Table 6 1 Comparison of MBL and MASP
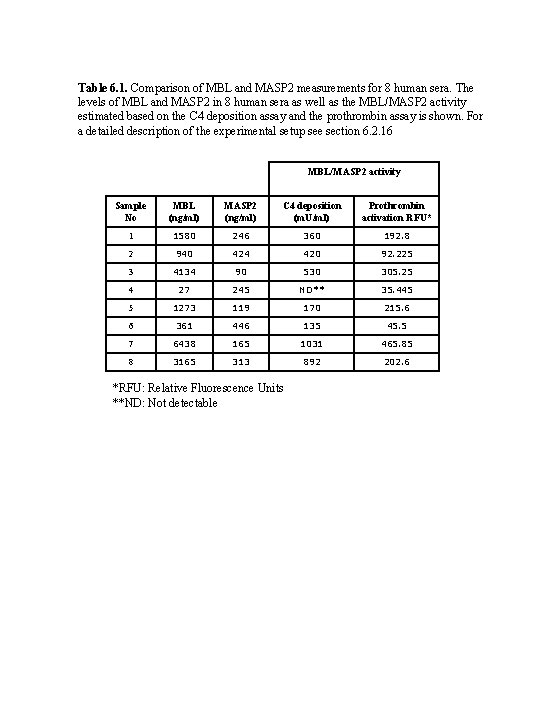
Table 6. 1. Comparison of MBL and MASP 2 measurements for 8 human sera. The levels of MBL and MASP 2 in 8 human sera as well as the MBL/MASP 2 activity estimated based on the C 4 deposition assay and the prothrombin assay is shown. For a detailed description of the experimental setup see section 6. 2. 16 MBL/MASP 2 activity Sample No MBL (ng/ml) MASP 2 (ng/ml) C 4 deposition (m. U/ml) Prothrombin activation RFU* 1 1580 246 360 192. 8 2 940 424 420 92. 225 3 4134 90 530 305. 25 4 27 245 ND** 35. 445 5 1273 119 170 215. 6 6 361 446 135 45. 5 7 6438 165 1031 465. 85 8 3165 313 892 202. 6 *RFU: Relative Fluorescence Units **ND: Not detectable
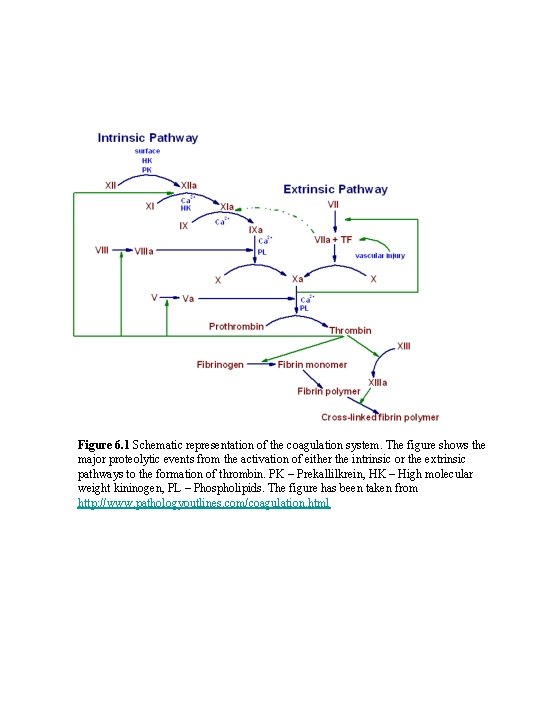
Figure 6. 1 Schematic representation of the coagulation system. The figure shows the major proteolytic events from the activation of either the intrinsic or the extrinsic pathways to the formation of thrombin. PK – Prekallilkrein, HK – High molecular weight kininogen, PL – Phospholipids. The figure has been taken from http: //www. pathologyoutlines. com/coagulation. html
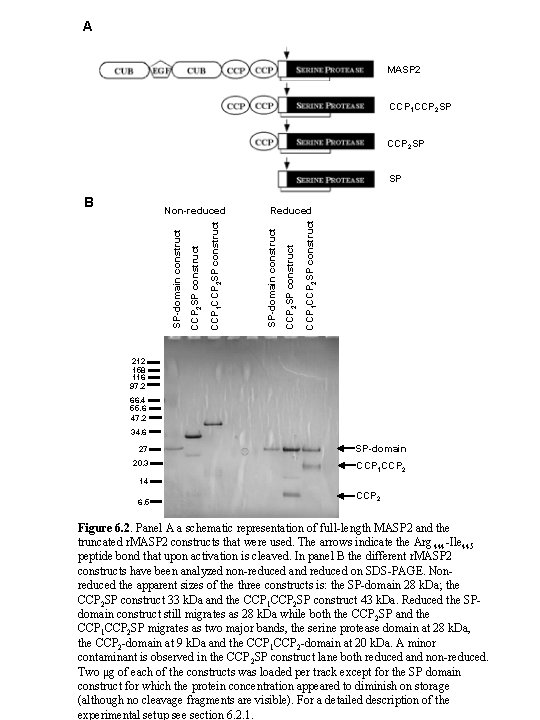
A MASP 2 CCP 1 CCP 2 SP SP CCP 1 CCP 2 SP construct Reduced CCP 2 SP construct CCP 1 CCP 2 SP construct SP-domain construct Non-reduced SP-domain construct B 212 158 116 97. 2 66. 4 55. 6 47. 2 34. 6 27 SP-domain 20. 3 CCP 1 CCP 2 14 6. 5 CCP 2 Figure 6. 2. Panel A a schematic representation of full-length MASP 2 and the truncated r. MASP 2 constructs that were used. The arrows indicate the Arg 444 -Ile 445 peptide bond that upon activation is cleaved. In panel B the different r. MASP 2 constructs have been analyzed non-reduced and reduced on SDS-PAGE. Nonreduced the apparent sizes of the three constructs is: the SP-domain 28 k. Da; the CCP 2 SP construct 33 k. Da and the CCP 1 CCP 2 SP construct 43 k. Da. Reduced the SPdomain construct still migrates as 28 k. Da while both the CCP 2 SP and the CCP 1 CCP 2 SP migrates as two major bands, the serine protease domain at 28 k. Da, the CCP 2 -domain at 9 k. Da and the CCP 1 CCP 2 -domain at 