Synthetic minimal cells toolbox for biosensing and monitoring

Synthetic minimal cells -toolbox for biosensing and monitoring biodiversity Kate Adamala MIT Media Lab, MIT Department of Biological Engineering

Kate Adamala Readout of biology – where? End-point Remove samples, analyze in the lab Continuous Analytes are continuously monitored in the natural context. End-point Continuous

Readout of biology in situ – how? Electronic sensors + Fast, reliable, remote readout + More sensitive and versatile, at the current state of technology - Electronic waste in the environment (if portable assay is available) - Needs a biolab (if no portable assay) Biosensors + Fast, in situ readout +/- Protein sensors, genetically encoded parts - More difficult to develop for each analyte - More sensitive to environmental stressors Kate Adamala
Biosensors Modified bacteria + very cheap to propagate, stable under wide array of conditions - They breed! Liposomal biosensors (synthetic minimal cells) + completely bio-orthogonal - More difficult to prepare, less stable Kate Adamala
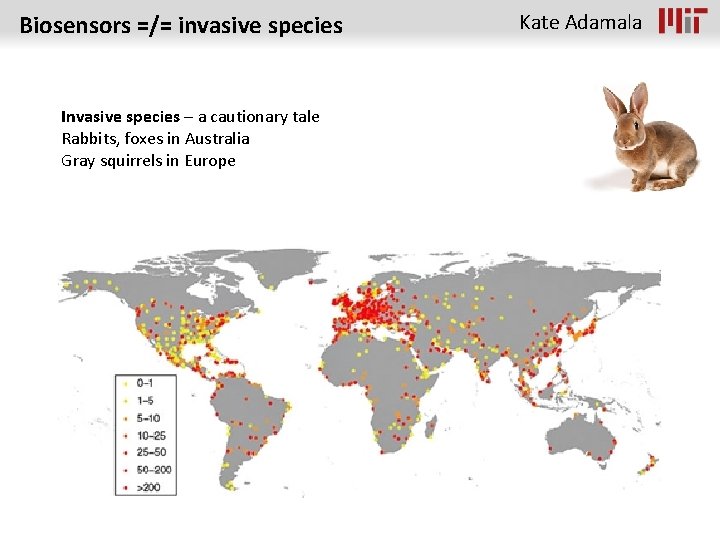
Biosensors =/= invasive species Invasive species – a cautionary tale Rabbits, foxes in Australia Gray squirrels in Europe Kate Adamala
Biosensors =/= invasive species Not in the Amazon! (at least not more from us…) Kate Adamala
Biosensors Modified bacteria + very cheap to propagate, stable under wide array of conditions - They breed! Liposomal biosensors (synthetic minimal cells) + completely bio-orthogonal - More difficult to prepare, less stable Kate Adamala
