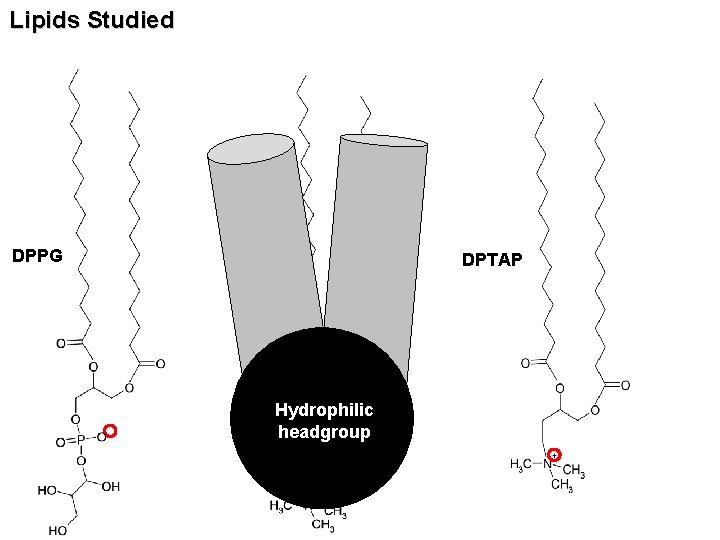
Lipids Studied DPPG DPPC Hydrophilic headgroup DPTAP
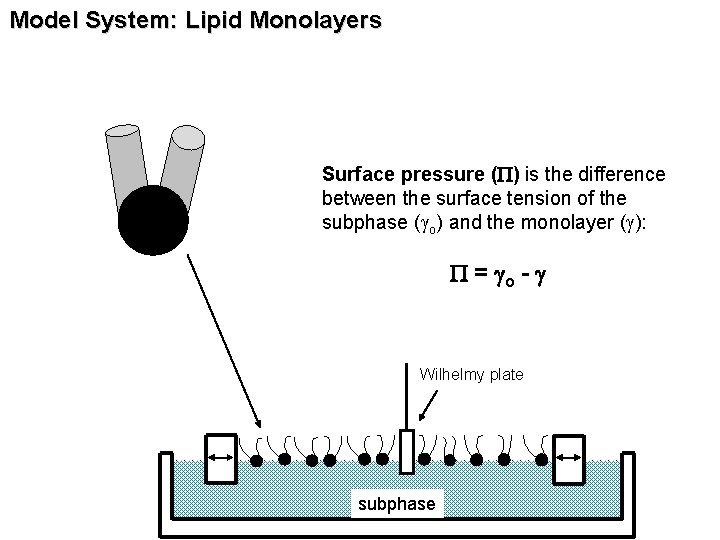
Model System: Lipid Monolayers Surface pressure (P) is the difference between the surface tension of the subphase (go) and the monolayer (g): P = go - g Wilhelmy plate subphase
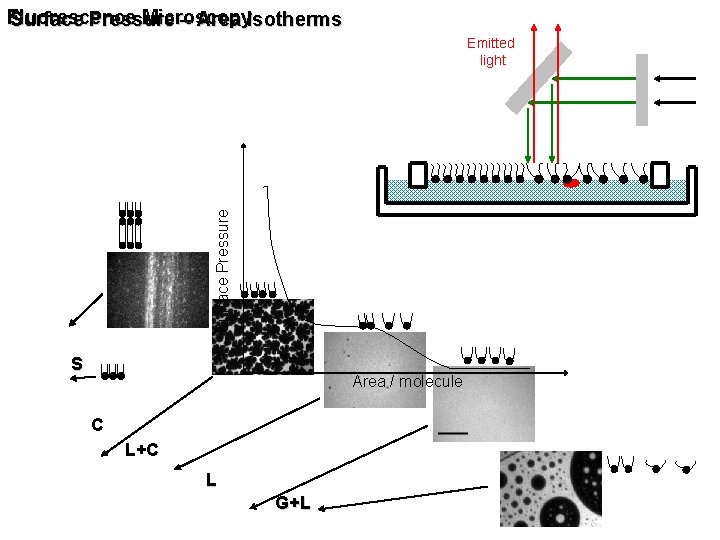
Fluorescence Microscopy Surface Pressure – Area Isotherms Surface Pressure Emitted light S Area / molecule C L+C L G+L
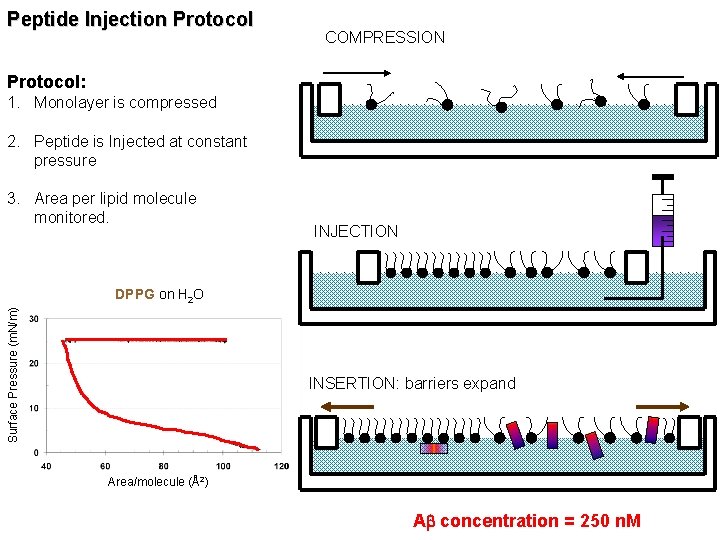
Peptide Injection Protocol COMPRESSION Protocol: 1. Monolayer is compressed 2. Peptide is Injected at constant pressure 3. Area per lipid