Teaching Genetic Linkage and Recombination through Mapping with

Teaching Genetic Linkage and Recombination through Mapping with Molecular Markers Lisa Mc. Donnell & Jennifer Klenz Note for instructors: Please see the notes field for comments that might be useful to you as an instructor when running this lesson.

Mapping with Molecular Markers Where is the drought tolerance locus?

Lesson Plan Part 1 • Assessing genetic linkage between a trait of interest and a molecular marker if we do a testcross (dihybrid F 1 x homozygous recessive). Part 2 • Predicting and assessing genetic linkage when we do a dihybrid x dihybrid cross (F 1 x F 1)

Learning Objectives • Explain how molecular markers (such as SNPs, microsatellites, VNTRs) can be used to map the location of genes/loci, including what crosses you would do, and why. • From analysis of data from a cross (such as gel banding patterns) determine if crosses involve linked genes. • Be able to justify your analysis by describing the information in the data that allows you to determine genes are linked. • Use linkage information between multiple loci to construct a genetic map

Brainstorm in your groups q Genes closer together on a chromosome are more often inherited together 1. If we know the location of some genes in a genome, how can we use that information to determine what DNA sequences cause specific traits of interest? 2. What are some examples of traits for which you (or someone) might want to know the location of a gene in the genome that controls that trait? 3. Why would you want to know the location of a gene? Write down your group answers and hand them in
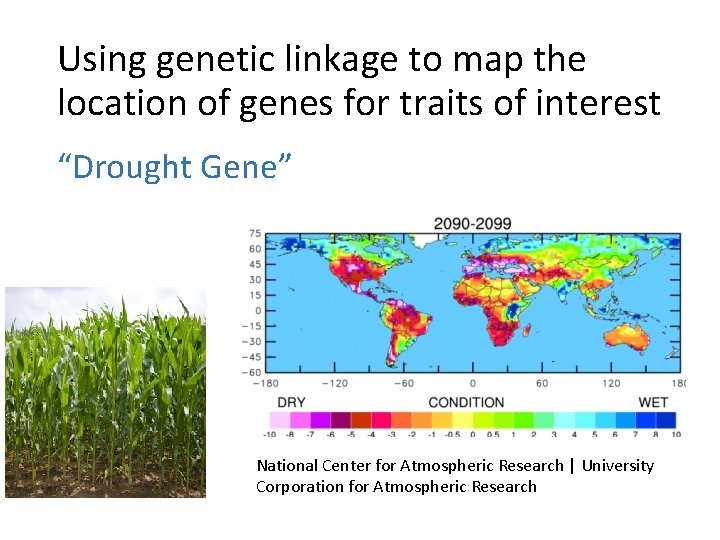
Using genetic linkage to map the location of genes for traits of interest “Drought Gene” National Center for Atmospheric Research | University Corporation for Atmospheric Research

Example: a drought tolerant plant. Most plants do not thrive in drought conditions. They may still reproduce, but they have lower yield.

Goals: 1) Is the drought tolerant phenotype heritable? 2) Is it caused by a single gene? 3) Mapping – where in the plant’s genome is the mutation/locus involved in drought tolerance?
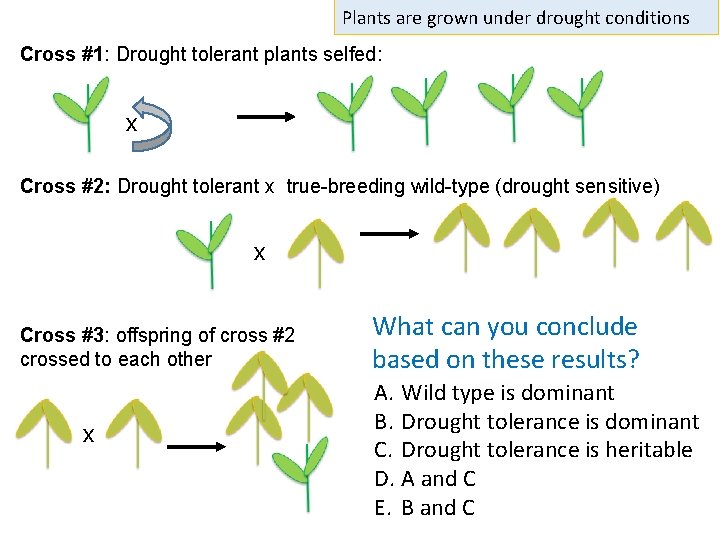
Plants are grown under drought conditions Cross #1: Drought tolerant plants selfed: x Cross #2: Drought tolerant x true-breeding wild-type (drought sensitive) x Cross #3: offspring of cross #2 crossed to each other x What can you conclude based on these results? A. Wild type is dominant B. Drought tolerance is dominant C. Drought tolerance is heritable D. A and C E. B and C

