Synthesizing Units for modelling cell physiology Bas Kooijman
Synthesizing Units for modelling cell physiology Bas Kooijman Research program: Dynamic Energy Budget theory Dept of Theoretical Biology Vrije Universiteit, Amsterdam http: //www. bio. vu. nl/thb/deb/ adult em br yo e l i n e v ju Leiden, 2004/06/24

Weird world at small scale Almost all transformations in cells are enzyme mediated Classic enzyme kinetics: based on chemical kinetics (industrial enzymes) • diffusion/convection • larger number of molecules • constant reactor volume • law of mass action: transformation rate product of conc. of substrates Problematic application in cellular metabolism: • definition of concentration (compartments, moving organelles) • transport mechanisms (proteins with address labels, targetting, allocation) • crowding (presence of many macro-molecules that do not partake in transformation) • intrinsic stochasticity due to small numbers of molecules • liquid crystalline properties • surface area - volume relationships: membrane-cytoplasm; polymer-liquid • connectivity (many metabolites are energy substrate & building block; dilution by growth) Alternative approach: reconstruction of transformation kinetics on the basis of cellular input/output kinetics
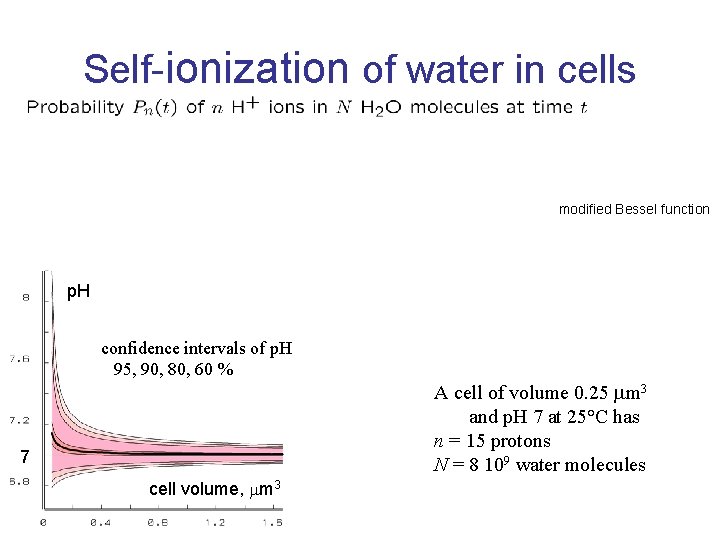
Self-ionization of water in cells modified Bessel function p. H confidence intervals of p. H 95, 90, 80, 60 % 7 cell volume, m 3 A cell of volume 0. 25 m 3 and p. H 7 at 25°C has n = 15 protons N = 8 109 water molecules

Diffusion cannot occur in cells

Crowding affects transport cytoskeletal polymers ribosomes nucleic acids proteins
ATP generation & use 5 106 ATP molecules in bacterial cell enough for 2 s of biosynthetic work Only used if energy generating & energy demanding transformations are at different site/time Processes that are not much faster than cell cycle, should be linked to large slow pools of metabolites, not to small fast pools DEB theory uses reserve as large slow pool for driving metabolism If ADP/ATP ratio varies, then rates of generation & use varies, but not necessarily the rates of transformations they drive
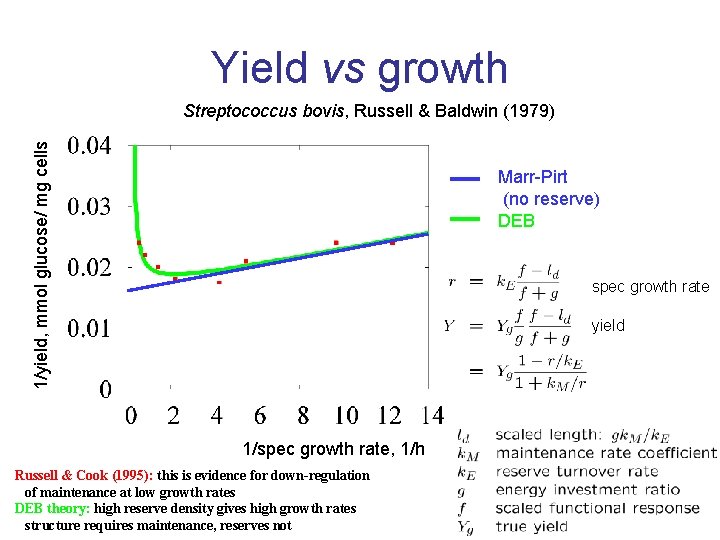
Yield vs growth 1/yield, mmol glucose/ mg cells Streptococcus bovis, Russell & Baldwin (1979) Marr-Pirt (no reserve) DEB spec growth rate yield 1/spec growth rate, 1/h Russell & Cook (1995): this is evidence for down-regulation of maintenance at low growth rates DEB theory: high reserve density gives high growth rates structure requires maintenance, reserves not