molecule monitored. INJECTION Surface Pressure (m. N/m) DPPG on H 2 O INSERTION: barriers expand Area/molecule (Å2) Ab concentration = 250 n. M
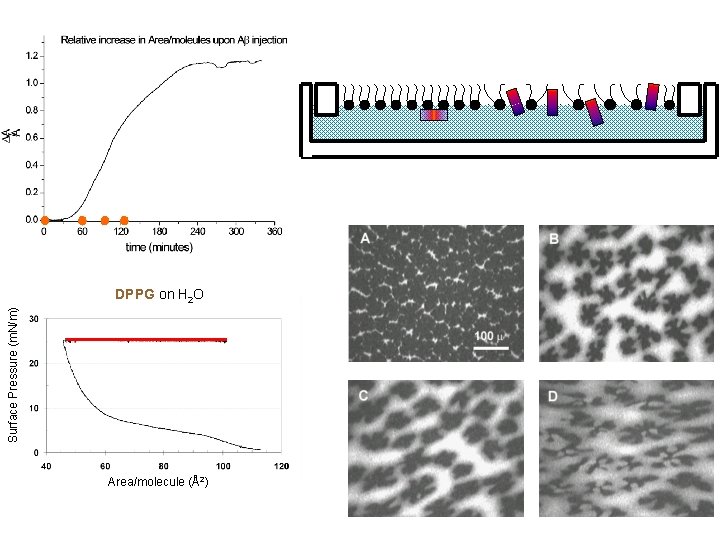
Surface Pressure (m. N/m) DPPG on H 2 O Area/molecule (Å2)
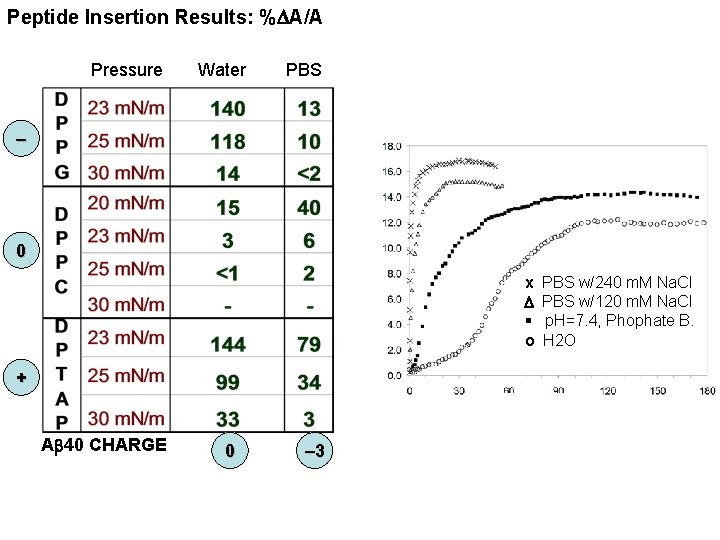
Peptide Insertion Results: %DA/A Pressure Water PBS – 0 x D § o + Ab 40 CHARGE 0 – 3 PBS w/240 m. M Na. Cl PBS w/120 m. M Na. Cl p. H=7. 4, Phophate B. H 2 O
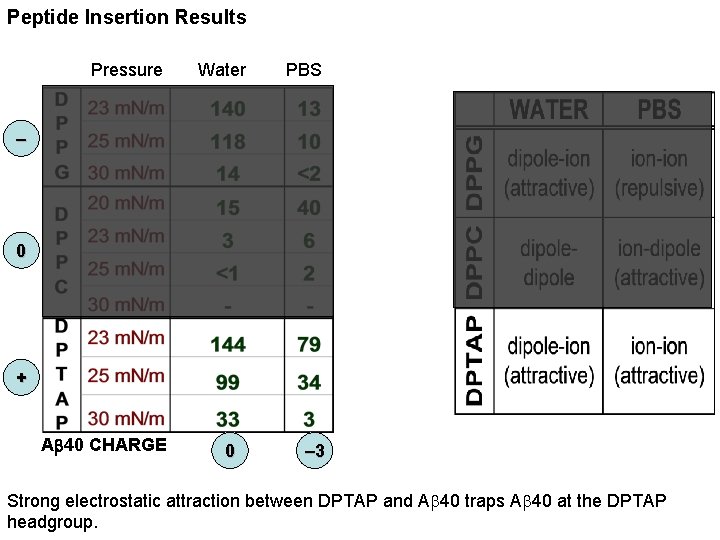
Peptide Insertion Results Pressure Water PBS – 0 + Ab 40 CHARGE 0 – 3 Strong electrostatic attraction between DPTAP and Ab 40 traps Ab 40 at the DPTAP headgroup.