We have a true-breeding drought tolerant population, and we know drought tolerance is: – recessive to wildtype (sensitive) – caused by a single gene What do we not know?
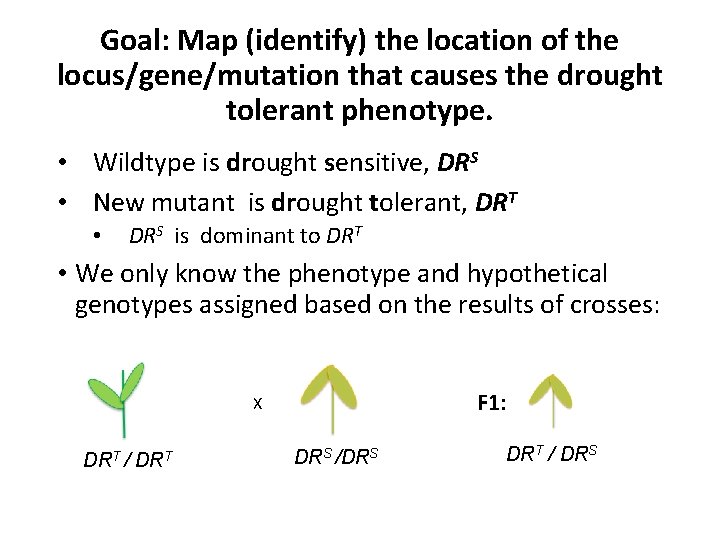
Goal: Map (identify) the location of the locus/gene/mutation that causes the drought tolerant phenotype. • Wildtype is drought sensitive, DRS • New mutant is drought tolerant, DRT • DRS is dominant to DRT • We only know the phenotype and hypothetical genotypes assigned based on the results of crosses: F 1: X DRT / DRT DRS /DRS DRT / DRS
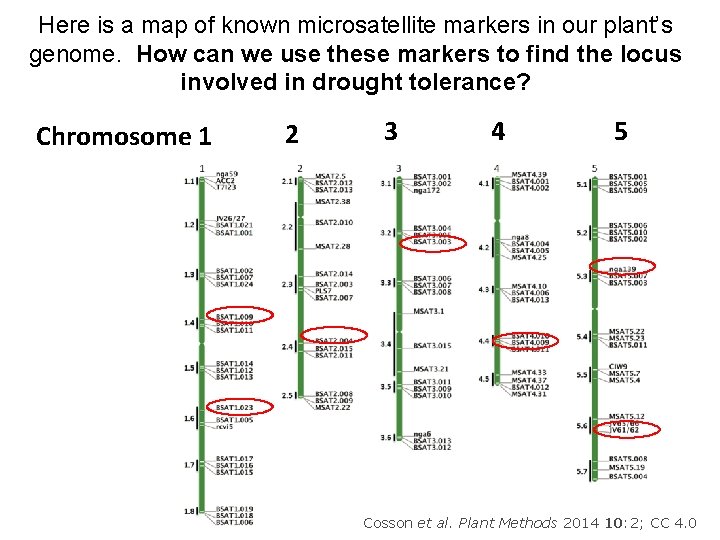
Here is a map of known microsatellite markers in our plant’s genome. How can we use these markers to find the locus involved in drought tolerance? Chromosome 1 2 3 4 5 12 Cosson et al. Plant Methods 2014 10: 2; CC 4. 0
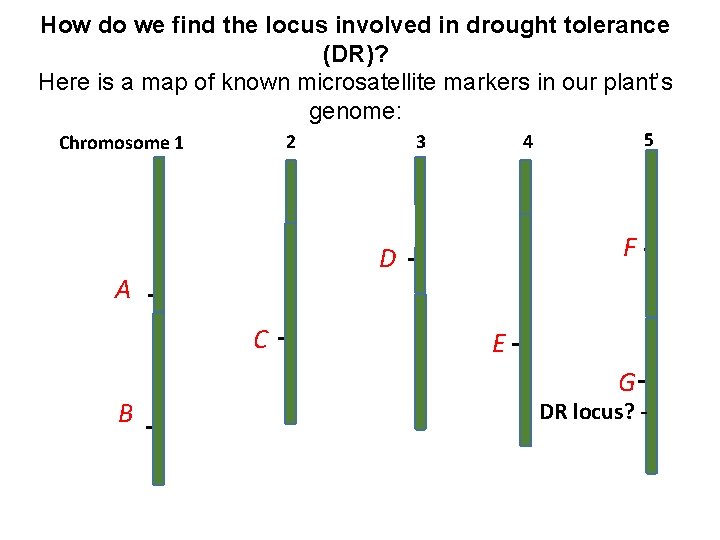
How do we find the locus involved in drought tolerance (DR)? Here is a map of known microsatellite markers in our plant’s genome: Chromosome 1 DR locus? 3 C DR locus? 5 4 DR locus? D A B 2 F E G DR locus? -

Does the drought tolerance phenotype segregate with a the G microsatellite locus? To measure segregation/linkage between 2 loci what do we look for? Frequency of parental and recombinant combinations of genotypes/phenotypes. For our experiment, this means segregation of drought tolerance with a G microsatellite allele (a specific banding pattern). We need two “parental” combinations of alleles: 1) Drought tolerance with a G microsatellite allele that gives one banding pattern 2) Drought sensitive with a different G microsatellite allele that gives a different banding pattern G microsatellite Image adapted from Cosson et al. Plant Methods 2014 10: 2; CC 4. 0

Is the drought tolerance locus linked to the G microsatellite? Experimental plan: a testcross to measure linkage 1. Identify a true-breeding drought sensitive strain that carries different microsatellite alleles (depending on the species these are available for purchase form research groups) 2. Generate a heterozygous plant, DRT G 200 /DRS G 400 3. Set up a testcross: heterozygote x homozygous recessive 4. Analyze offspring of the testcross, look for segregation of the drought tolerance phenotype with G 200 and G 400 alleles.
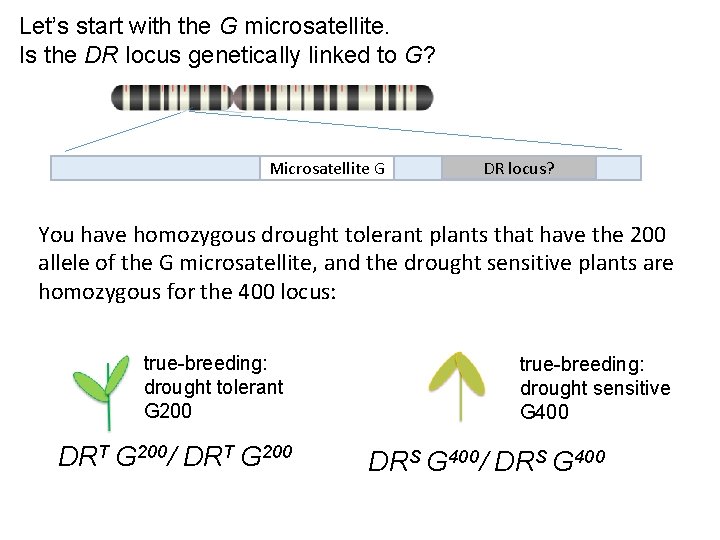
Let’s start with the G microsatellite. Is the DR locus genetically linked to G? Microsatellite G DR locus? You have homozygous drought tolerant plants that have the 200 allele of the G microsatellite, and the drought sensitive plants are homozygous for the 400 locus: true-breeding: drought tolerant G 200 DRT G 200/ DRT G 200 true-breeding: drought sensitive G 400 DRS G 400/ DRS G 400