Methanotrophy Macroscopic transformation (variable yield coefficients and indices): 0 0 0 Assim (catabolic) 0 1 0 0 1 -1 0 Growth (catabolic) 0 1 -1 0 GA Growth (anabolic) 0 0 -1 1 C Carbon 1 1 0 0 0 1 1 H Hydrogen 4 0 2 0 3 O Oxygen 0 2 1 2 0 N Nitrogen 0 0 1 rate V: structure GC E: reserve Maintenance N: ammonia M O: dioxygen -1 H: water C: carbon dioxide Assim (anabolic) -2 process AA 2 symbol 1 Yield coefficients. T -1 AC Chemical indices X: methane Microscopic transformations (constant coefficients and indices): reserve density m. E = ME/MV
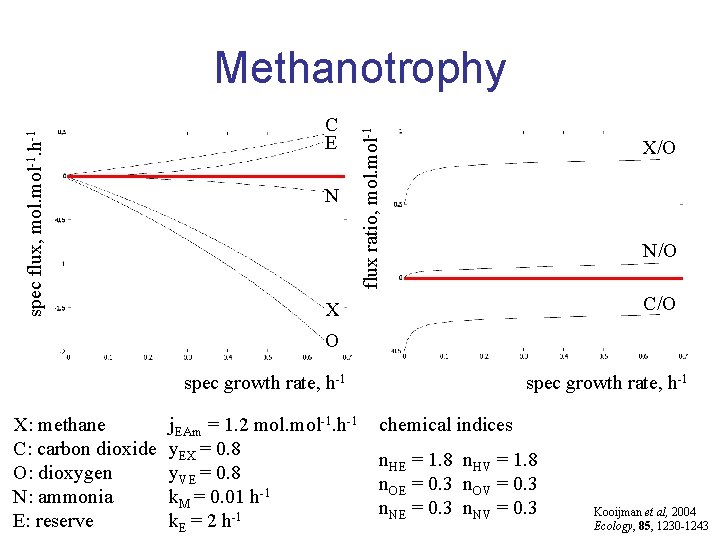
C E N flux ratio, mol-1 spec flux, mol-1. h-1 Methanotrophy X/O N/O C/O X O spec growth rate, h-1 X: methane C: carbon dioxide O: dioxygen N: ammonia E: reserve j. EAm = 1. 2 mol-1. h-1 y. EX = 0. 8 y. VE = 0. 8 k. M = 0. 01 h-1 k. E = 2 h-1 spec growth rate, h-1 chemical indices n. HE = 1. 8 n. HV = 1. 8 n. OE = 0. 3 n. OV = 0. 3 n. NE = 0. 3 n. NV = 0. 3 Kooijman et al, 2004 Ecology, 85, 1230 -1243