DPTAP (30 m. N/m) DPPC (23 m. N/m) Dual probe fluorescence in PBS:
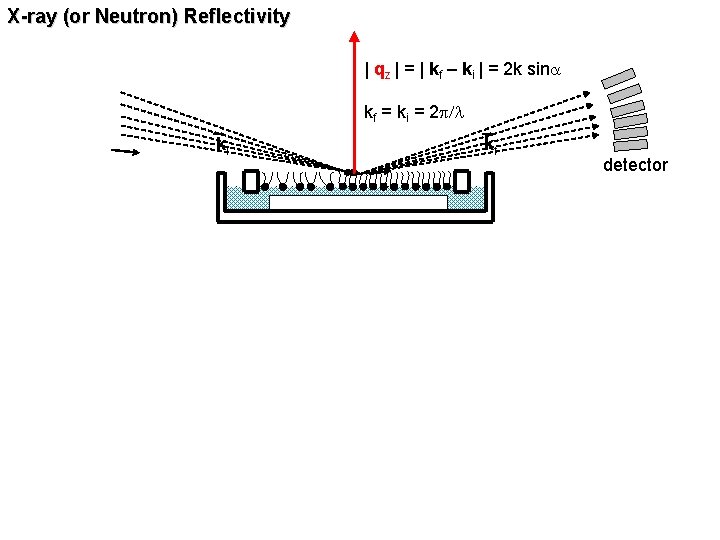
X-ray (or Neutron) Reflectivity | qz | = | kf – ki | = 2 k sina kf = ki = 2 p/l ki kf detector
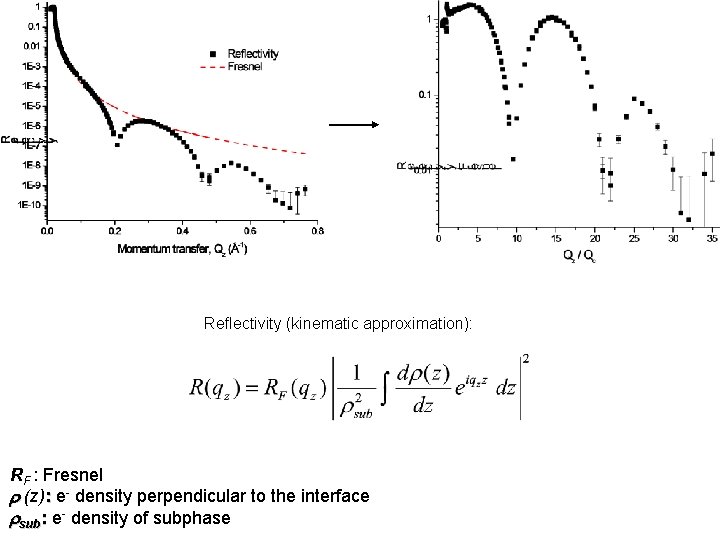
Reflectivity (kinematic approximation): RF : Fresnel r (z): e- density perpendicular to the interface rsub: e- density of subphase
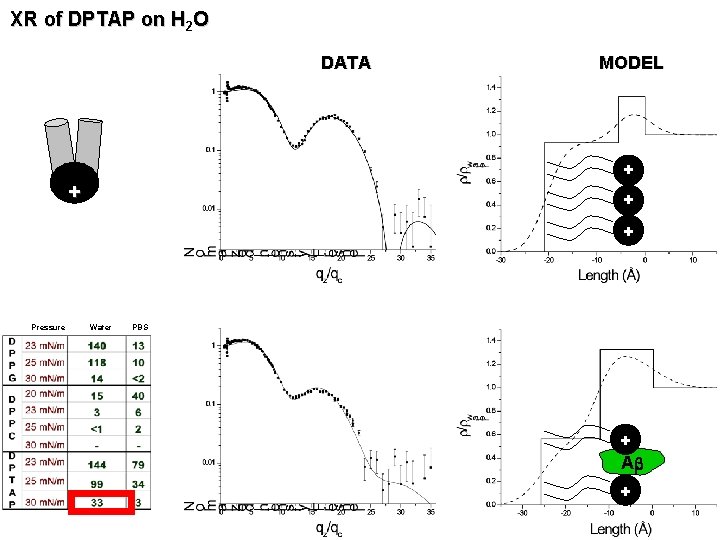
XR of DPTAP on H 2 O DATA MODEL + + Pressure Water PBS + Ab +
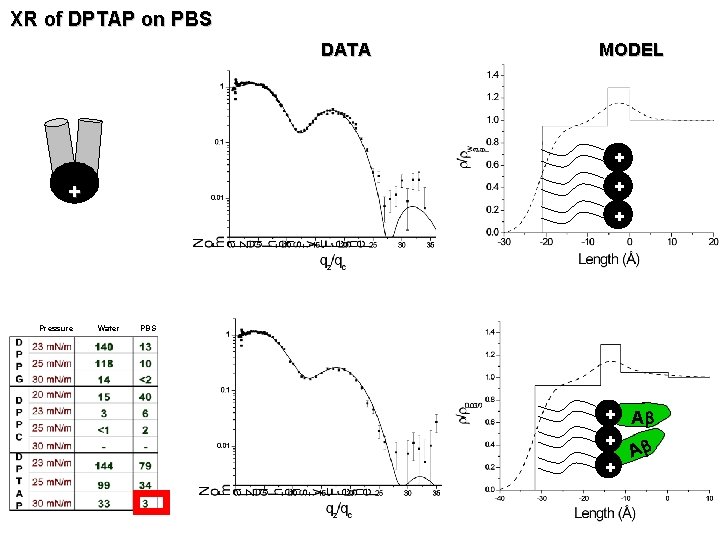
XR of DPTAP on PBS DATA MODEL + + Pressure Water PBS + Ab +
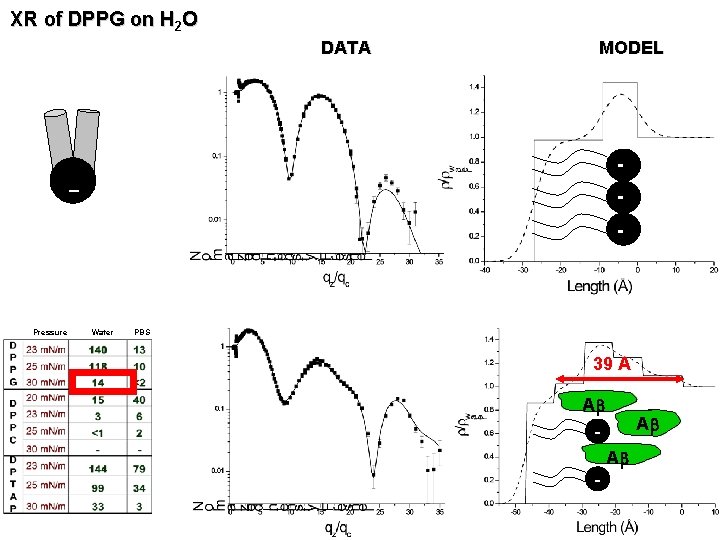
XR of DPPG on H 2 O DATA MODEL – - Pressure Water PBS 39 A Ab - Ab Ab -