20 k. Da. A minor contaminant is observed in the CCP 2 SP construct lane both reduced and non-reduced. Two μg of each of the constructs was loaded per track except for the SP domain construct for which the protein concentration appeared to diminish on storage (although no cleavage fragments are visible). For a detailed description of the experimental setup see section 6. 2. 1.
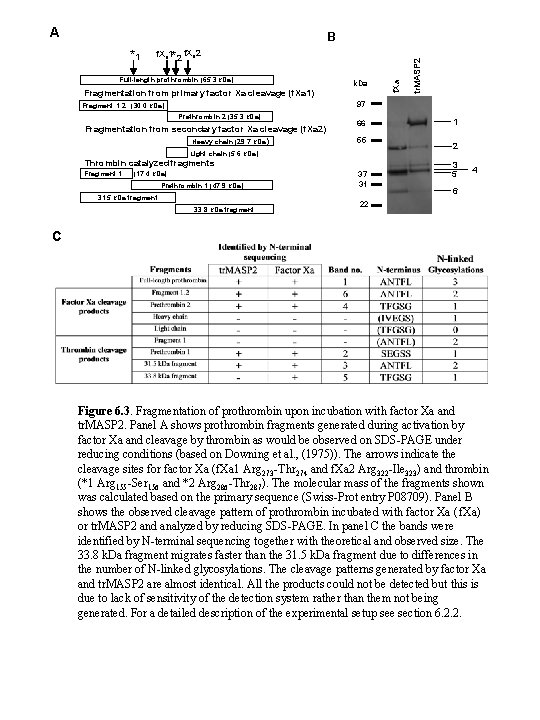
A f. Xa 1*2 f. Xa 2 Full-length prothrombin (65. 3 k. Da) Fragmentation from primary factor Xa cleavage (f. Xa 1) k. Da 97 Fragment 1. 2 (30. 0 k. Da) Prethrombin 2 (35. 3 k. Da) Fragmentation from secondary factor Xa cleavage (f. Xa 2) Heavy chain (29. 7 k. Da) 66 55 Light chain (5. 6 k. Da) Thrombin catalyzed fragments Fragment 1 f. Xa *1 tr. MASP 2 B (17. 4 k. Da) Prethrombin 1 (47. 9 k. Da) 31. 5 k. Da fragment 33. 8 k. Da fragment 37 31 1 2 3 5 4 6 22 C Figure 6. 3. Fragmentation of prothrombin upon incubation with factor Xa and tr. MASP 2. Panel A shows prothrombin fragments generated during activation by factor Xa and cleavage by thrombin as would be observed on SDS-PAGE under reducing conditions (based on Downing et al. , (1975)). The arrows indicate the cleavage sites for factor Xa (f. Xa 1 Arg 273 -Thr 274 and f. Xa 2 Arg 322 -Ile 323) and thrombin (*1 Arg 155 -Ser 156 and *2 Arg 286 -Thr 287). The molecular mass of the fragments shown was calculated based on the primary sequence (Swiss-Prot entry P 08709). Panel B shows the observed cleavage pattern of prothrombin incubated with factor Xa ( f. Xa) or tr. MASP 2 and analyzed by reducing SDS-PAGE. In panel C the bands were identified by N-terminal sequencing together with theoretical and observed size. The 33. 8 k. Da fragment migrates faster than the 31. 5 k. Da fragment due to differences in the number of N-linked glycosylations. The cleavage patterns generated by factor Xa and tr. MASP 2 are almost identical. All the products could not be detected but this is due to lack of sensitivity of the detection system rather than them not being generated. For a detailed description of the experimental setup see section 6. 2. 2.