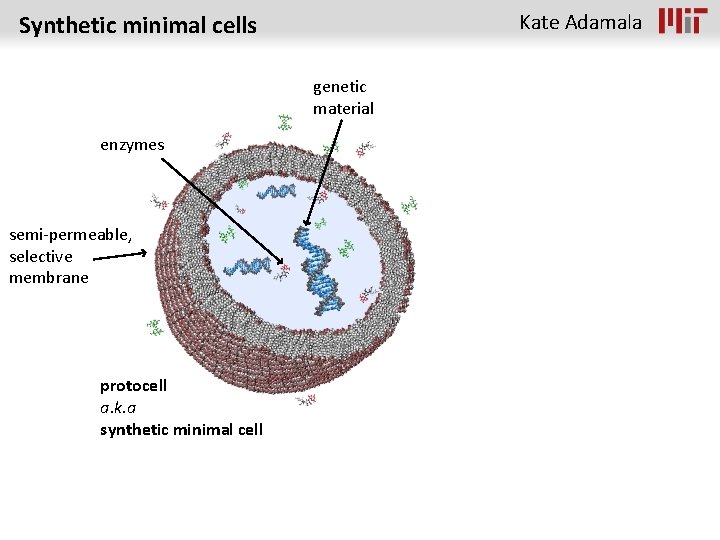
Kate Adamala Synthetic minimal cells genetic material enzymes semi-permeable, selective membrane protocell a. k. a synthetic minimal cell
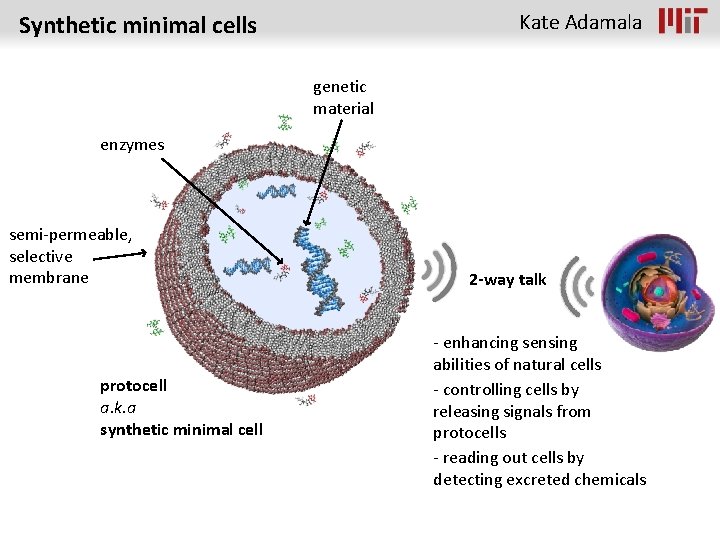
Kate Adamala Synthetic minimal cells genetic material enzymes semi-permeable, selective membrane protocell a. k. a synthetic minimal cell 2 -way talk - enhancing sensing abilities of natural cells - controlling cells by releasing signals from protocells - reading out cells by detecting excreted chemicals
Kate Adamala Synthetic minimal cells cell
Kate Adamala Synthetic minimal cells cell
Kate Adamala Synthetic minimal cells cell
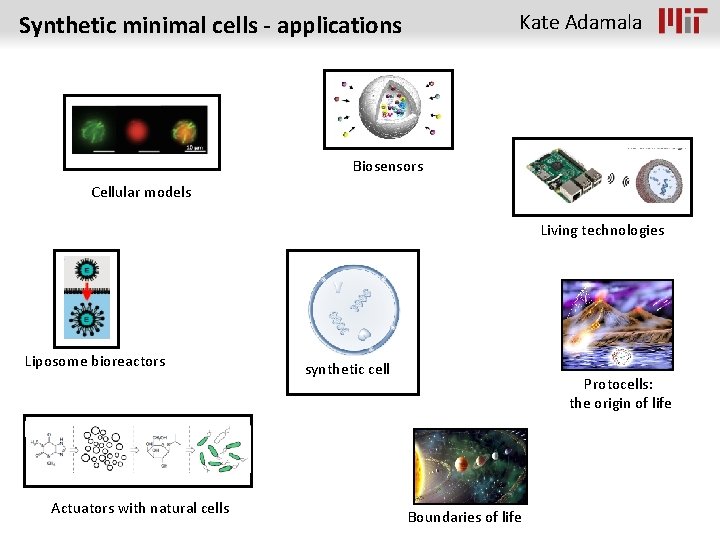
Kate Adamala Synthetic minimal cells - applications Biosensors Cellular models Living technologies Liposome bioreactors Actuators with natural cells synthetic cell Protocells: the origin of life Boundaries of life
The anatomy of the synthetic cell The compartment Kate Adamala The encapsulated machinery
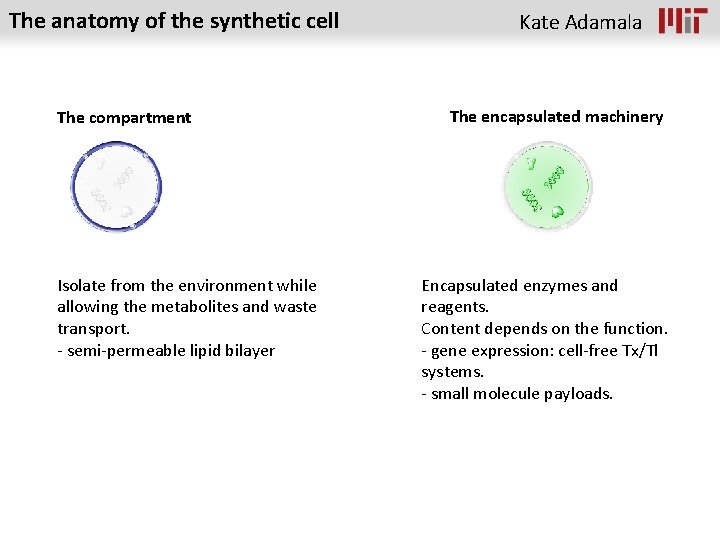
The anatomy of the synthetic cell The compartment Isolate from the environment while allowing the metabolites and waste transport. - semi-permeable lipid bilayer Kate Adamala The encapsulated machinery Encapsulated enzymes and reagents. Content depends on the function. - gene expression: cell-free Tx/Tl systems. - small molecule payloads.