Is the drought tolerance locus linked to the G microsatellite? Parent plants: Drought tolerant Drought sensitive DRTG 200 / DRTG 200 DRS G 400 / DRSG 400 Ultimately we need to look for recombination (or independent assortment) between the DR and G loci. What crosses will we do next?
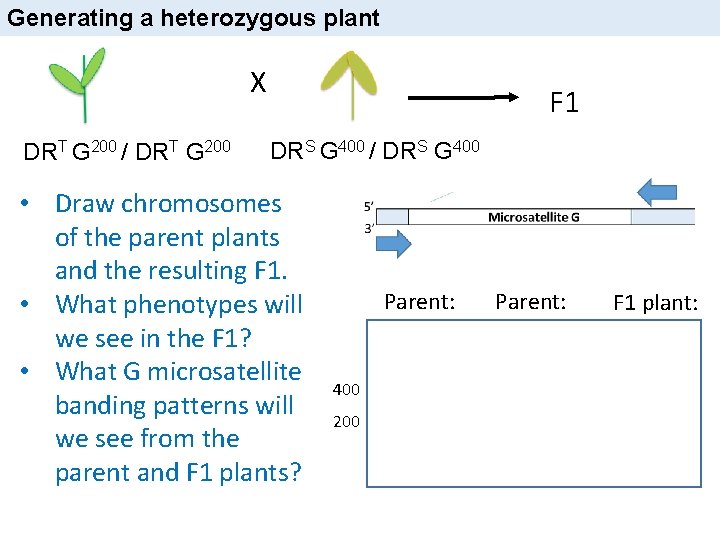
Generating a heterozygous plant X DRT G 200 / DRT G 200 F 1 DRS G 400 / DRS G 400 • Draw chromosomes of the parent plants and the resulting F 1. • What phenotypes will we see in the F 1? • What G microsatellite banding patterns will we see from the parent and F 1 plants? Parent: 400 200 Parent: F 1 plant:
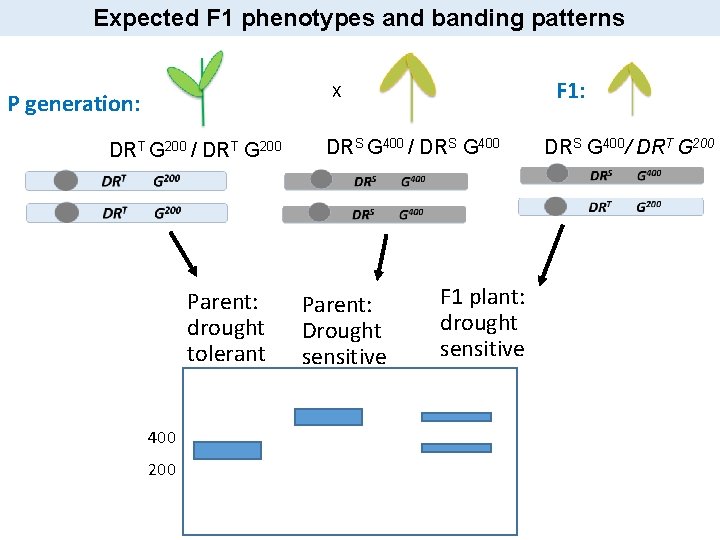
Expected F 1 phenotypes and banding patterns F 1: X P generation: DRT G 200 / DRT G 200 Parent: drought tolerant 400 200 DRS G 400 / DRS G 400 Parent: Drought sensitive F 1 plant: drought sensitive DRS G 400/ DRT G 200
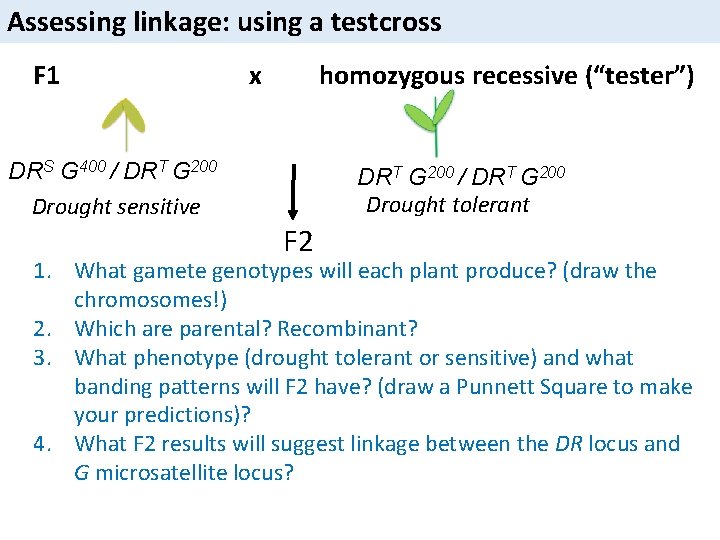
Assessing linkage: using a testcross F 1 x homozygous recessive (“tester”) DRS G 400 / DRT G 200 Drought sensitive DRT G 200 / DRT G 200 Drought tolerant F 2 1. What gamete genotypes will each plant produce? (draw the chromosomes!) 2. Which are parental? Recombinant? 3. What phenotype (drought tolerant or sensitive) and what banding patterns will F 2 have? (draw a Punnett Square to make your predictions)? 4. What F 2 results will suggest linkage between the DR locus and G microsatellite locus?
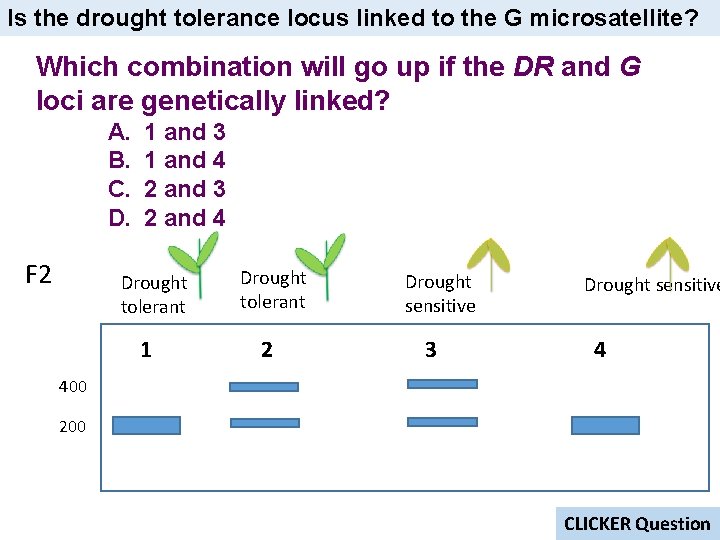
Is the drought tolerance locus linked to the G microsatellite? Which combination will go up if the DR and G loci are genetically linked? A. B. C. D. F 2 1 and 3 1 and 4 2 and 3 2 and 4 Drought tolerant 1 Drought tolerant 2 Drought sensitive 3 Drought sensitive 4 400 200 CLICKER Question
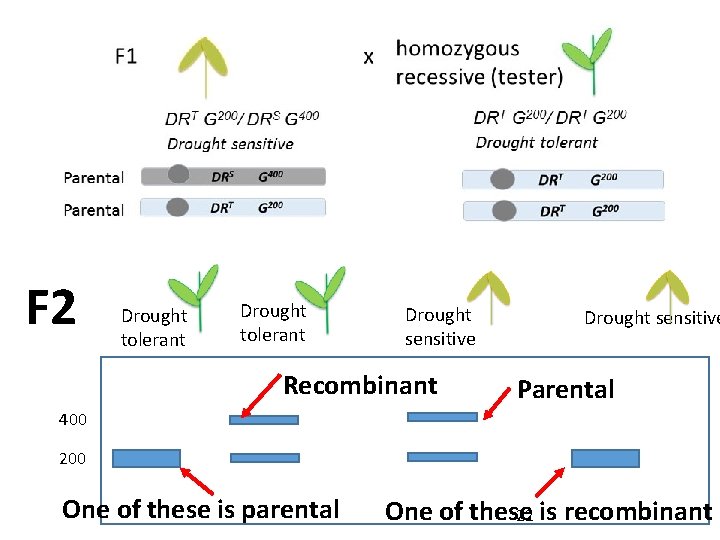
F 2 Drought tolerant Drought sensitive Recombinant Drought sensitive Parental 400 200 One of these is parental One of these 22 is recombinant
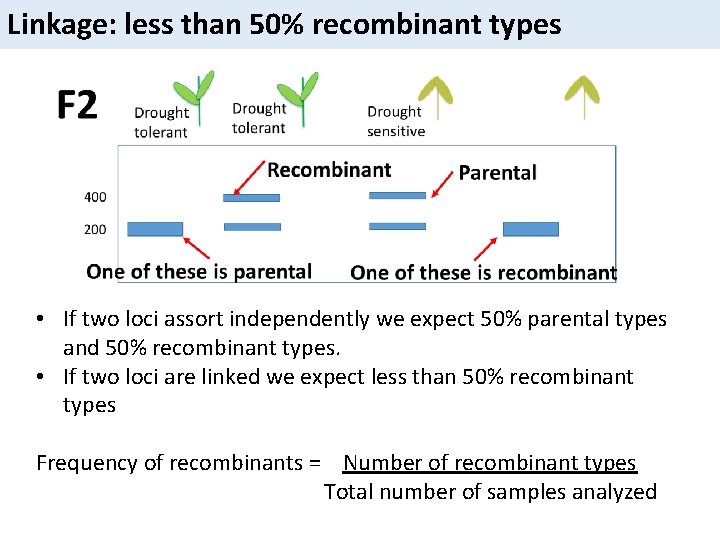
Linkage: less than 50% recombinant types • If two loci assort independently we expect 50% parental types and 50% recombinant types. • If two loci are linked we expect less than 50% recombinant types Frequency of recombinants = Number of recombinant types Total number of samples analyzed