Enzyme kinetics A+B C : conc of compounds A, B, C : dissociation rates : association rates constant Rejection Unit Synthesizing Unit : fractions of bounded enzymes
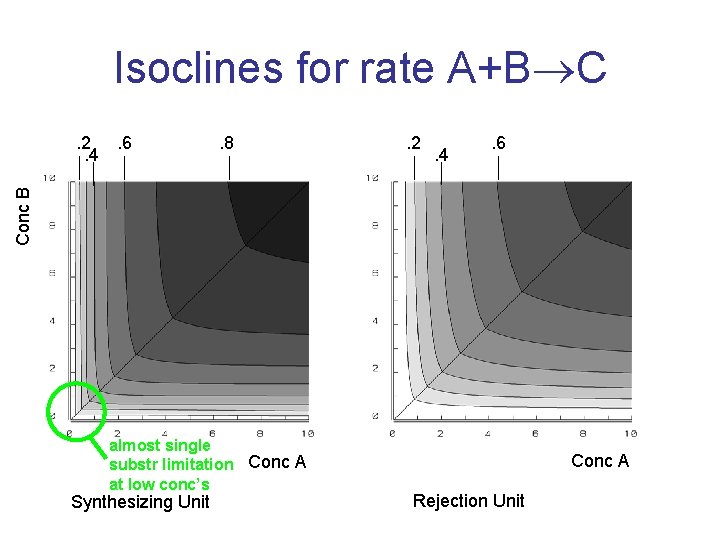
Isoclines for rate A+B C. 8 . 2 . 4 . 6 Conc B . 2. 6. 4 almost single substr limitation Conc A at low conc’s Synthesizing Unit Conc A Rejection Unit

Synthesizing units Generalized enzymes that process generalized substrates and follow classic enzyme kinetics E + S EP E + P with two modifications: • back flux is negligibly small E + S EP E + P • specification of transformation is on the basis of arrival fluxes of substrates rather than concentrations In spatially homogeneous environments: arrival fluxes concentrations
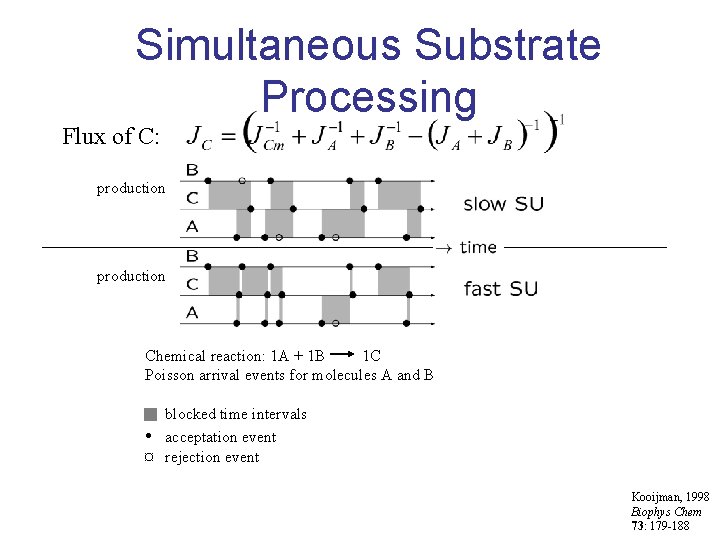
Simultaneous Substrate Processing Flux of C: production Chemical reaction: 1 A + 1 B 1 C Poisson arrival events for molecules A and B blocked time intervals • acceptation event ¤ rejection event Kooijman, 1998 Biophys Chem 73: 179 -188