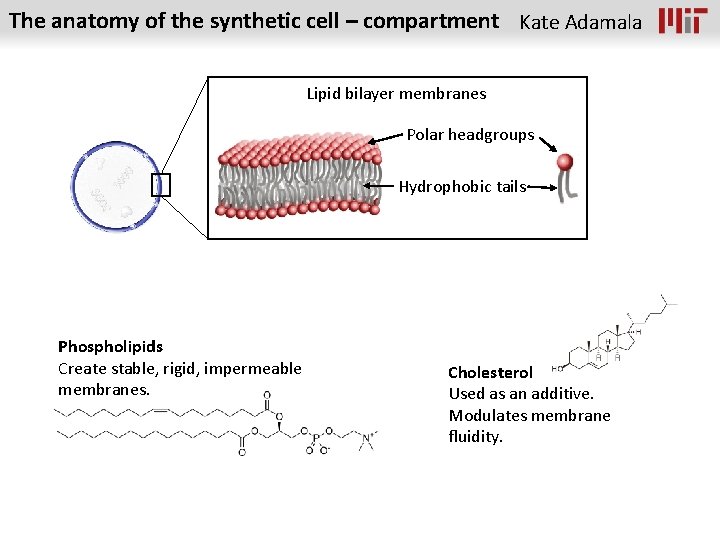
The anatomy of the synthetic cell – compartment Kate Adamala Lipid bilayer membranes Polar headgroups Hydrophobic tails Phospholipids Create stable, rigid, impermeable membranes. Cholesterol Used as an additive. Modulates membrane fluidity.
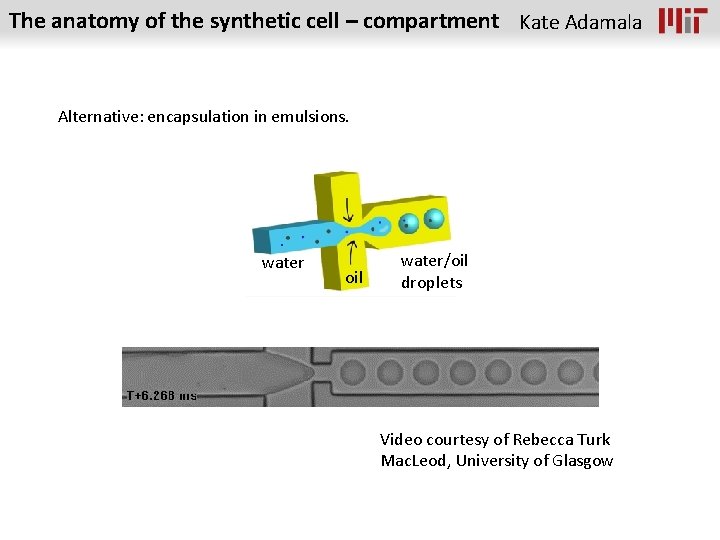
The anatomy of the synthetic cell – compartment Kate Adamala Alternative: encapsulation in emulsions. water oil water/oil droplets Video courtesy of Rebecca Turk Mac. Leod, University of Glasgow
The anatomy of the synthetic cell The compartment Isolate from the environment while allowing the metabolites and waste transport. - semi-permeable lipid bilayer Kate Adamala The encapsulated machinery Encapsulated enzymes and reagents. Content depends on the function. - gene expression: cell-free Tx/Tl systems. - small molecule payloads.