Is the drought tolerance locus linked to the G microsatellite? Now that we’ve made predictions, let’s analyze some data Our testcross scenario: F 1 x homozygous recessive F 2 Examine the four gels and determine which (if any) indicates the drought tolerance locus is genetically linked to the G microsatellite? (gels are on the next two slides) What is the map distance?
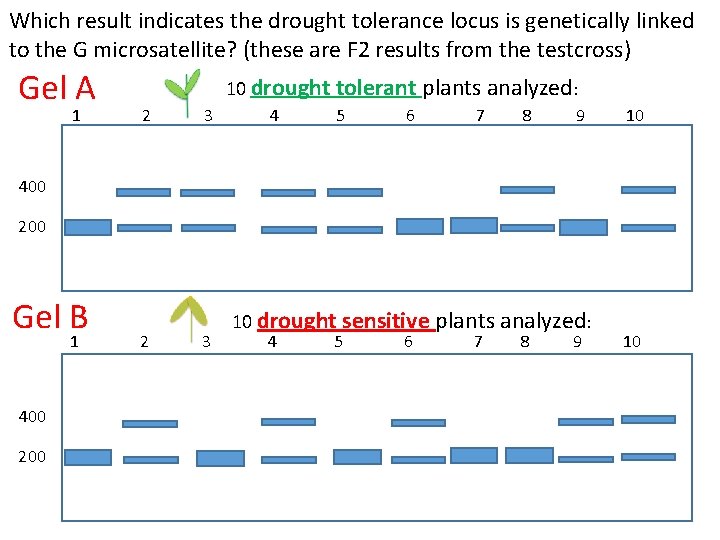
Which result indicates the drought tolerance locus is genetically linked to the G microsatellite? (these are F 2 results from the testcross) Gel A 1 10 drought tolerant plants analyzed: 2 3 4 5 6 7 8 9 10 400 200 Gel B 1 400 2 3 10 drought sensitive plants analyzed: 4 5 6 7 8 9 10
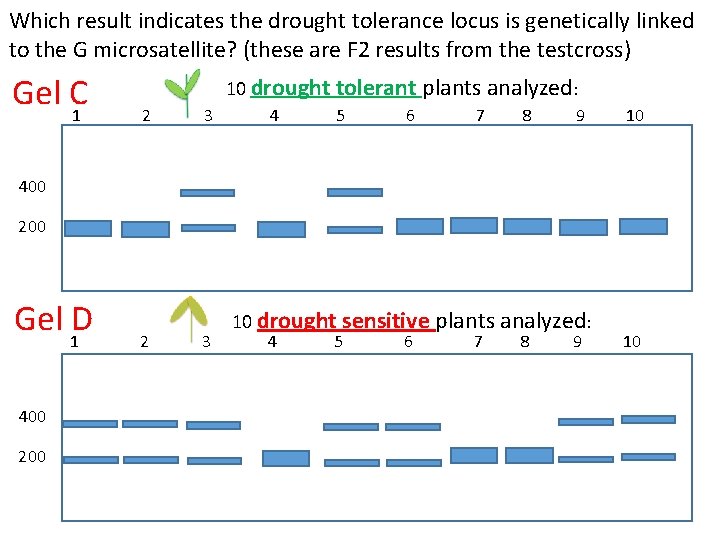
Which result indicates the drought tolerance locus is genetically linked to the G microsatellite? (these are F 2 results from the testcross) Gel C 1 10 drought tolerant plants analyzed: 2 3 4 5 6 7 8 9 10 400 200 Gel 1 D 400 2 3 10 drought sensitive plants analyzed: 4 5 6 7 8 9 10