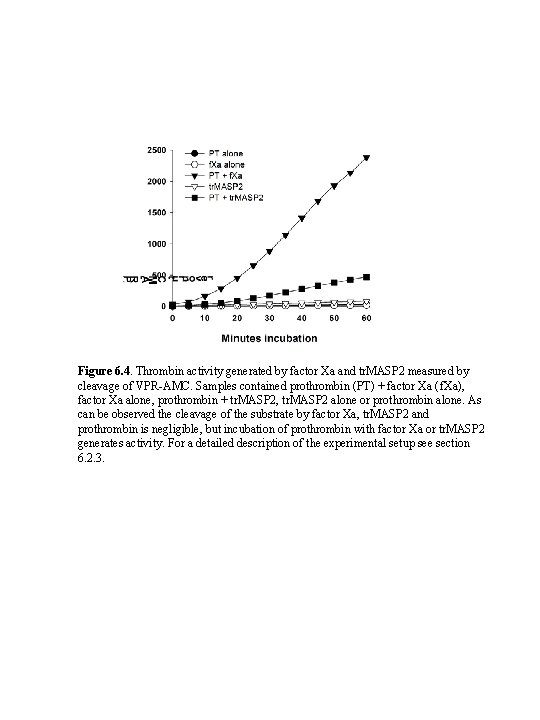
Figure 6. 4. Thrombin activity generated by factor Xa and tr. MASP 2 measured by cleavage of VPR-AMC. Samples contained prothrombin (PT) + factor Xa (f. Xa), factor Xa alone, prothrombin + tr. MASP 2, tr. MASP 2 alone or prothrombin alone. As can be observed the cleavage of the substrate by factor Xa, tr. MASP 2 and prothrombin is negligible, but incubation of prothrombin with factor Xa or tr. MASP 2 generates activity. For a detailed description of the experimental setup see section 6. 2. 3.
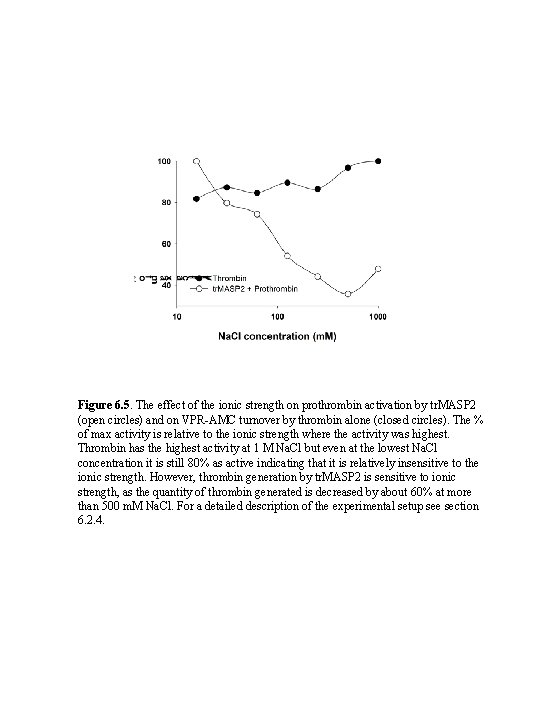
Figure 6. 5. The effect of the ionic strength on prothrombin activation by tr. MASP 2 (open circles) and on VPR-AMC turnover by thrombin alone (closed circles). The % of max activity is relative to the ionic strength where the activity was highest. Thrombin has the highest activity at 1 M Na. Cl but even at the lowest Na. Cl concentration it is still 80% as active indicating that it is relatively insensitive to the ionic strength. However, thrombin generation by tr. MASP 2 is sensitive to ionic strength, as the quantity of thrombin generated is decreased by about 60% at more than 500 m. M Na. Cl. For a detailed description of the experimental setup see section 6. 2. 4.
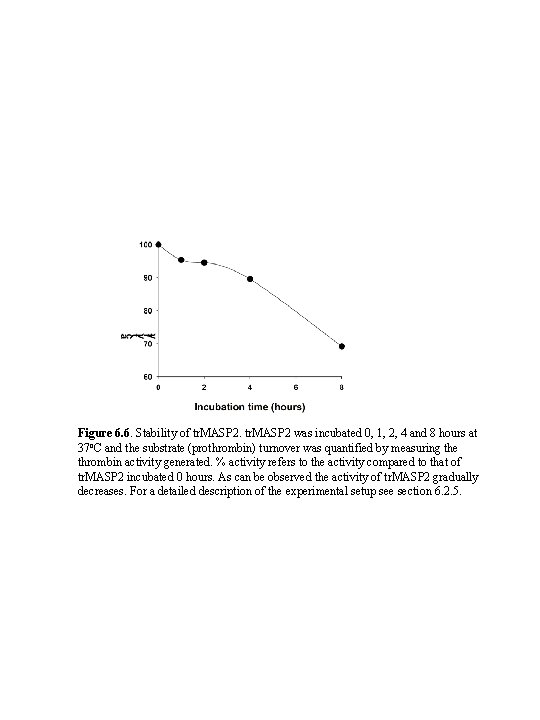
Figure 6. 6. Stability of tr. MASP 2 was incubated 0, 1, 2, 4 and 8 hours at 37 o. C and the substrate (prothrombin) turnover was quantified by measuring the thrombin activity generated. % activity refers to the activity compared to that of tr. MASP 2 incubated 0 hours. As can be observed the activity of tr. MASP 2 gradually decreases. For a detailed description of the experimental setup see section 6. 2. 5.