Cell-free Tx/Tl Cell free transcription and translation (Tx/Tl) - No transfection, cell culture, protein purification Kate Adamala
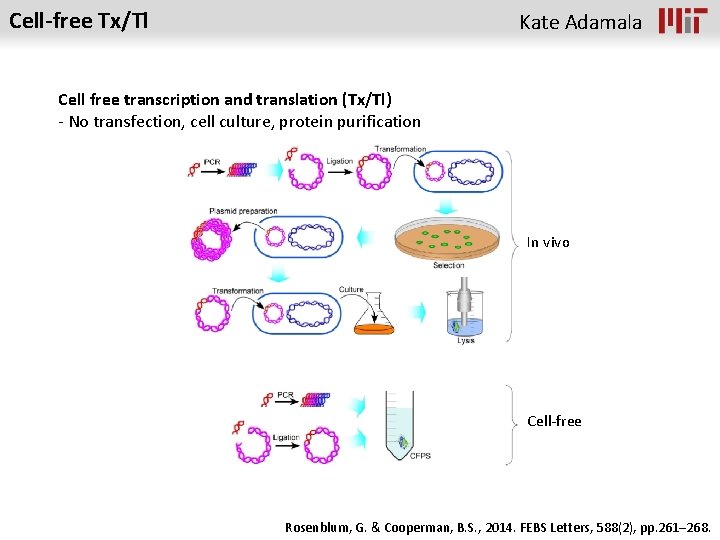
Cell-free Tx/Tl Kate Adamala Cell free transcription and translation (Tx/Tl) - No transfection, cell culture, protein purification In vivo Cell-free Rosenblum, G. & Cooperman, B. S. , 2014. FEBS Letters, 588(2), pp. 261– 268.

Cell-free Tx/Tl Kate Adamala Cell free transcription and translation (Tx/Tl) - No transfection, cell culture, protein purification - Easier to add control factors, chaperones, inhibitors, binding and folding partners - Easier to remove proteases, nucleases - Rapid, small scale production of isotope labeled proteins (for NMR and kinetics studies) and selenomethionine labeled proteins (for X-ray crystallography) - Possible to make toxic proteins - Avoid inclusion bodies
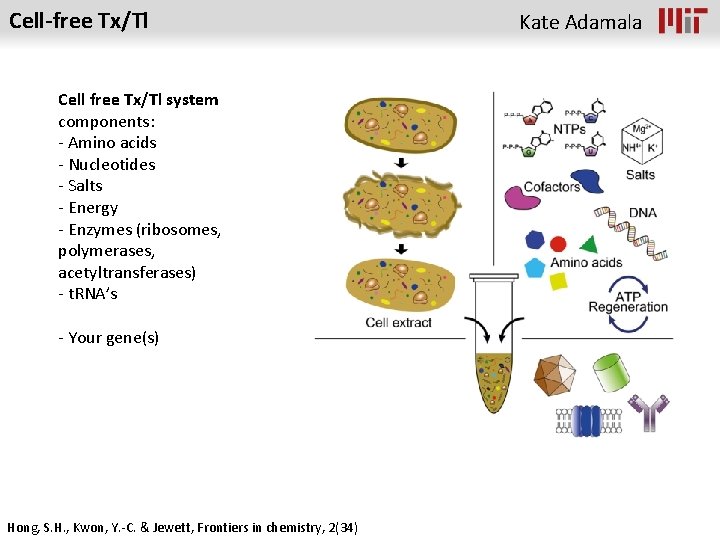
Cell-free Tx/Tl Cell free Tx/Tl system components: - Amino acids - Nucleotides - Salts - Energy - Enzymes (ribosomes, polymerases, acetyltransferases) - t. RNA’s - Your gene(s) Hong, S. H. , Kwon, Y. -C. & Jewett, Frontiers in chemistry, 2(34) Kate Adamala