Is the drought tolerance locus linked to the G microsatellite? F 1 x tester F 2 From your analysis of the four gels, which (if any) indicates the drought tolerance locus is genetically linked to the G microsatellite locus. A. B. C. D. E. Gels A & B Gels C & D Gels A & D Gels B & C None of the results indicate linkage What is the map distance? CLICKER Question

Summary Always determine what allele combination is considered parental, and which is recombinant. Draw the chromosomes of the parents and F 1 to be sure you keep track of information correctly. If you do an F 1 testcross you can analyze the entire F 2 population to determine linkage. You always know what gamete genotype the tester donates, so the F 2 phenotype proportions are representative of the recombination events (and frequencies) that happened in the F 1.

Part 2: Determining linkage when we do a dihybrid cross
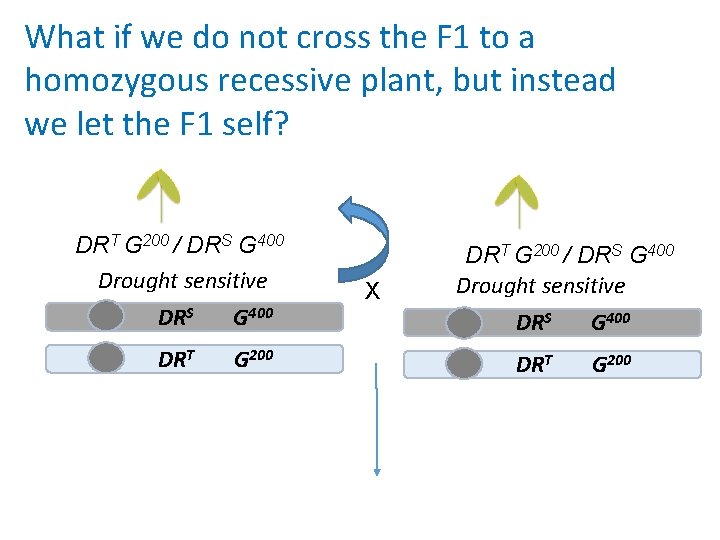
What if we do not cross the F 1 to a homozygous recessive plant, but instead we let the F 1 self? DRT G 200 / DRS G 400 Drought sensitive DRS G 400 DRT G 200 X DRT G 200 / DRS G 400 Drought sensitive DRS G 400 DRT G 200
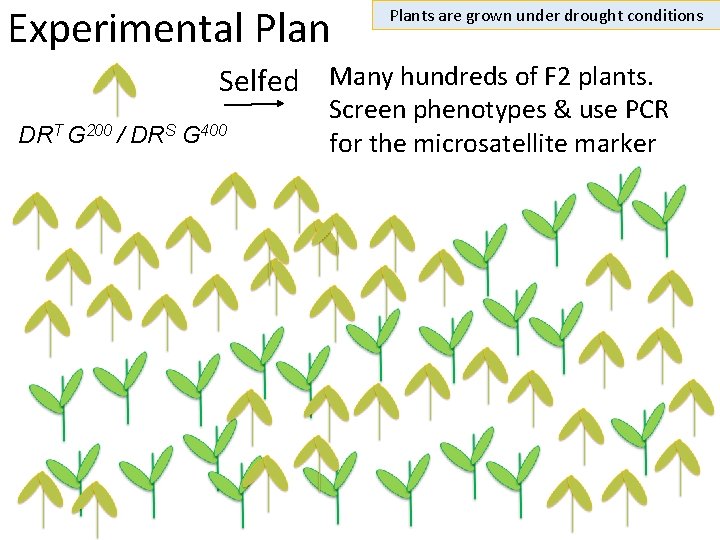
Experimental Plants are grown under drought conditions Selfed Many hundreds of F 2 plants. DRT G 200 / DRS G 400 Screen phenotypes & use PCR for the microsatellite marker