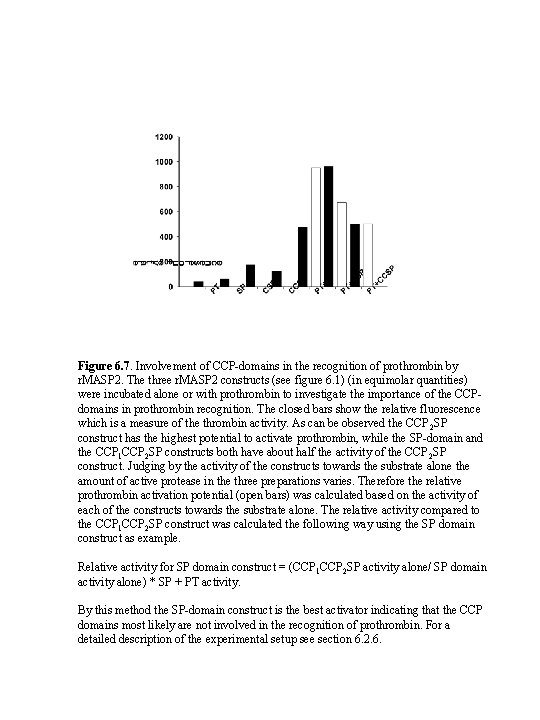
Figure 6. 7. Involvement of CCP-domains in the recognition of prothrombin by r. MASP 2. The three r. MASP 2 constructs (see figure 6. 1) (in equimolar quantities) were incubated alone or with prothrombin to investigate the importance of the CCPdomains in prothrombin recognition. The closed bars show the relative fluorescence which is a measure of the thrombin activity. As can be observed the CCP 2 SP construct has the highest potential to activate prothrombin, while the SP-domain and the CCP 1 CCP 2 SP constructs both have about half the activity of the CCP 2 SP construct. Judging by the activity of the constructs towards the substrate alone the amount of active protease in the three preparations varies. Therefore the relative prothrombin activation potential (open bars) was calculated based on the activity of each of the constructs towards the substrate alone. The relative activity compared to the CCP 1 CCP 2 SP construct was calculated the following way using the SP domain construct as example. Relative activity for SP domain construct = (CCP 1 CCP 2 SP activity alone/ SP domain activity alone) * SP + PT activity. By this method the SP-domain construct is the best activator indicating that the CCP domains most likely are not involved in the recognition of prothrombin. For a detailed description of the experimental setup see section 6. 2. 6.
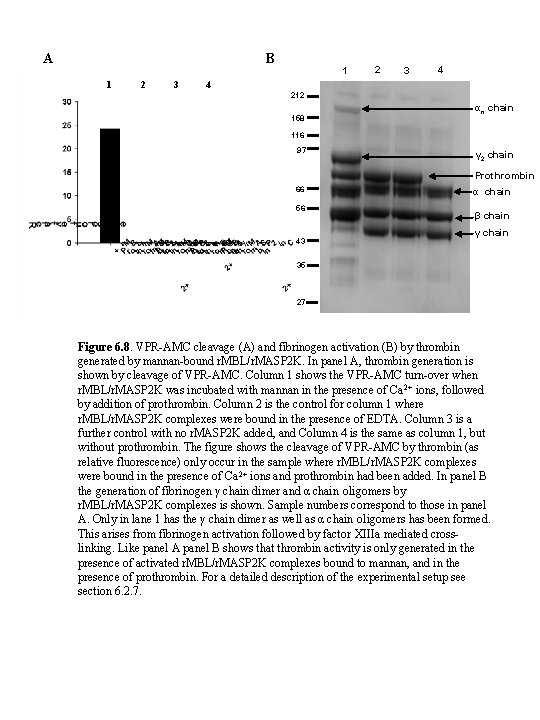
A B 1 2 3 4 4 212 158 αn chain 116 97 γ 2 chain Prothrombin 66 56 43 α chain β chain γ chain 35 27 Figure 6. 8. VPR-AMC cleavage (A) and fibrinogen activation (B) by thrombin generated by mannan-bound r. MBL/r. MASP 2 K. In panel A, thrombin generation is shown by cleavage of VPR-AMC. Column 1 shows the VPR-AMC turn-over when r. MBL/r. MASP 2 K was incubated with mannan in the presence of Ca 2+ ions, followed by addition of prothrombin. Column 2 is the control for column 1 where r. MBL/r. MASP 2 K complexes were bound in the presence of EDTA. Column 3 is a further control with no r. MASP 2 K added, and Column 4 is the same as column 1, but without prothrombin. The figure shows the cleavage of VPR-AMC by thrombin (as relative fluorescence) only occur in the sample where r. MBL/r. MASP 2 K complexes were bound in the presence of Ca 2+ ions and prothrombin had been added. In panel B the generation of fibrinogen γ chain dimer and α chain oligomers by r. MBL/r. MASP 2 K complexes is shown. Sample numbers correspond to those in panel A. Only in lane 1 has the γ chain dimer as well as α chain oligomers has been formed. This arises from fibrinogen activation followed by factor XIIIa mediated crosslinking. Like panel A panel B shows that thrombin activity is only generated in the presence of activated r. MBL/r. MASP 2 K complexes bound to mannan, and in the presence of prothrombin. For a detailed description of the experimental setup see section 6. 2. 7.