SU kinetics: n 1 X 1+n 2 X 2 X product release flux of X max flux of X Expected value of tb Survivor function of tb product release 0 tb binding tc time prod. cycle Period between subsequent arrivals is exponentially distributed Sum of exponentially distributed vars is gamma distributed Production flux not very sensitive for details of stoichiometry Stoichiometry mainly affects arrival rates Kooijman, 1998 Biophys Chem 73: 179 -188
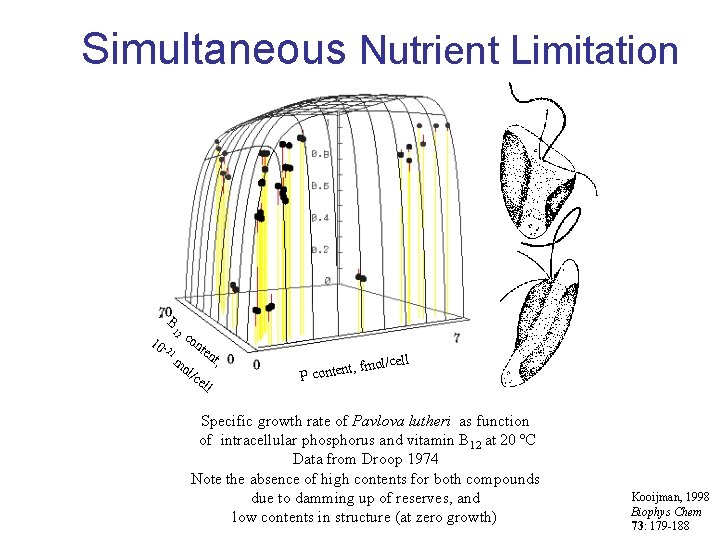
Simultaneous Nutrient Limitation B 12 10 - 21 co nte mo l/c e nt, ll l/cell t, fmo P conten Specific growth rate of Pavlova lutheri as function of intracellular phosphorus and vitamin B 12 at 20 ºC Data from Droop 1974 Note the absence of high contents for both compounds due to damming up of reserves, and low contents in structure (at zero growth) Kooijman, 1998 Biophys Chem 73: 179 -188
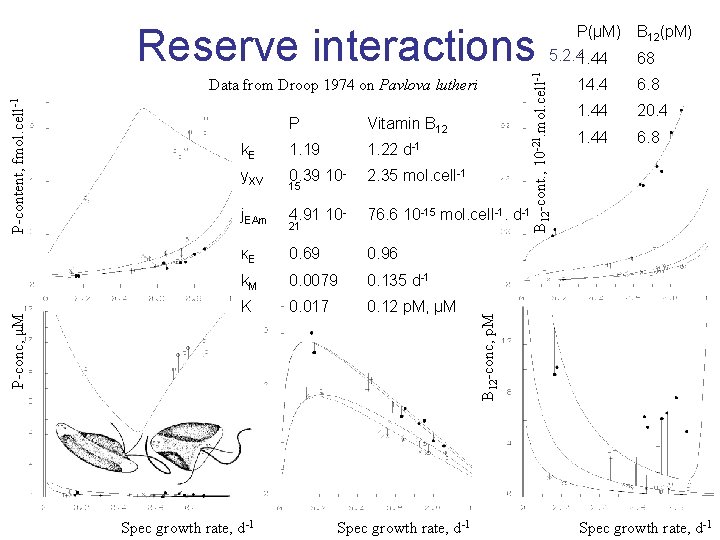
P Vitamin B 12 k. E 1. 19 1. 22 d-1 y. XV 0. 39 10 - 2. 35 mol. cell-1 j. EAm 4. 91 10 - 76. 6 10 -15 mol. cell-1. d-1 κE 0. 69 0. 96 k. M 0. 0079 0. 135 d-1 K 0. 017 0. 12 p. M, μM Spec growth rate, d-1 21 Spec growth rate, d-1 B 12(p. M) 68 14. 4 6. 8 1. 44 20. 4 1. 44 6. 8 B 12 -conc, p. M P-conc, μM P-content, fmol. cell-1 Data from Droop 1974 on Pavlova lutheri 15 5. 2. 41. 44 B 12 -cont. , 10 -21. mol. cell-1 Reserve interactions P(μM) Spec growth rate, d-1
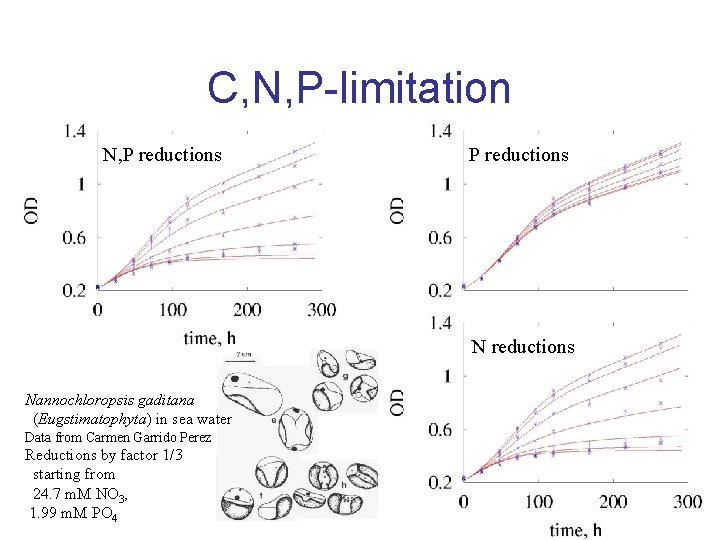
C, N, P-limitation N, P reductions Nannochloropsis gaditana (Eugstimatophyta) in sea water Data from Carmen Garrido Perez Reductions by factor 1/3 starting from 24. 7 m. M NO 3, 1. 99 m. M PO 4