Insert a picture of a growth chamber, such as the picture found at: http: //www. rochmec hatronics. com/Humid ity-enviornmentalchamber. html
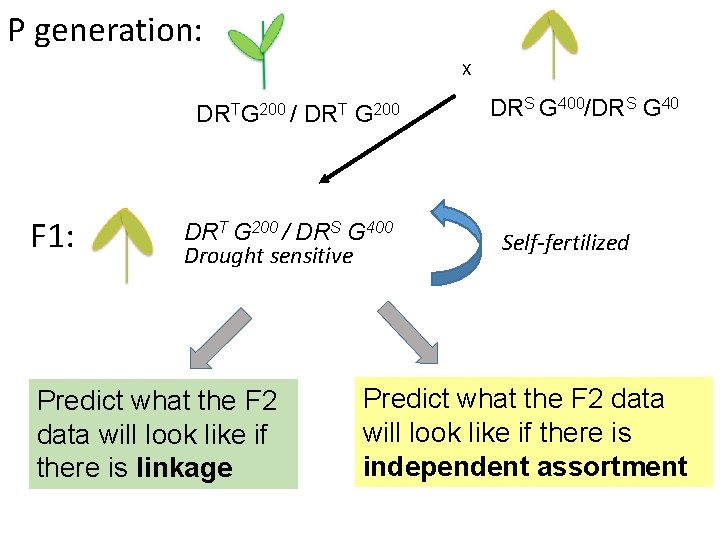
P generation: X DRTG 200 / DRT G 200 F 1: DRT G 200 / DRS G 400 Drought sensitive Predict what the F 2 data will look like if there is linkage DRS G 400/DRS G 40 Self-fertilized Predict what the F 2 data will look like if there is independent assortment

Prediction Activity Students assemble into groups of 6 and then break into 2 mini-groups: 3 people in each mini-group. You have 3 people will “predict what the F 2 will look like, and banding patterns, if marker and DR locus are linked” Class Slide 1 15 minut e s! Each group, on the flip chart 3 people will predict paper: draw gametes, what the F 2 will look chromosomes with alleles like, and banding labelled, banding patterns on patterns, if marker a gel. Put your work on the and DR locus are not wall when you are done. linked (assort COMPLETE HANDOUT A independently) Come back together in your group of 6. Teach other about your predictions and identify how they are different, and how they are different from when we did an F 1 x tester cross. COMPLETE HANDBOUT B.
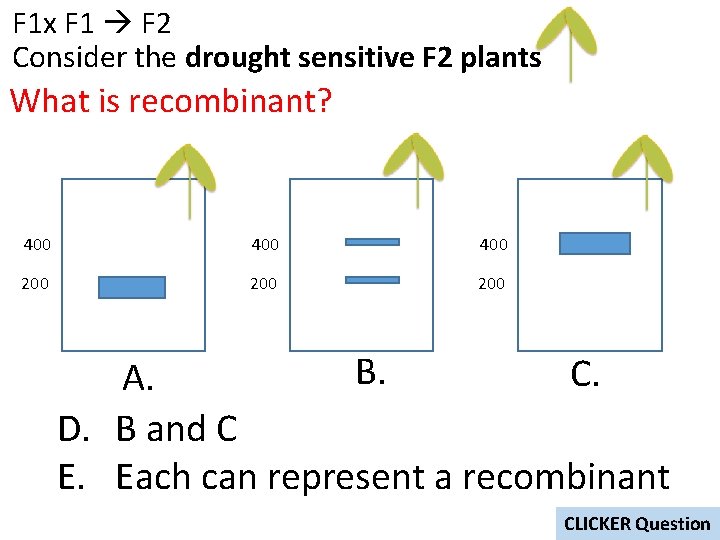
F 1 x F 1 F 2 Consider the drought sensitive F 2 plants What is recombinant? 400 400 200 200 B. C. A. D. B and C E. Each can represent a recombinant CLICKER Question
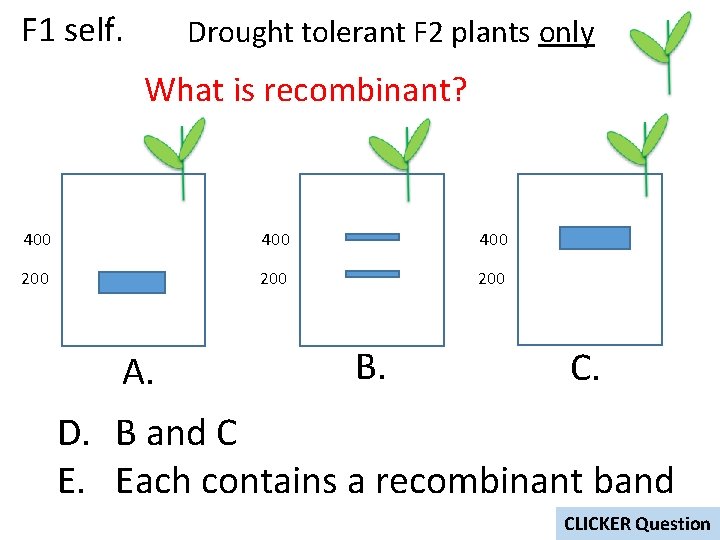
F 1 self. Drought tolerant F 2 plants only What is recombinant? 400 400 200 200 A. B. C. D. B and C E. Each contains a recombinant band CLICKER Question