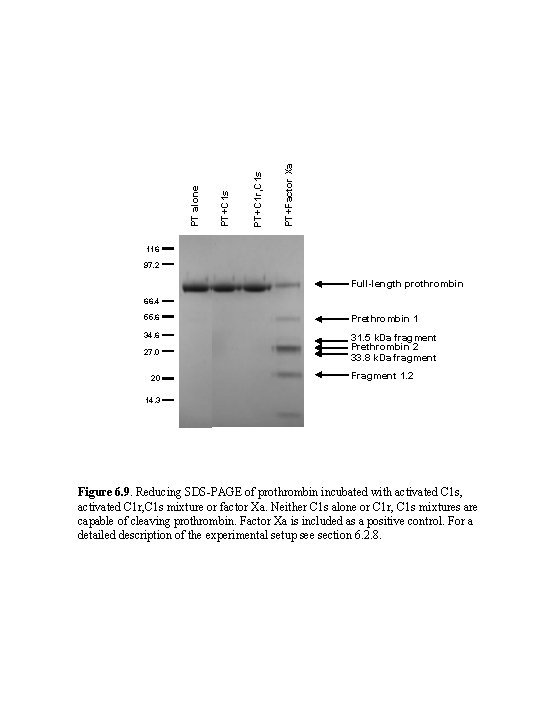
PT+Factor Xa PT+C 1 r, C 1 s PT+C 1 s PT alone 116 97. 2 Full-length prothrombin 66. 4 55. 6 Prethrombin 1 34. 6 31. 5 k. Da fragment Prethrombin 2 33. 8 k. Da fragment 27. 0 20 Fragment 1. 2 14. 3 Figure 6. 9. Reducing SDS-PAGE of prothrombin incubated with activated C 1 s, activated C 1 r, C 1 s mixture or factor Xa. Neither C 1 s alone or C 1 r, C 1 s mixtures are capable of cleaving prothrombin. Factor Xa is included as a positive control. For a detailed description of the experimental setup see section 6. 2. 8.
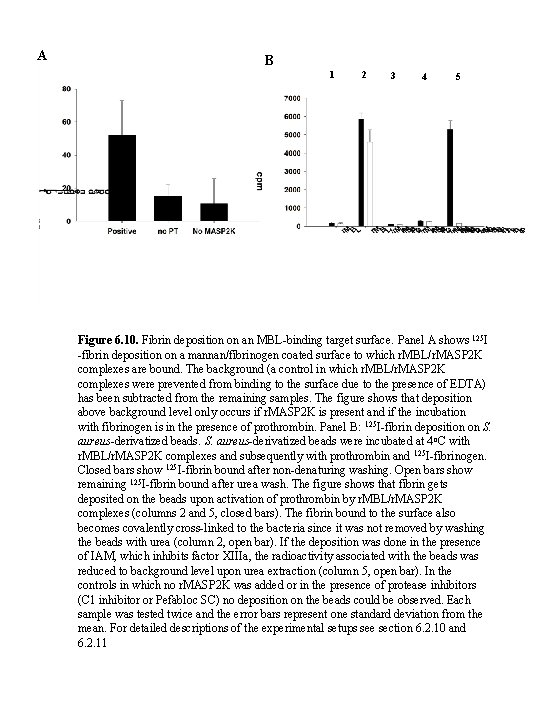
A B 1 2 3 4 5 Figure 6. 10. Fibrin deposition on an MBL-binding target surface. Panel A shows 125 I -fibrin deposition on a mannan/fibrinogen coated surface to which r. MBL/r. MASP 2 K complexes are bound. The background (a control in which r. MBL/r. MASP 2 K complexes were prevented from binding to the surface due to the presence of EDTA) has been subtracted from the remaining samples. The figure shows that deposition above background level only occurs if r. MASP 2 K is present and if the incubation with fibrinogen is in the presence of prothrombin. Panel B: 125 I-fibrin deposition on S. aureus-derivatized beads were incubated at 4 o. C with r. MBL/r. MASP 2 K complexes and subsequently with prothrombin and 125 I-fibrinogen. Closed bars show 125 I-fibrin bound after non-denaturing washing. Open bars show remaining 125 I-fibrin bound after urea wash. The figure shows that fibrin gets deposited on the beads upon activation of prothrombin by r. MBL/r. MASP 2 K complexes (columns 2 and 5, closed bars). The fibrin bound to the surface also becomes covalently cross-linked to the bacteria since it was not removed by washing the beads with urea (column 2, open bar). If the deposition was done in the presence of IAM, which inhibits factor XIIIa, the radioactivity associated with the