C, N, P-limitation Nannochloropsis gaditana in sea water DIC nitrate phosphate res. dens. structure spec growth For uptake rate spec growth rate

Producer/consumer dynamics producer consumer : hazard rate nutr reserve of producer : total nutrient in closed system spec growth of consumer special case: consumer is not nutrient limited Kooijman et al 2004 Ecology, 85, 1230 -1243
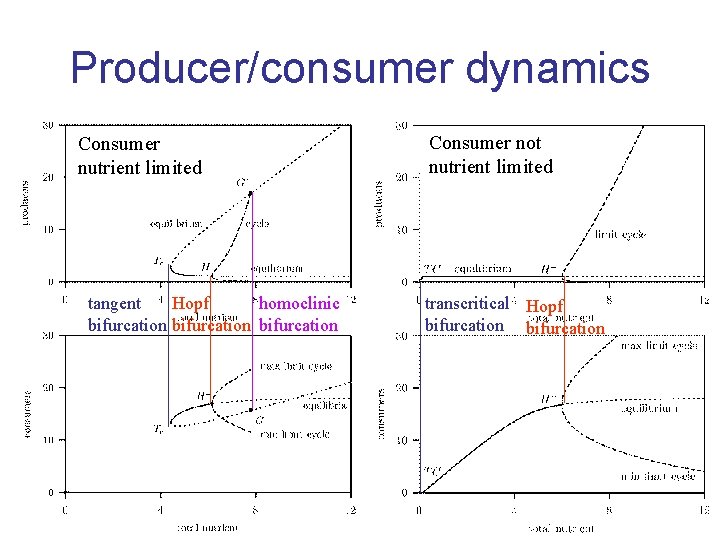
Producer/consumer dynamics Consumer nutrient limited tangent Hopf homoclinic bifurcation Consumer not nutrient limited transcritical Hopf bifurcation

Interactions of substrates Kooijman, 2001 Phil Trans R Soc B 356: 331 -349
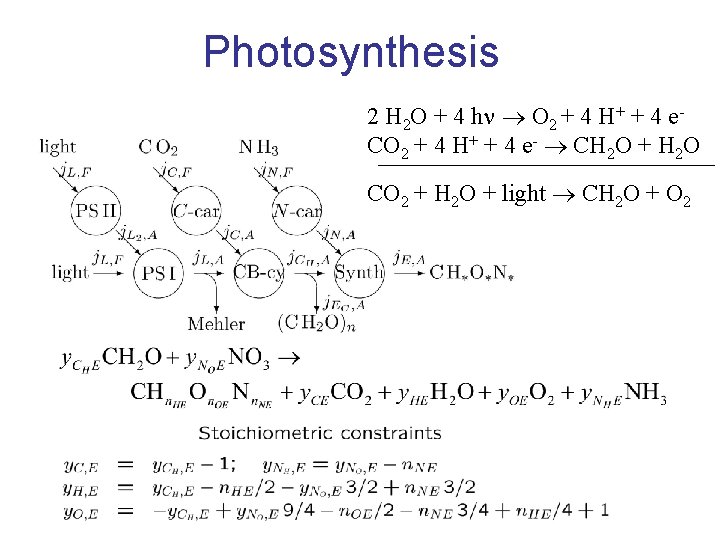
Photosynthesis 2 H 2 O + 4 h O 2 + 4 H+ + 4 e. CO 2 + 4 H+ + 4 e- CH 2 O + H 2 O CO 2 + H 2 O + light CH 2 O + O 2