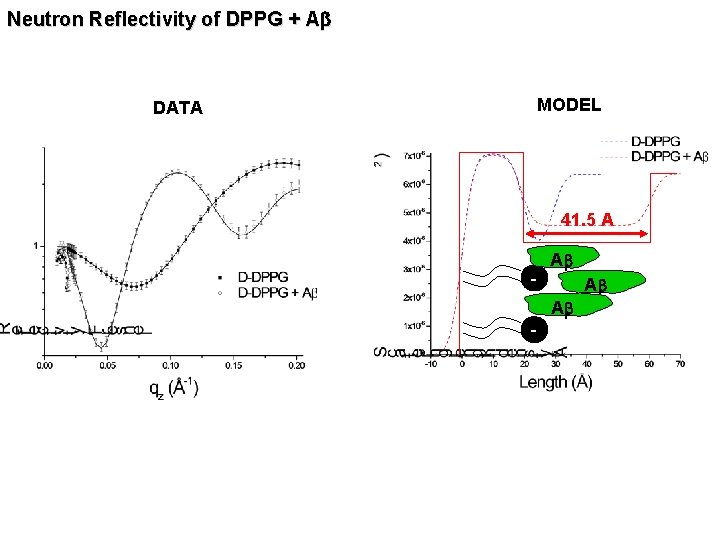
Neutron Reflectivity of DPPG + Ab DATA MODEL 41. 5 A - Ab Ab Ab -
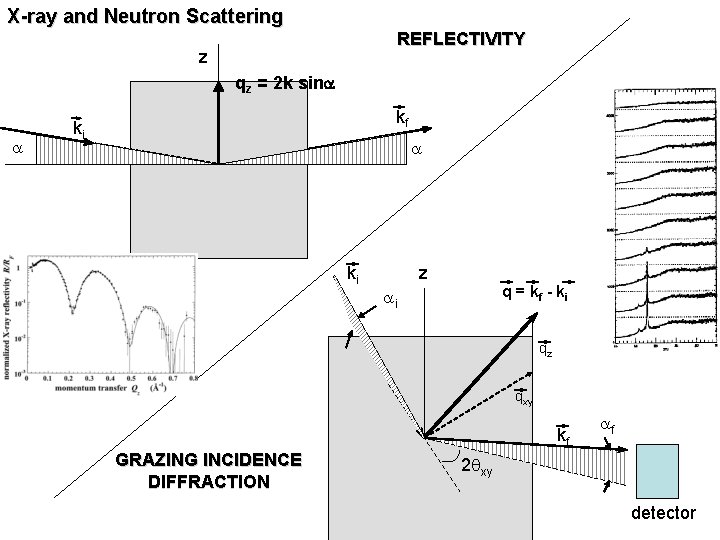
X-ray and Neutron Scattering REFLECTIVITY z qz = 2 k sina a kf ki a ki z q = kf - ki ai qz qxy kf GRAZING INCIDENCE DIFFRACTION af 2 qxy detector
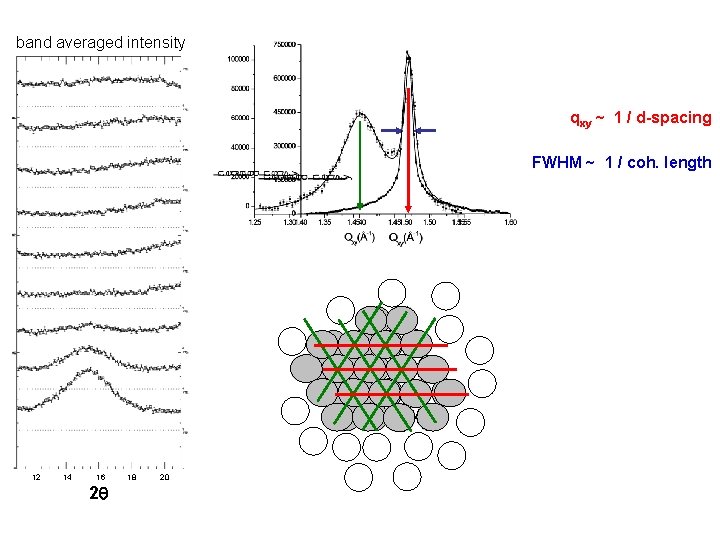
band averaged intensity qxy ~ 1 / d-spacing FWHM ~ 1 / coh. length 12 14 16 2 q 18 20
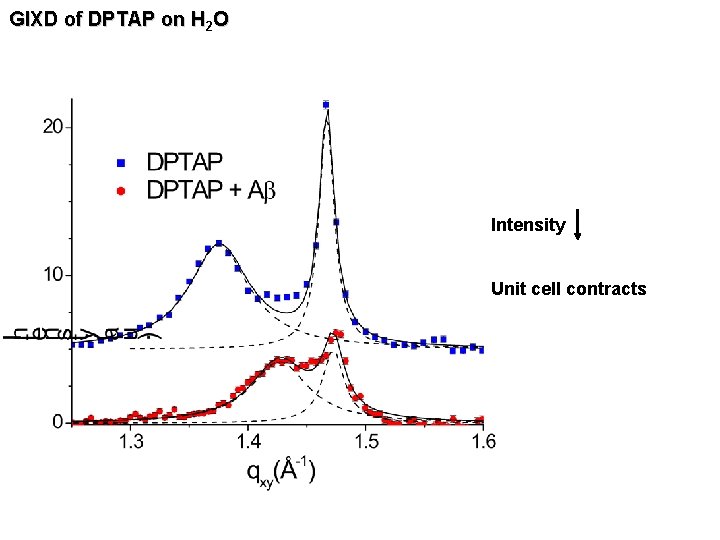
GIXD of DPTAP on H 2 O Intensity Unit cell contracts
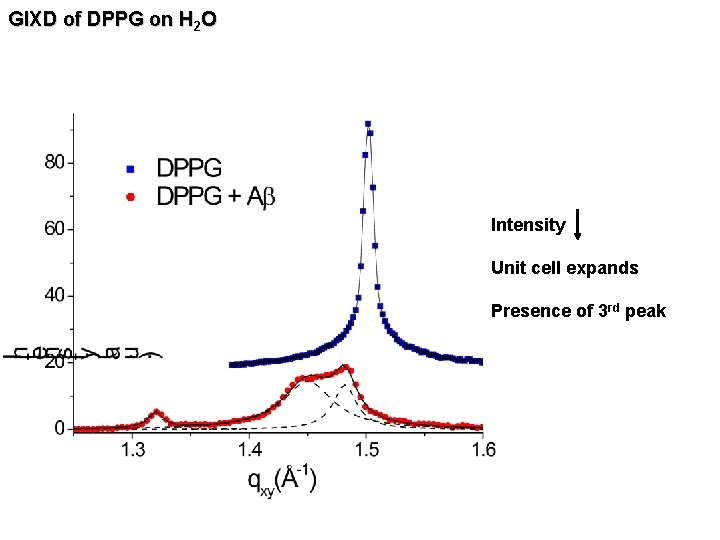
GIXD of DPPG on H 2 O Intensity Unit cell expands Presence of 3 rd peak
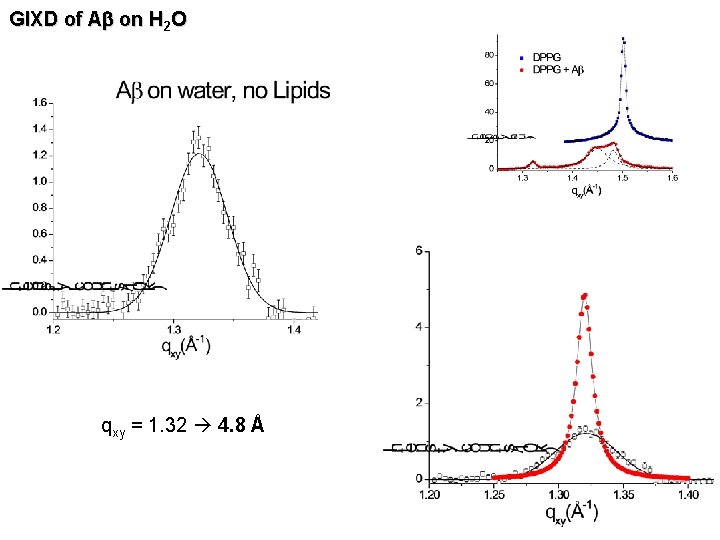