beads was reduced to background level upon urea extraction (column 5, open bar). In the controls in which no r. MASP 2 K was added or in the presence of protease inhibitors (C 1 inhibitor or Pefabloc SC) no deposition on the beads could be observed. Each sample was tested twice and the error bars represent one standard deviation from the mean. For detailed descriptions of the experimental setups see section 6. 2. 10 and 6. 2. 11
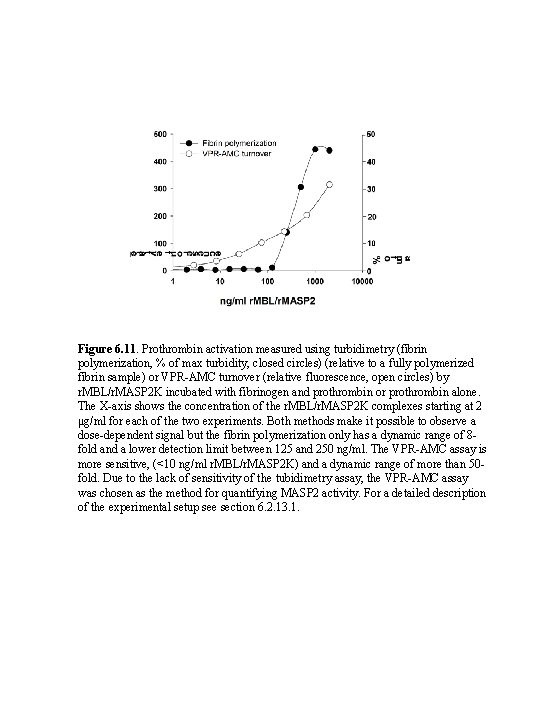
Figure 6. 11. Prothrombin activation measured using turbidimetry (fibrin polymerization, % of max turbidity, closed circles) (relative to a fully polymerized fibrin sample) or VPR-AMC turnover (relative fluorescence, open circles) by r. MBL/r. MASP 2 K incubated with fibrinogen and prothrombin or prothrombin alone. The X-axis shows the concentration of the r. MBL/r. MASP 2 K complexes starting at 2 μg/ml for each of the two experiments. Both methods make it possible to observe a dose-dependent signal but the fibrin polymerization only has a dynamic range of 8 fold and a lower detection limit between 125 and 250 ng/ml. The VPR-AMC assay is more sensitive, (<10 ng/ml r. MBL/r. MASP 2 K) and a dynamic range of more than 50 fold. Due to the lack of sensitivity of the tubidimetry assay, the VPR-AMC assay was chosen as the method for quantifying MASP 2 activity. For a detailed description of the experimental setup see section 6. 2. 13. 1.
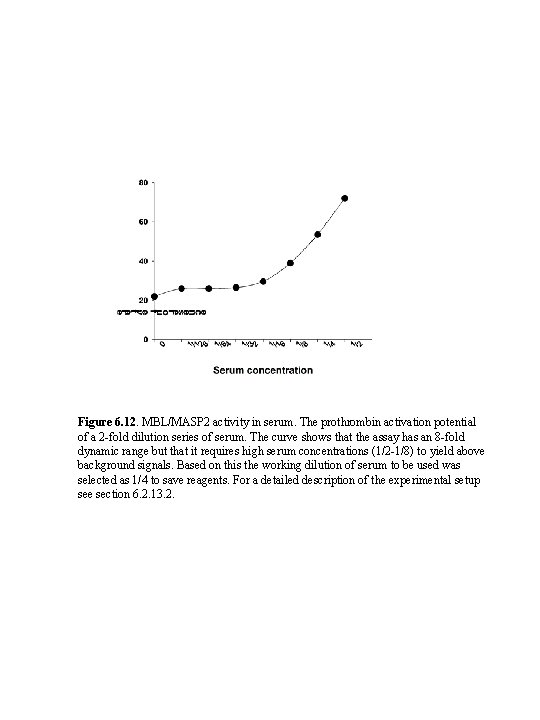
Figure 6. 12. MBL/MASP 2 activity in serum. The prothrombin activation potential of a 2 -fold dilution series of serum. The curve shows that the assay has an 8 -fold dynamic range but that it requires high serum concentrations (1/2 -1/8) to yield above background signals. Based on this the working dilution of serum to be used was selected as 1/4 to save reagents. For a detailed description of the experimental setup see section 6. 2. 13. 2.