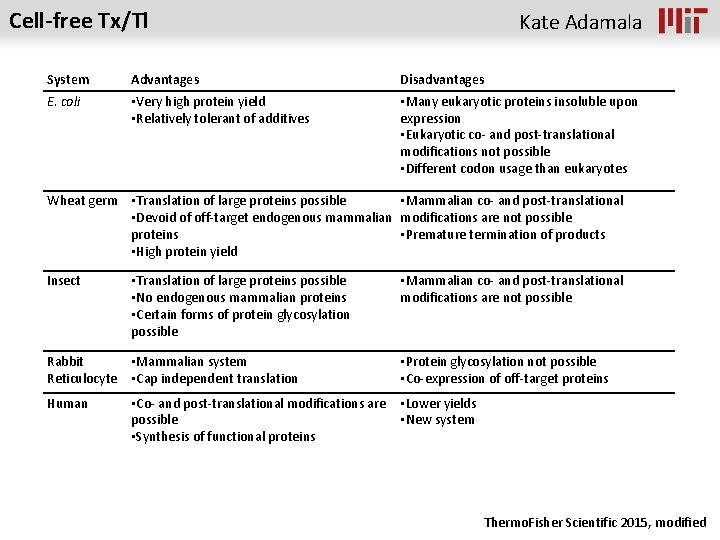
Cell-free Tx/Tl Kate Adamala System Advantages Disadvantages E. coli • Very high protein yield • Relatively tolerant of additives • Many eukaryotic proteins insoluble upon expression • Eukaryotic co- and post-translational modifications not possible • Different codon usage than eukaryotes Wheat germ • Translation of large proteins possible • Mammalian co- and post-translational • Devoid of off-target endogenous mammalian modifications are not possible proteins • Premature termination of products • High protein yield Insect • Translation of large proteins possible • No endogenous mammalian proteins • Certain forms of protein glycosylation possible • Mammalian co- and post-translational modifications are not possible Rabbit • Mammalian system Reticulocyte • Cap independent translation • Protein glycosylation not possible • Co-expression of off-target proteins Human • Lower yields • New system • Co- and post-translational modifications are possible • Synthesis of functional proteins Thermo. Fisher Scientific 2015, modified
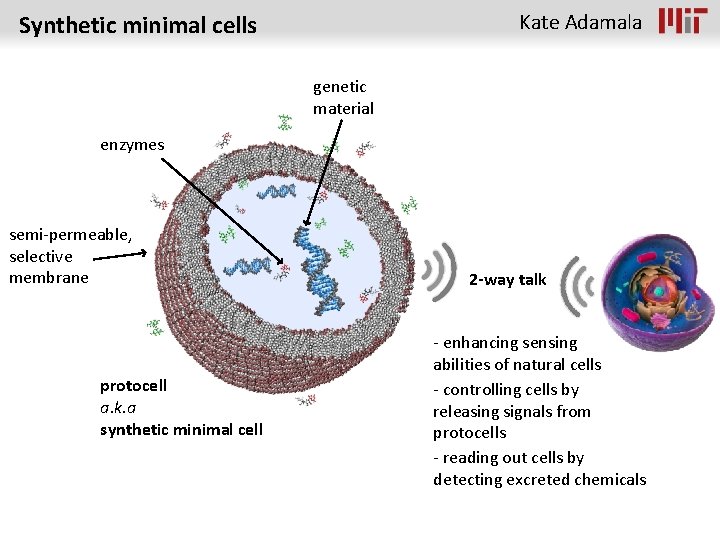