GIXD of Ab on H 2 O qxy = 1. 32 4. 8 Å
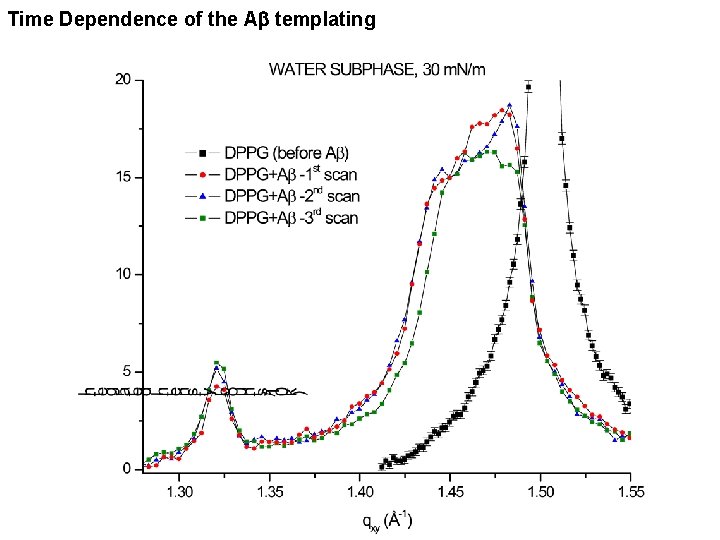
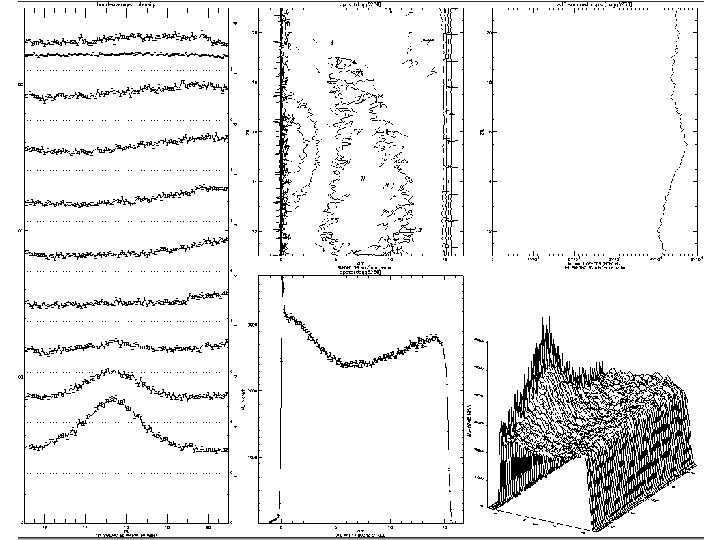
Time Dependence of the Ab templating
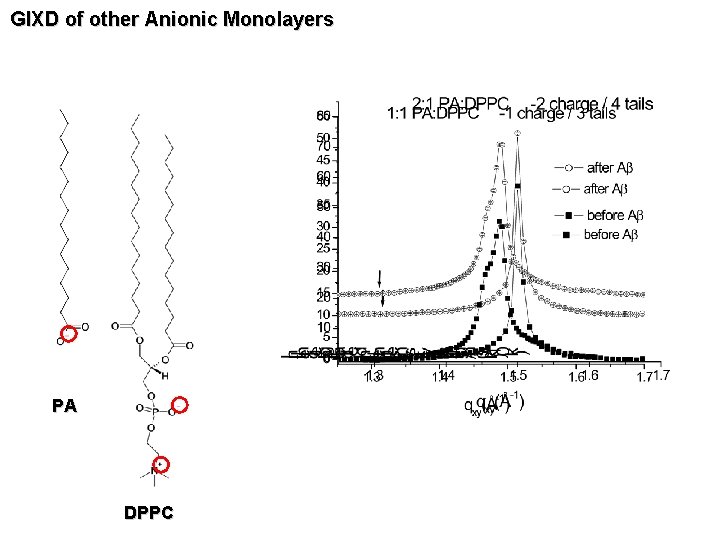
GIXD of other Anionic Monolayers PA DPPC
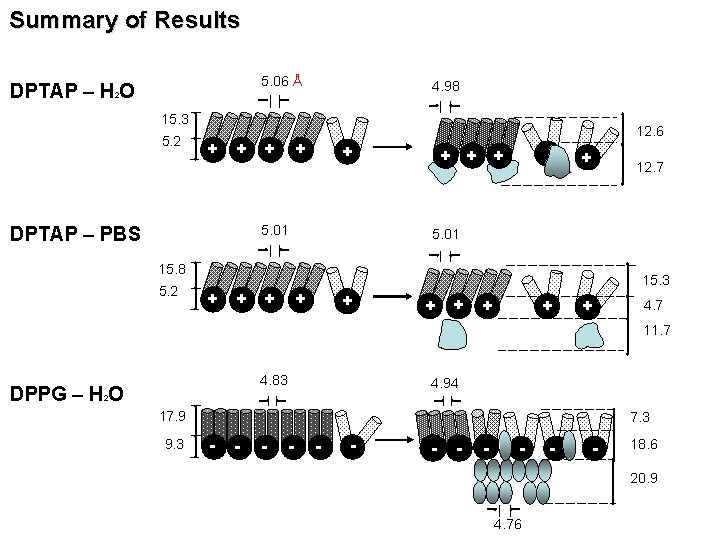
Summary of Results 5. 06 Å DPTAP – H O 4. 98 2 15. 3 5. 2 + + + 12. 6 + + 5. 01 DPTAP – PBS + + 12. 7 5. 01 15. 8 5. 2 15. 3 + + + + + 4. 7 11. 7 4. 83 DPPG – H O 4. 94 2 17. 9 9. 3 7. 3 - - - 18. 6 20. 9 4. 76
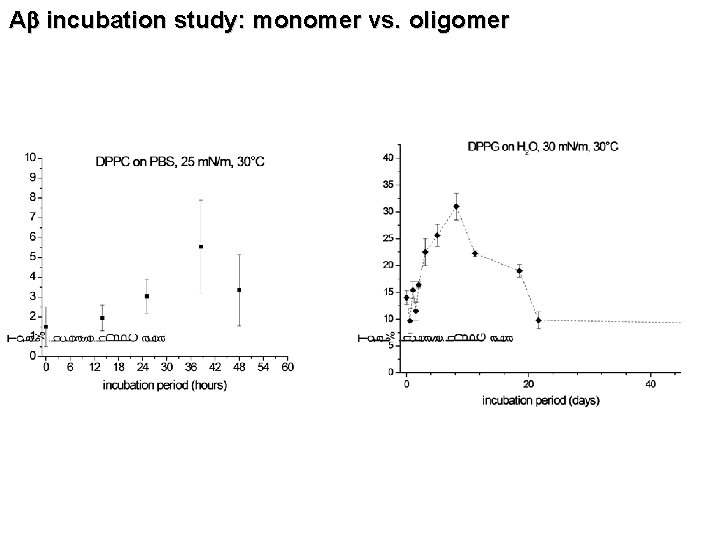
Ab incubation study: monomer vs. oligomer

CONCLUSIONS • Alzheimer’s Ab peptides inserts into lipid membranes to the point of destroying membrane morphology • Electrostatic arguments can be used to understand the insertion behavior • Anionic DPPG templates the ordering of Ab • Ab undergoes a rearrangement whereby it’s surface activity peaks at 8 days • SPECULATION: Negatively charged lipids, normally residing in the inner leaflet of the cell membrane get exposed to the surface, initiating the aggregation of Ab

FUTURE DIRECTIONS Peptide Interaction with Phosphatidyl Serine, an anionic lipid present physiologically Peptide interactions with membranes containing cholesterol and gangliosides Characterize the oligomerization of the peptide by measuring Ab surface activity, and size (via AFM) as a function of incubation time.