Photorespiration (C 5 + C 2 C 3) (C 5 C 3 + C 2 ) Ru. P 2 ribulose 1, 5 -biphosphate Transformations are catalized by Rubisco, which evolved in anaerobic environments O 2 competes with CO 2 which gives an oxidation, rather than a reduction no synthesis of hydrocarbons at compensation point

Co-metabolism Consider coupled transformations A C and B D Binding probability of B to free SU differs from that to SU-A complex
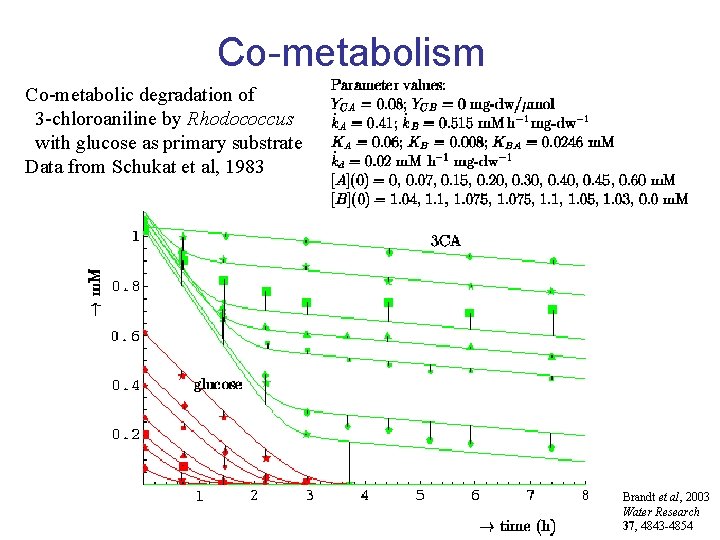
Co-metabolism Co-metabolic degradation of 3 -chloroaniline by Rhodococcus with glucose as primary substrate Data from Schukat et al, 1983 Brandt et al, 2003 Water Research 37, 4843 -4854
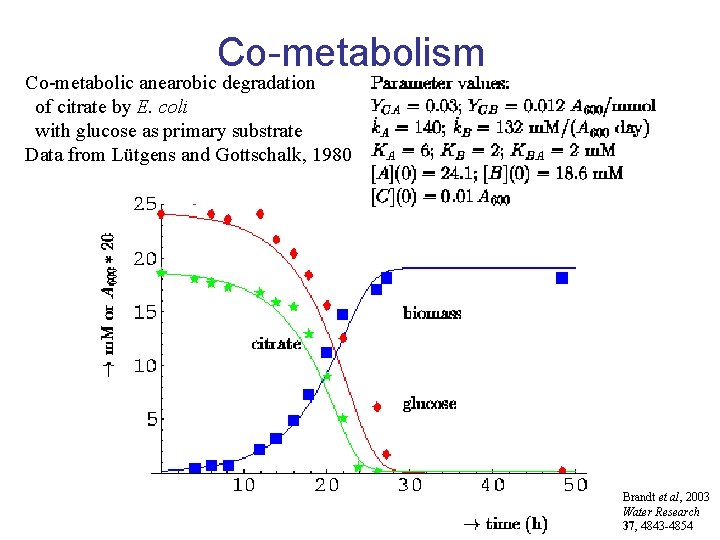
Co-metabolism Co-metabolic anearobic degradation of citrate by E. coli with glucose as primary substrate Data from Lütgens and Gottschalk, 1980 Brandt et al, 2003 Water Research 37, 4843 -4854
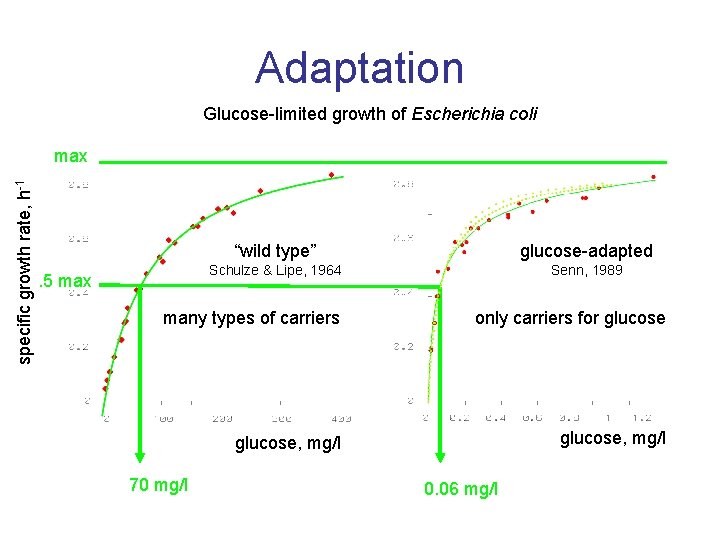
Adaptation Glucose-limited growth of Escherichia coli specific growth rate, h-1 max . 5 max “wild type” glucose-adapted Schulze & Lipe, 1964 Senn, 1989 many types of carriers only carriers for glucose, mg/l 70 mg/l 0. 06 mg/l