Kate Adamala Synthetic minimal cells genetic material enzymes semi-permeable, selective membrane protocell a. k. a synthetic minimal cell 2 -way talk - enhancing sensing abilities of natural cells - controlling cells by releasing signals from protocells - reading out cells by detecting excreted chemicals
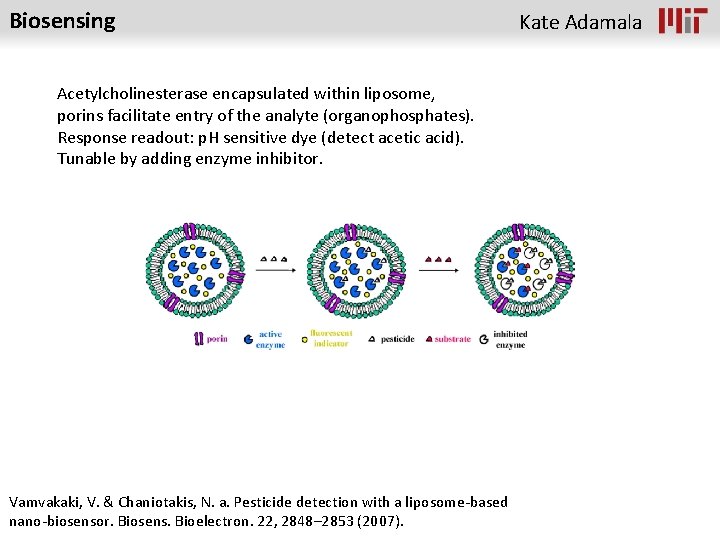
Biosensing Acetylcholinesterase encapsulated within liposome, porins facilitate entry of the analyte (organophosphates). Response readout: p. H sensitive dye (detect acetic acid). Tunable by adding enzyme inhibitor. Vamvakaki, V. & Chaniotakis, N. a. Pesticide detection with a liposome-based nano-biosensor. Biosens. Bioelectron. 22, 2848– 2853 (2007). Kate Adamala
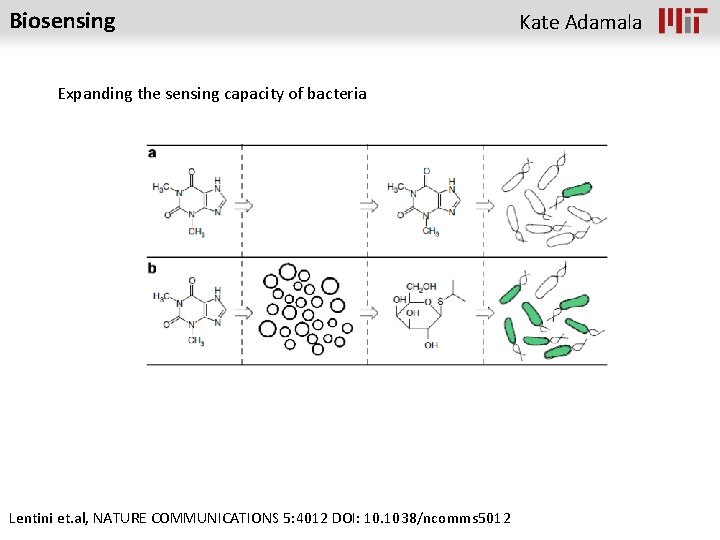
Biosensing Expanding the sensing capacity of bacteria Lentini et. al, NATURE COMMUNICATIONS 5: 4012 DOI: 10. 1038/ncomms 5012 Kate Adamala
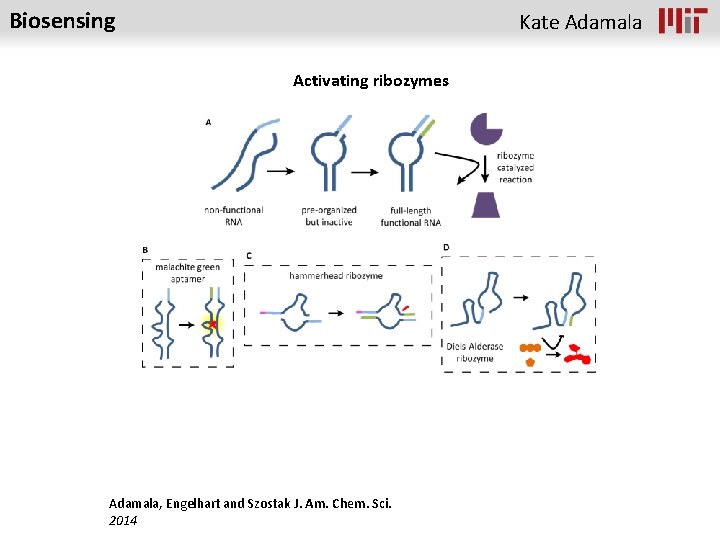
Biosensing Kate Adamala Activating ribozymes Adamala, Engelhart and Szostak J. Am. Chem. Sci. 2014
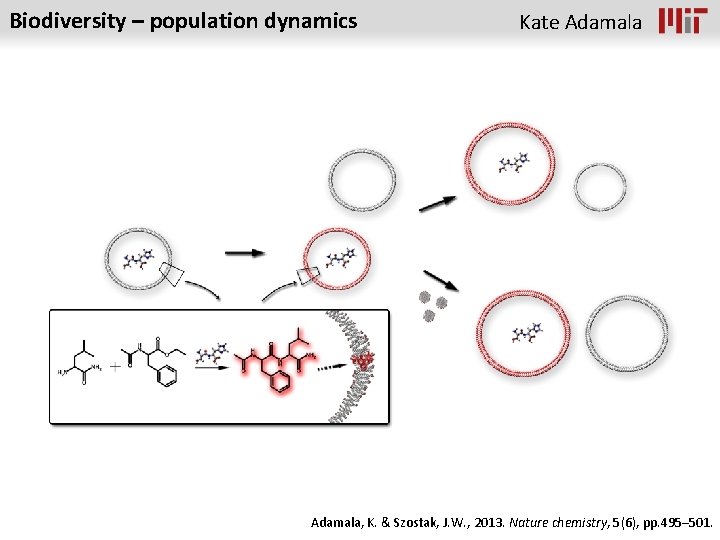
Biodiversity – population dynamics Kate Adamala, K. & Szostak, J. W. , 2013. Nature chemistry, 5(6), pp. 495– 501.