ACKNOWLEDGEMENTS Committee Members: Ka Yee Lee, James Norris, Stuart Rice Lee Group Members: (Dr. ) Ajay Gopal, Guohui Wu, Yuji Ishitsuka, Steve Danauskas, Maria Ratajczak, Kinlok Lam, Shelli Frey, Luka Pocivavsek, Mark Kittisopikul, Donna Wilson, Dr. Eva Chi. Past Members: Dr. Haim Diamant, Dr. David Gidalevitz, Dr. Josh Kurutz, Dr. Toan Nguyen, Dr. Adrian Muresan, Stacey Maskarinec, Katie Lemberg. Collaborators: Jarek Majewski (Los Alamos National Labs, NM), Kristian Kjaer (Risø National Labs, Denmark) Sushil Satija (NIST, MD) Fluorescently Labeled Peptides: Charles Glabe University of California, Irvine Funding: Alzheimer’s Association American Health Assistance Foundation MRSEC Packard Family: Parents, grandparents, sister, aunt Others: Sachin Rane, Fazil Zobu, Melita Morton, Aytek Erdil, Arash Bakhtyari, and Laura Syzmczyk
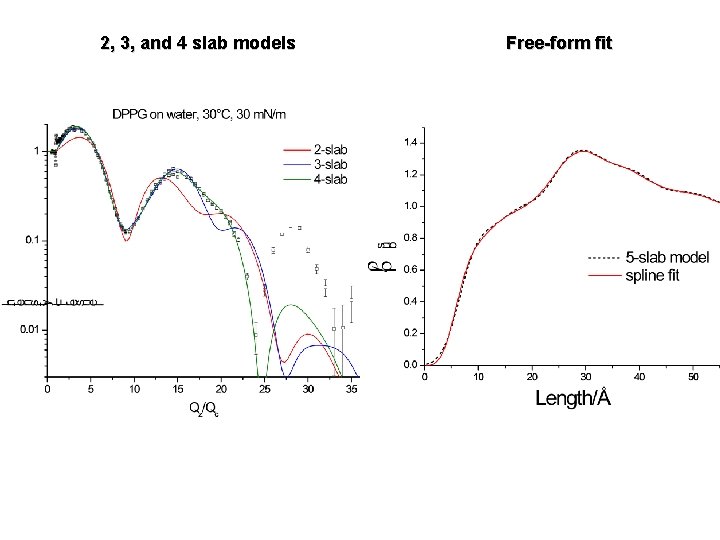
2, 3, and 4 slab models Free-form fit
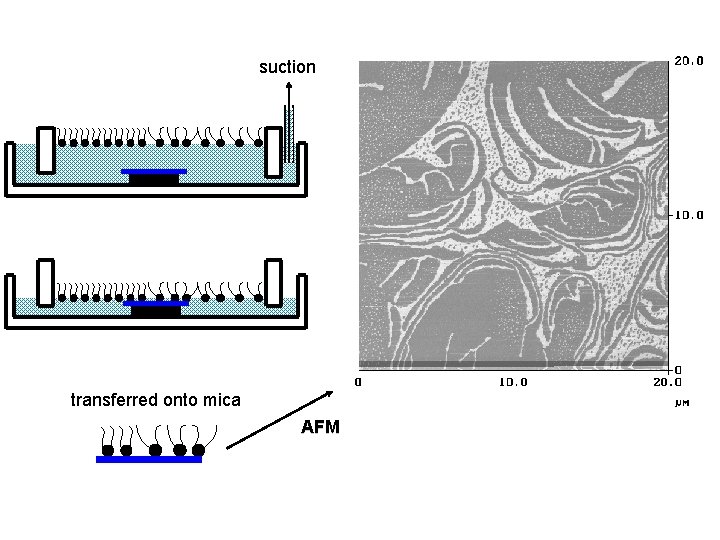
suction transferred onto mica AFM
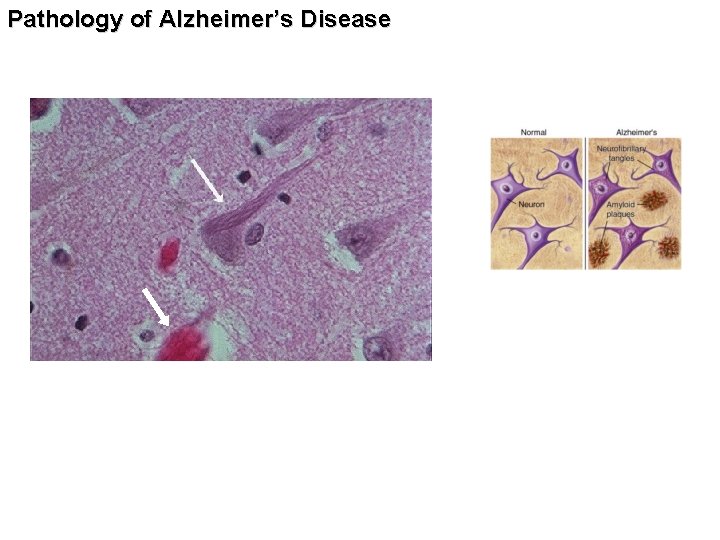
Pathology of Alzheimer’s Disease
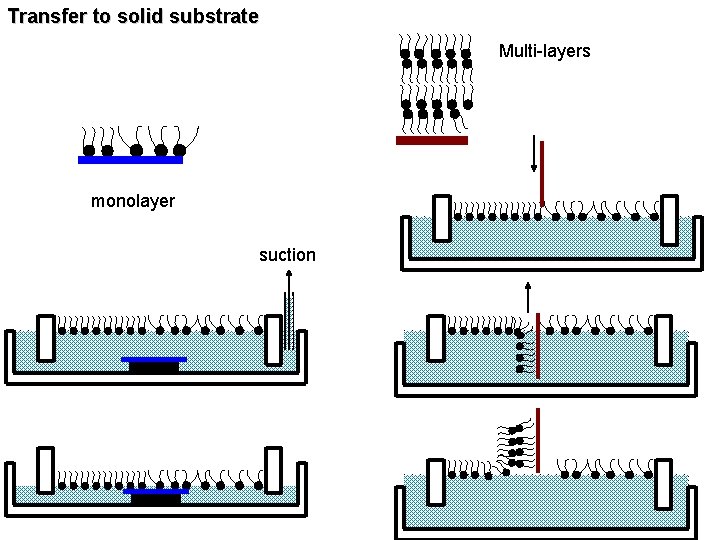
Transfer to solid substrate Multi-layers monolayer suction

DPTAP (30 m. N/m) DPPG (30 m. N/m) Dual probe fluorescence in WATER: 7
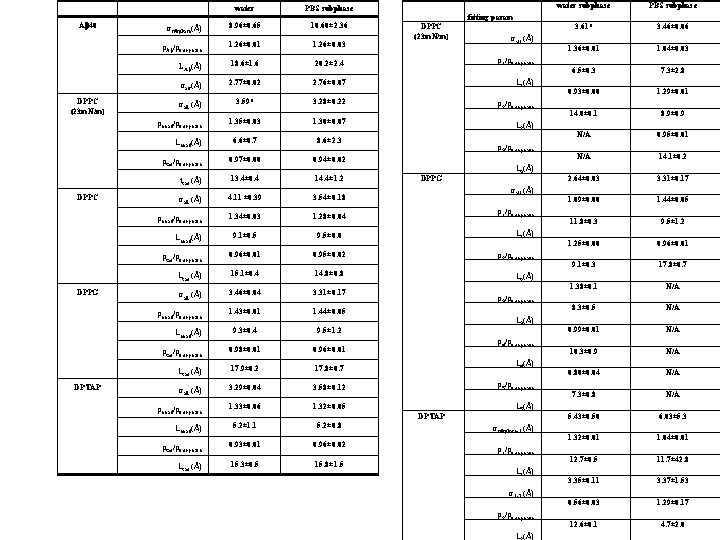
Ab 40 DPPC (23 m. N/m) water PBS subphase ssubphase(Å) 8. 96± 0. 65 10. 60± 2. 36 r. Ab/rsubphase 1. 26± 0. 01 18. 6± 1. 6 20. 2± 2. 4 sair(Å) 2. 77± 0. 02 2. 76± 0. 07 L 1(Å) sall (Å) 3. 59* 3. 28± 0. 22 r 2/rsubphase 1. 35± 0. 03 1. 30± 0. 07 6. 6± 0. 7 8. 6± 2. 3 rtail/rsubphase 0. 