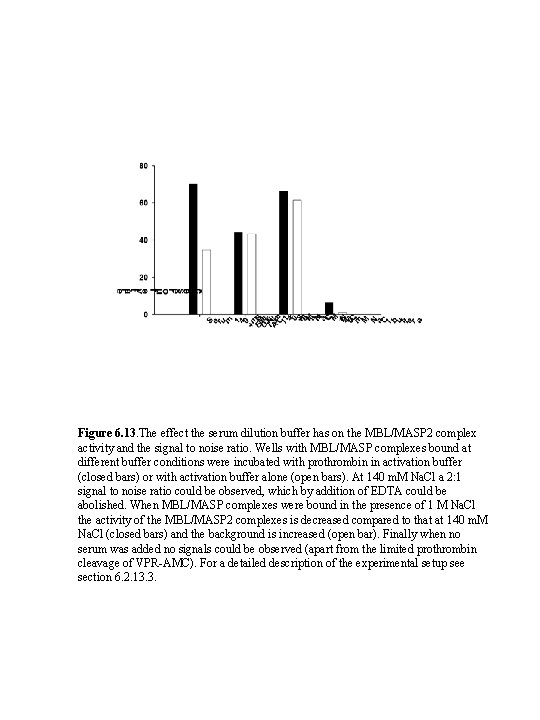
Figure 6. 13. The effect the serum dilution buffer has on the MBL/MASP 2 complex activity and the signal to noise ratio. Wells with MBL/MASP complexes bound at different buffer conditions were incubated with prothrombin in activation buffer (closed bars) or with activation buffer alone (open bars). At 140 m. M Na. Cl a 2: 1 signal to noise ratio could be observed, which by addition of EDTA could be abolished. When MBL/MASP complexes were bound in the presence of 1 M Na. Cl the activity of the MBL/MASP 2 complexes is decreased compared to that at 140 m. M Na. Cl (closed bars) and the background is increased (open bar). Finally when no serum was added no signals could be observed (apart from the limited prothrombin cleavage of VPR-AMC). For a detailed description of the experimental setup see section 6. 2. 13. 3.
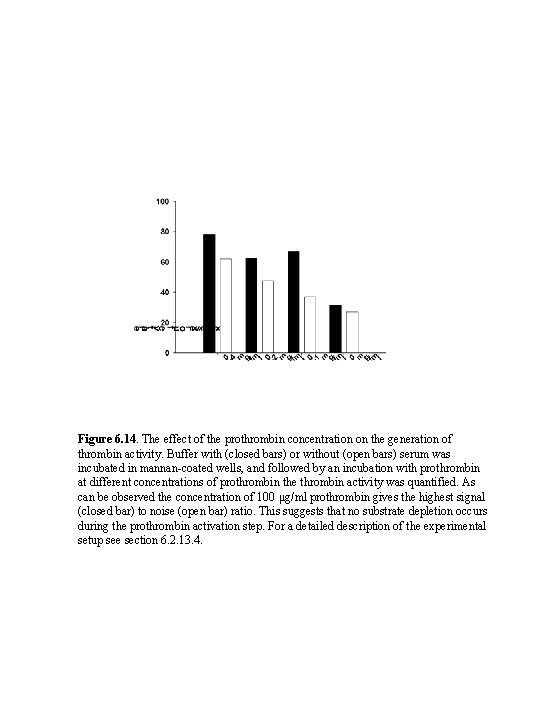
Figure 6. 14. The effect of the prothrombin concentration on the generation of thrombin activity. Buffer with (closed bars) or without (open bars) serum was incubated in mannan-coated wells, and followed by an incubation with prothrombin at different concentrations of prothrombin the thrombin activity was quantified. As can be observed the concentration of 100 μg/ml prothrombin gives the highest signal (closed bar) to noise (open bar) ratio. This suggests that no substrate depletion occurs during the prothrombin activation step. For a detailed description of the experimental setup see section 6. 2. 13. 4.
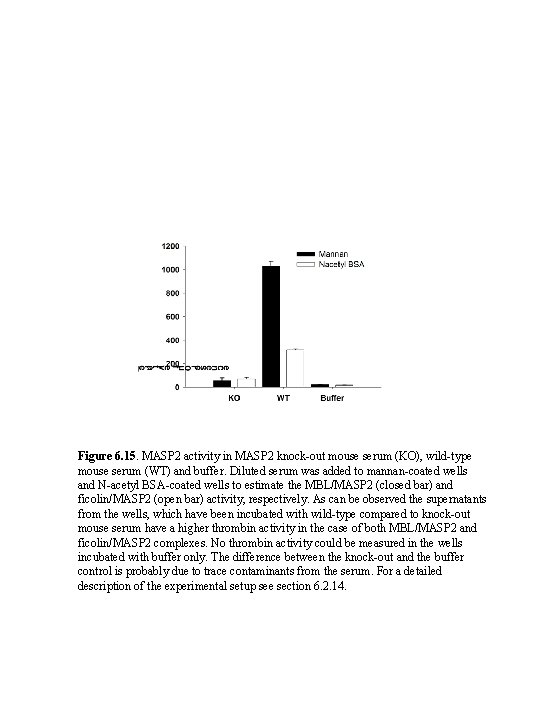
Figure 6. 15. MASP 2 activity in MASP 2 knock-out mouse serum (KO), wild-type mouse serum (WT) and buffer. Diluted serum was added to mannan-coated wells and N-acetyl BSA-coated wells to estimate the MBL/MASP 2 (closed bar) and ficolin/MASP 2 (open bar) activity, respectively. As can be observed the supernatants from the wells, which have been incubated with wild-type compared to knock-out mouse serum have a higher thrombin activity in the case of both MBL/MASP 2 and ficolin/MASP 2 complexes. No thrombin activity could be measured in the wells incubated with buffer only. The difference between the knock-out and the buffer control is probably due to trace contaminants from the serum. For a detailed description of the experimental setup see section 6. 2. 14.