Which plants would you use to test for genetic linkage? A. The entire F 2 population B. The F 2 drought tolerant plants only C. The F 2 drought sensitive plants only CLICKER Question
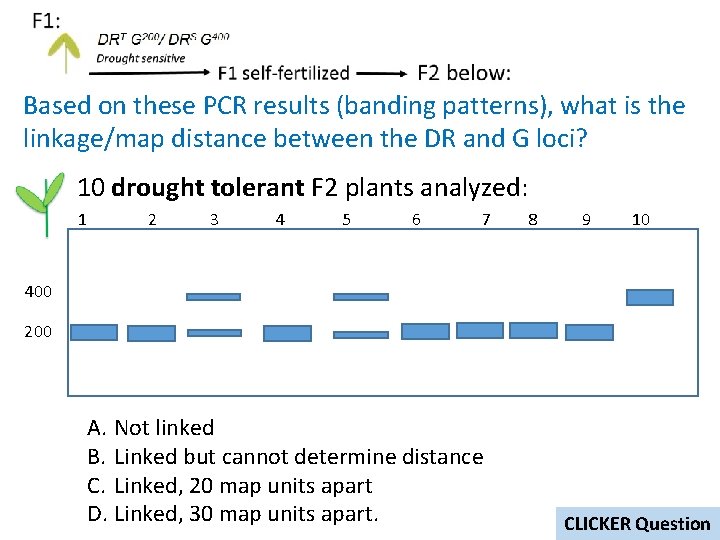
Based on these PCR results (banding patterns), what is the linkage/map distance between the DR and G loci? 10 drought tolerant F 2 plants analyzed: 1 2 3 4 5 6 7 8 9 10 400 200 A. Not linked B. Linked but cannot determine distance C. Linked, 20 map units apart D. Linked, 30 map units apart. CLICKER Question
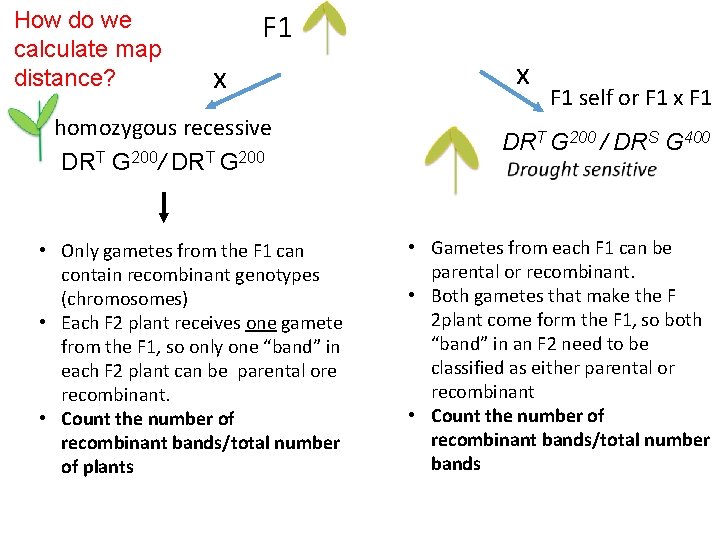
How do we calculate map distance? F 1 x homozygous recessive DRT G 200/ DRT G 200 • Only gametes from the F 1 can contain recombinant genotypes (chromosomes) • Each F 2 plant receives one gamete from the F 1, so only one “band” in each F 2 plant can be parental ore recombinant. • Count the number of recombinant bands/total number of plants x F 1 self or F 1 x F 1 DRT G 200 / DRS G 400 • Gametes from each F 1 can be parental or recombinant. • Both gametes that make the F 2 plant come form the F 1, so both “band” in an F 2 need to be classified as either parental or recombinant • Count the number of recombinant bands/total number bands
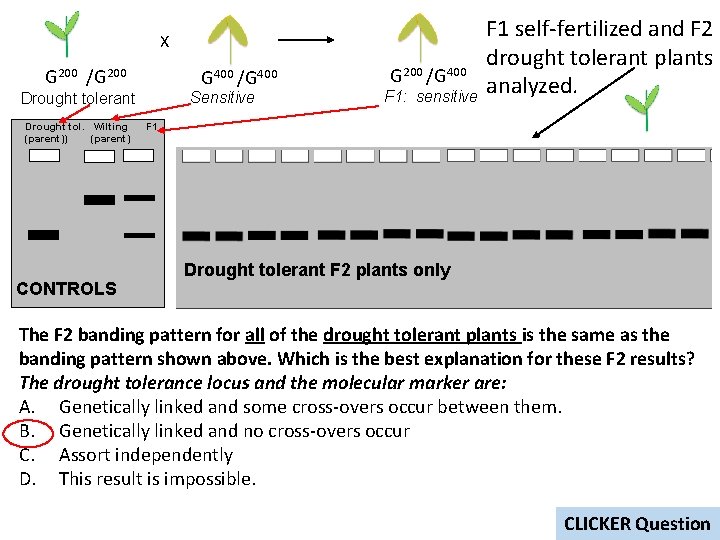
X G 200 /G 200 G 400 /G 400 Sensitive Drought tolerant Drought tol. Wilting (parent)) (parent) CONTROLS G 200 /G 400 F 1: sensitive F 1 self-fertilized and F 2 drought tolerant plants analyzed. F 1 Drought tolerant F 2 plants only The F 2 banding pattern for all of the drought tolerant plants is the same as the banding pattern shown above. Which is the best explanation for these F 2 results? The drought tolerance locus and the molecular marker are: A. Genetically linked and some cross-overs occur between them. B. Genetically linked and no cross-overs occur C. Assort independently D. This result is impossible. CLICKER Question