Inhibition A does not affect B in y. ACA C; B inhibits binding of A A inhibits binding of B in y. ACA C; B inhibits binding of A unbounded fraction binding prob of A arrival rate of A dissociation rate of A yield of C on A

Adaptation Batch culture, Monod special case Model elements: • uptake of substrate by specific carriers • carrier densities n. A and n. B • metabolic signals from uptake fini • relative signal s. A = p. A f. An. A/ i pi fini • carrier production by SUs that are fed by relative signals that inhibit reciprocally • carriers have a common turnover rate Result: Expression fraction 0 asymptotically in absence of substrate biomass density substrate i conc scaled func response saturation coeff for i yield of biom on substr Brandt et al, 2004 Water Research, 38, 1003 - 1013 spec growth rate max spec growth rate on i expression fraction for i carrier turnover rate preference ratio
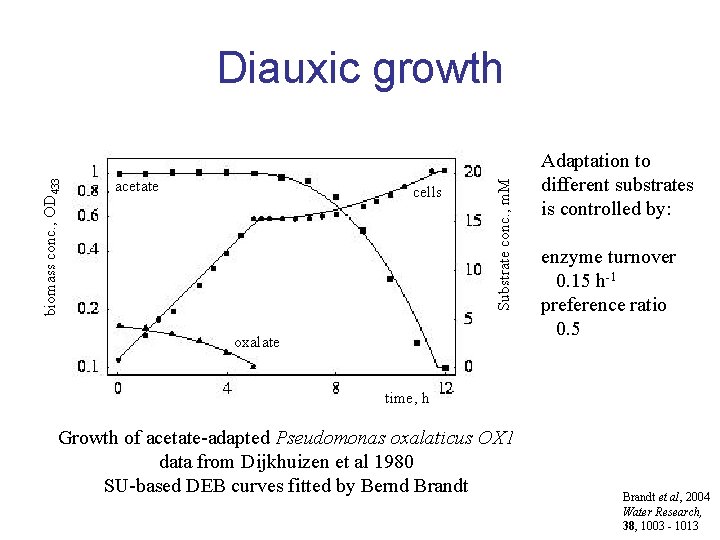
acetate cells Substrate conc. , m. M biomass conc. , OD 433 Diauxic growth oxalate Adaptation to different substrates is controlled by: enzyme turnover 0. 15 h-1 preference ratio 0. 5 time, h Growth of acetate-adapted Pseudomonas oxalaticus OX 1 data from Dijkhuizen et al 1980 SU-based DEB curves fitted by Bernd Brandt et al, 2004 Water Research, 38, 1003 - 1013