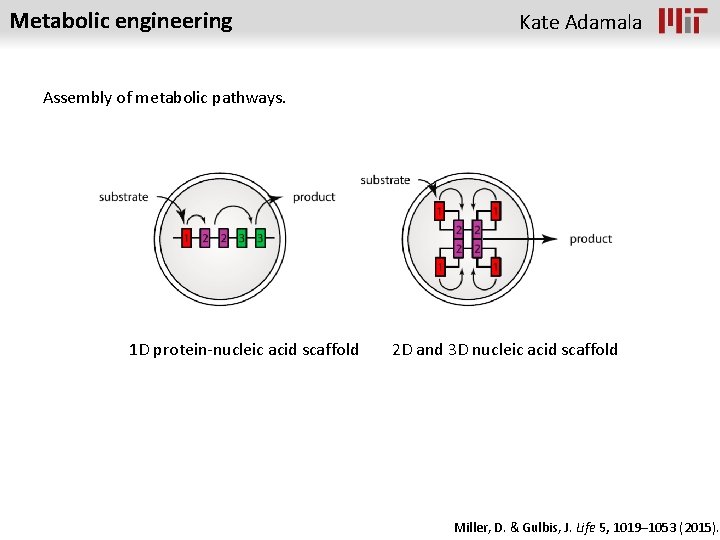
Metabolic engineering Kate Adamala Assembly of metabolic pathways. 1 D protein-nucleic acid scaffold 2 D and 3 D nucleic acid scaffold Miller, D. & Gulbis, J. Life 5, 1019– 1053 (2015).
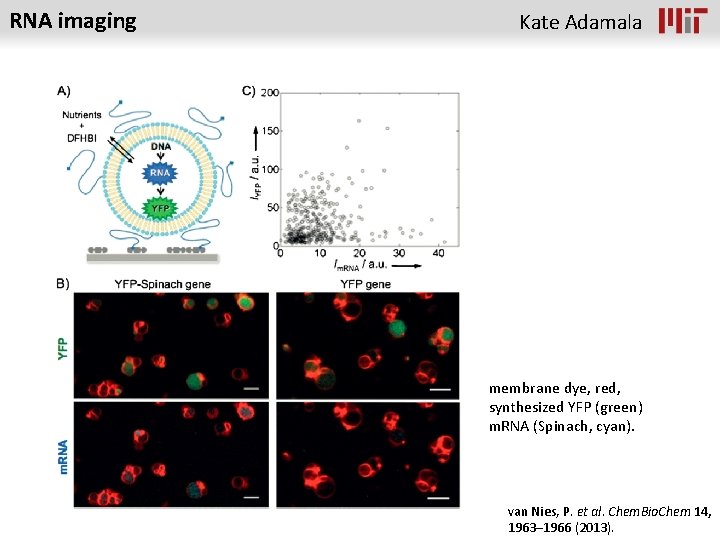
RNA imaging Kate Adamala membrane dye, red, synthesized YFP (green) m. RNA (Spinach, cyan). van Nies, P. et al. Chem. Bio. Chem 14, 1963– 1966 (2013).

Kate Adamala Thank you! Thanks to Beno Juarez @ Fab. Lab Lima! Thanks to all members of Boyden Lab, especially Ed Boyden, Daniel Martin-Alarcon, Kiryl Piatkevich and Daniel Schmidt Thanks to all members of Szostak Lab, especially Jack Szostak, Aaron Engelhart, Neha Kamat and Anders Bjorkbom Thanks to Pierluigi Luisi and Pasquale Stano, and everyone in Luisi Lab and big thanks to everyone else for comments and support! Funding from NSF CBET, Howard Hughes Medical Institute and NASA Exobiology

Kate Adamala
- Slides: 32