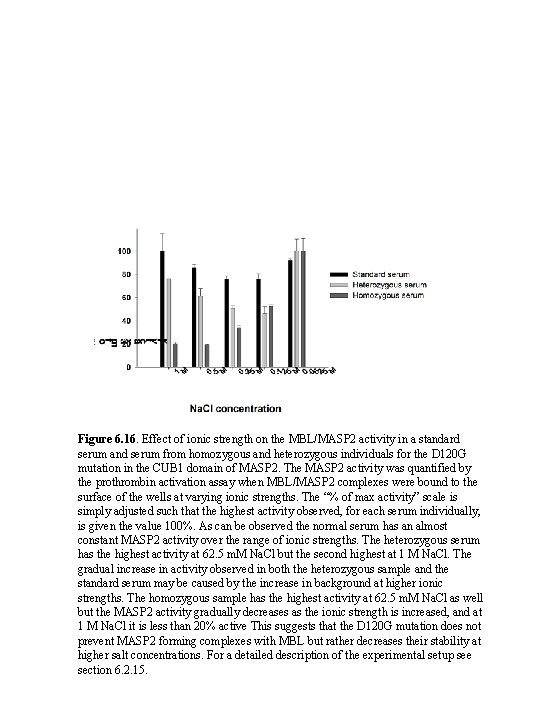
Figure 6. 16. Effect of ionic strength on the MBL/MASP 2 activity in a standard serum and serum from homozygous and heterozygous individuals for the D 120 G mutation in the CUB 1 domain of MASP 2. The MASP 2 activity was quantified by the prothrombin activation assay when MBL/MASP 2 complexes were bound to the surface of the wells at varying ionic strengths. The “% of max activity” scale is simply adjusted such that the highest activity observed, for each serum individually, is given the value 100%. As can be observed the normal serum has an almost constant MASP 2 activity over the range of ionic strengths. The heterozygous serum has the highest activity at 62. 5 m. M Na. Cl but the second highest at 1 M Na. Cl. The gradual increase in activity observed in both the heterozygous sample and the standard serum may be caused by the increase in background at higher ionic strengths. The homozygous sample has the highest activity at 62. 5 m. M Na. Cl as well but the MASP 2 activity gradually decreases as the ionic strength is increased, and at 1 M Na. Cl it is less than 20% active This suggests that the D 120 G mutation does not prevent MASP 2 forming complexes with MBL but rather decreases their stability at higher salt concentrations. For a detailed description of the experimental setup see section 6. 2. 15.
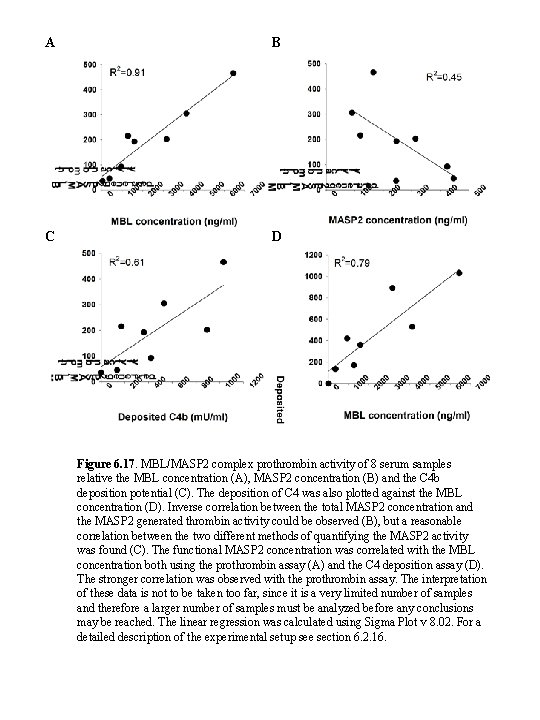
A B C D Figure 6. 17. MBL/MASP 2 complex prothrombin activity of 8 serum samples relative the MBL concentration (A), MASP 2 concentration (B) and the C 4 b deposition potential (C). The deposition of C 4 was also plotted against the MBL concentration (D). Inverse correlation between the total MASP 2 concentration and the MASP 2 generated thrombin activity could be observed (B), but a reasonable correlation between the two different methods of quantifying the MASP 2 activity was found (C). The functional MASP 2 concentration was correlated with the MBL concentration both using the prothrombin assay (A) and the C 4 deposition assay (D). The stronger correlation was observed with the prothrombin assay. The interpretation of these data is not to be taken too far, since it is a very limited number of samples and therefore a larger number of samples must be analyzed before any conclusions may be reached. The linear regression was calculated using Sigma Plot v 8. 02. For a detailed description of the experimental setup see section 6. 2. 16.
- Slides: 18