97± 0. 00 0. 94± 0. 02 ttail (Å) 13. 4± 0. 4 sall (Å) 4. 11 ± 0. 39 14. 4± 1. 2 L 2(Å) r 3/rsubphase DPPG L 3(Å) sall (Å) 3. 54± 0. 18 r 1/rsubphase 1. 34± 0. 03 1. 28± 0. 04 9. 1± 0. 5 9. 5± 0. 0 rtail/rsubphase 0. 96± 0. 01 0. 95± 0. 02 r 2/rsubphase Ltail (Å) 15. 1± 0. 4 14. 8± 0. 8 L 2(Å) sall (Å) 3. 46± 0. 04 3. 31± 0. 17 rhead/rsubphase 1. 43± 0. 01 1. 44± 0. 05 9. 3± 0. 4 9. 5± 1. 2 rhead/rsubphase Lhead(Å) rtail/rsubphase Ltail (Å) DPTAP sall (Å) LAb(Å) Lhead(Å) DPPG DPPC (23 m. N/m) r 1/rsubphase rhead/rsubphase DPPC 1. 26± 0. 03 0. 98± 0. 01 17. 9± 0. 2 L 1(Å) r 3/rsubphase L 3(Å) r 4/rsubphase 0. 96± 0. 01 17. 8± 0. 7 L 4(Å) sall (Å) 3. 29± 0. 04 3. 58± 0. 12 r 5/rsubphase rhead/rsubphase 1. 33± 0. 06 1. 32± 0. 05 L 5(Å) 5. 2± 1. 1 5. 2± 0. 8 rtail/rsubphase 0. 93± 0. 01 0. 96± 0. 02 Ltail (Å) 15. 3± 0. 5 15. 8± 1. 5 Lhead(Å) water subphase PBS subphase 3. 61* 3. 46± 0. 06 1. 36± 0. 01 1. 04± 0. 03 6. 5± 0. 3 7. 3± 2. 8 0. 93± 0. 00 1. 29± 0. 01 14. 0± 0. 1 8. 9± 0. 9 N/A 0. 95± 0. 01 N/A 14. 1± 0. 2 2. 64± 0. 03 3. 31± 0. 17 1. 09± 0. 00 1. 44± 0. 05 11. 8± 0. 3 9. 5± 1. 25± 0. 00 0. 96± 0. 01 9. 1± 0. 3 17. 8± 0. 7 1. 38± 0. 1 N/A 8. 3± 0. 5 N/A 0. 99± 0. 01 N/A 10. 3± 0. 9 N/A 0. 80± 0. 04 N/A 7. 3± 0. 8 N/A 5. 43± 0. 50 6. 03± 5. 3 1. 32± 0. 01 1. 04± 0. 01 12. 7± 0. 5 11. 7± 42. 8 3. 35± 0. 11 3. 37± 1. 53 0. 56± 0. 03 1. 29± 0. 17 12. 6± 0. 1 4. 7± 2. 0 fitting param DPTAP ssubphase-1 (Å) r 1/rsubphase L 1(Å) s 1 -2 (Å) r 2/rsubphase L (Å)
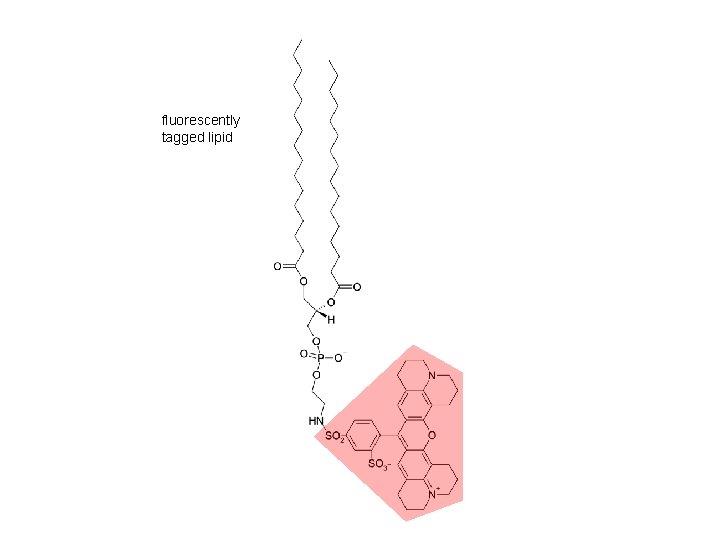
fluorescently tagged lipid
- Slides: 41