Practice Assessing Linkage by analyzing results of an F 1 x F 1 cross New trait of interest: potatoes that are resistant to the Colorado Potato Beetle. a pest that destroys millions of dollars of potato crops each year (and the beetle is resistant to most insecticides) You have found a potato plant that potato beetles do not attack. Is the plant producing something that deters attack from the beetle (or kills the beetle)? © 2008, Pilise Gábor. Creative Commons 3. 0
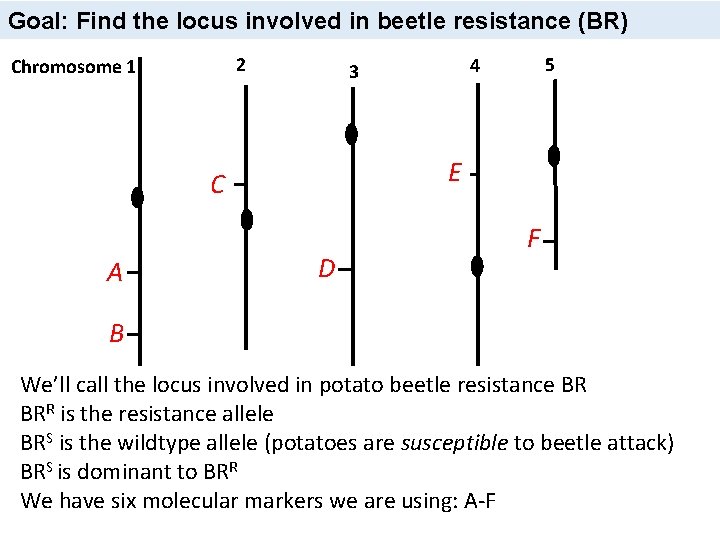
Goal: Find the locus involved in beetle resistance (BR) 2 Chromosome 1 E C A 5 4 3 D F B We’ll call the locus involved in potato beetle resistance BR BRR is the resistance allele BRS is the wildtype allele (potatoes are susceptible to beetle attack) BRS is dominant to BRR We have six molecular markers we are using: A-F 43
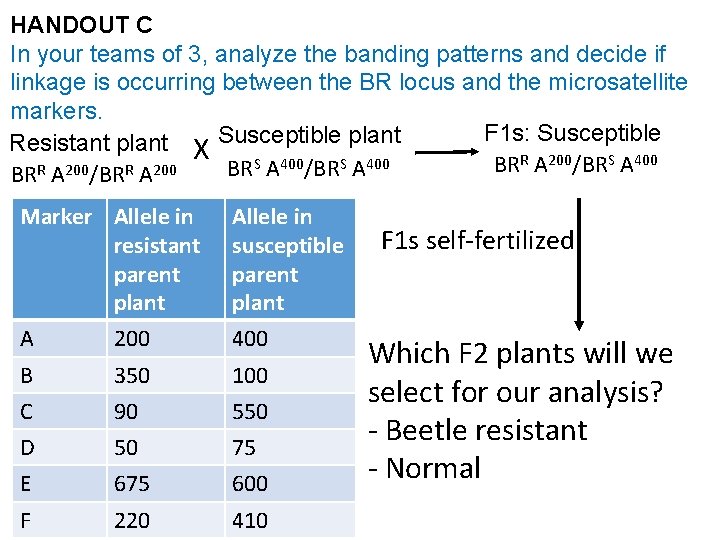
HANDOUT C In your teams of 3, analyze the banding patterns and decide if linkage is occurring between the BR locus and the microsatellite markers. F 1 s: Susceptible Resistant plant X Susceptible plant R A 200/BRS A 400/BRS A 400 BR R 200 BR BR A /BR A Marker Allele in resistant parent plant A 200 Allele in susceptible parent plant 400 B 350 100 C 90 550 D 50 75 E 675 600 F 220 410 F 1 s self-fertilized Which F 2 plants will we select for our analysis? - Beetle resistant - Normal

Use handouts with gel data from different markers to determine the location • Determine linkage • Calculate map distance • First group of 3 with the correct answer wins bragging rights!
For your interest: What do you do next? If the genome for this plant is sequenced: Look at the genes in this approximate location. At a specific map location there could be anywhere from 20 -500 genes depending on the genome. Depending on the phenotype of interest it could be difficult or easy to make educated guesses as to possible target genes. A B BR, Beetle resistant locus For species like Arabidopsis there are databases with additional sources of helpful information such as whether any gene is expressed under specific conditions, including attack by pests such as the beetle. Genes in this region could be checked to see what their wild-type expression pattern looks like under pest attack. You could compare the DNA sequence of your beetle resistant plant to wildtype for your most likely candidate genes to look for sequence differences causing the mutant phenotype. You could take a wild copy of a likely candidate gene and add it to the mutant to see if the wild type phenotype is restored. 46
- Slides: 45