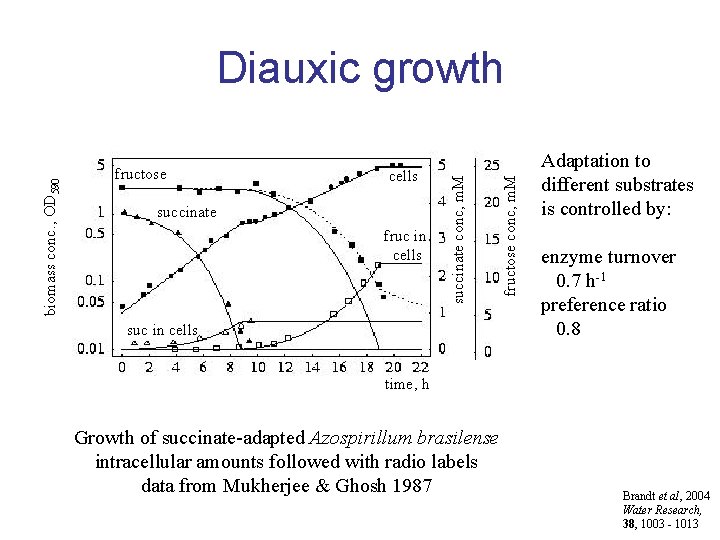
cells succinate fruc in cells suc in cells fructose conc, m. M fructose succinate conc, m. M biomass conc. , OD 590 Diauxic growth Adaptation to different substrates is controlled by: enzyme turnover 0. 7 h-1 preference ratio 0. 8 time, h Growth of succinate-adapted Azospirillum brasilense intracellular amounts followed with radio labels data from Mukherjee & Ghosh 1987 Brandt et al, 2004 Water Research, 38, 1003 - 1013
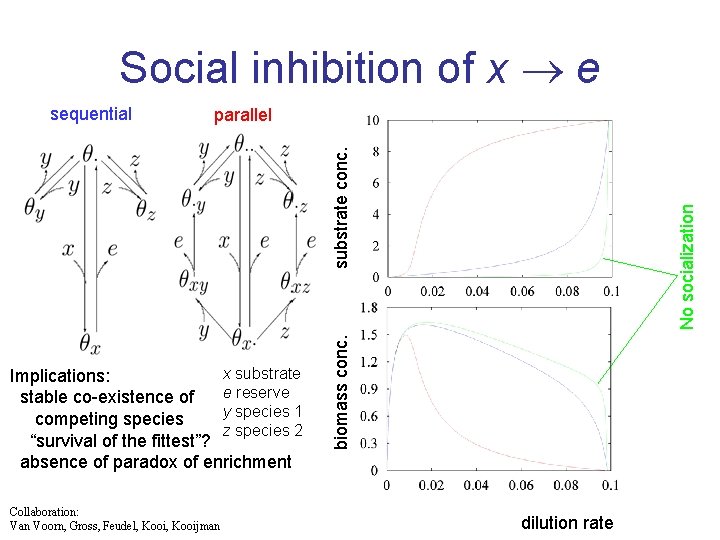
Social inhibition of x e parallel Collaboration: Van Voorn, Gross, Feudel, Kooijman biomass conc. x substrate Implications: e reserve stable co-existence of y species 1 competing species z species 2 “survival of the fittest”? absence of paradox of enrichment No socialization substrate conc. sequential dilution rate

Aggressive competition JEM, JVM V structure; E reserve; M maintenance substrate priority E M; posteriority V M JE flux mobilized from reserve specified by DEB theory JV flux mobilized from structure amount of structure (part of maint. ) excess returns to structure k. V dissociation rate SU-V complex k. E dissociation rate SU-E complex k. V k. E depend on such that k. M = y. MEk. E( E. + EV)+y. MVk. V is constant Collaboration: Tolla, Poggiale, Auger, Kooijman k. V = k. E k. V < k. E JE
- Slides: 33