Synapses Signal is carried chemically across the synaptic




![MAP and ML MAP: s* which maximizes p[s|r] ML: s* which maximizes p[r|s] Difference MAP and ML MAP: s* which maximizes p[s|r] ML: s* which maximizes p[r|s] Difference](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-14.jpg)
![Need to know full P[r|s] Assume Poisson: Assume independent: Population response of 11 cells Need to know full P[r|s] Assume Poisson: Assume independent: Population response of 11 cells](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-16.jpg)
![Apply ML: maximise P[r|s] with respect to s Set derivative to zero, use sum Apply ML: maximise P[r|s] with respect to s Set derivative to zero, use sum](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-17.jpg)
![Apply MAP: maximise p[s|r] with respect to s Set derivative to zero, use sum Apply MAP: maximise p[s|r] with respect to s Set derivative to zero, use sum](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-18.jpg)




- Slides: 25

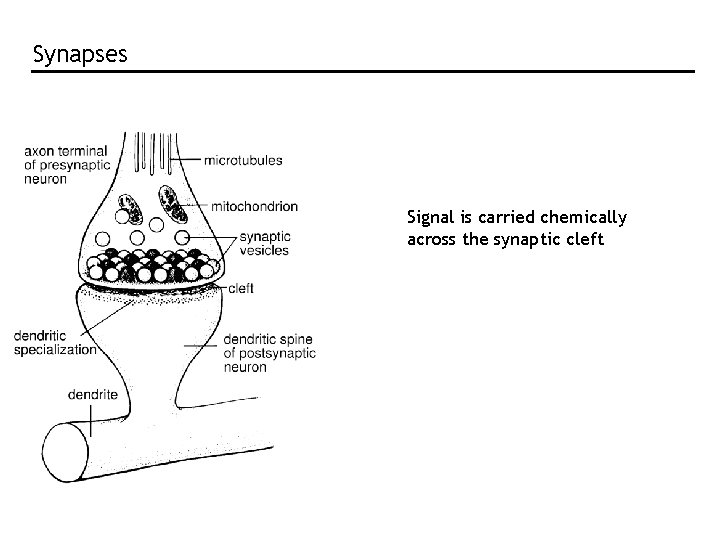
Synapses Signal is carried chemically across the synaptic cleft

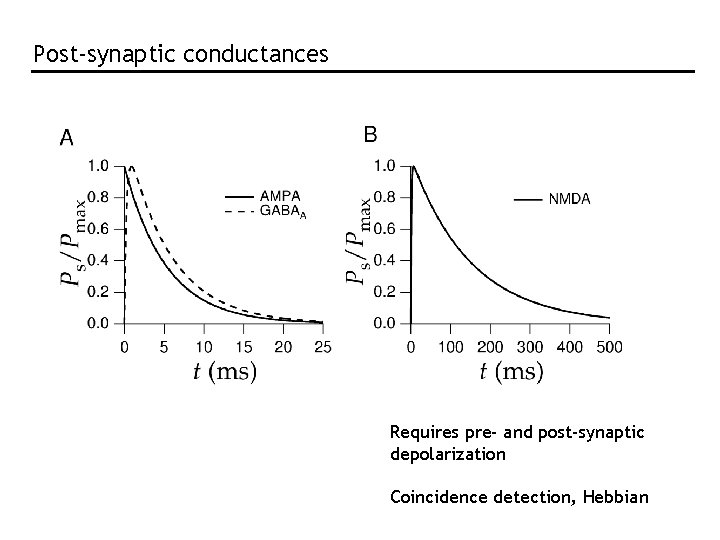
Post-synaptic conductances Requires pre- and post-synaptic depolarization Coincidence detection, Hebbian

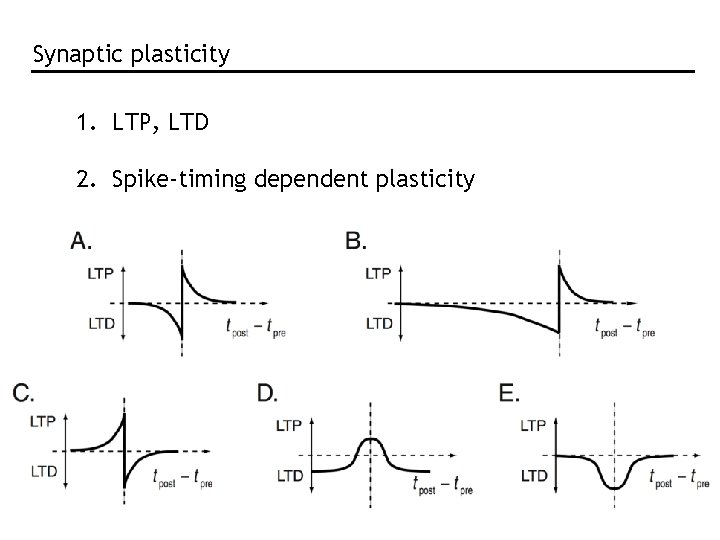
Synaptic plasticity 1. LTP, LTD 2. Spike-timing dependent plasticity

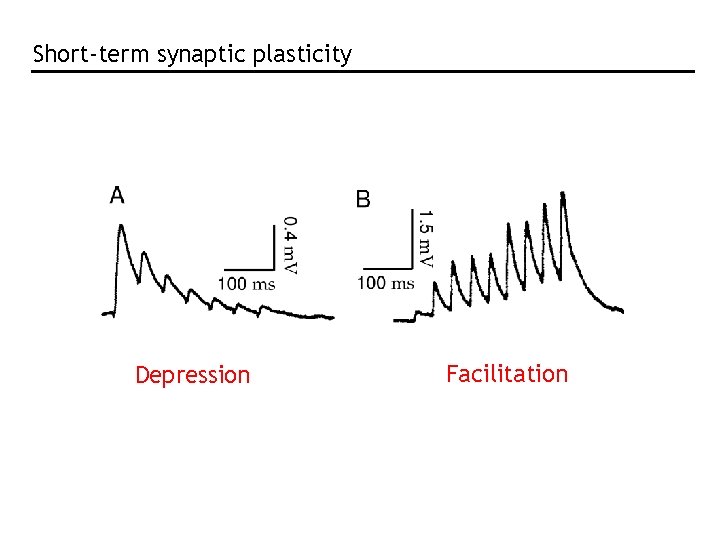
Short-term synaptic plasticity Depression Facilitation

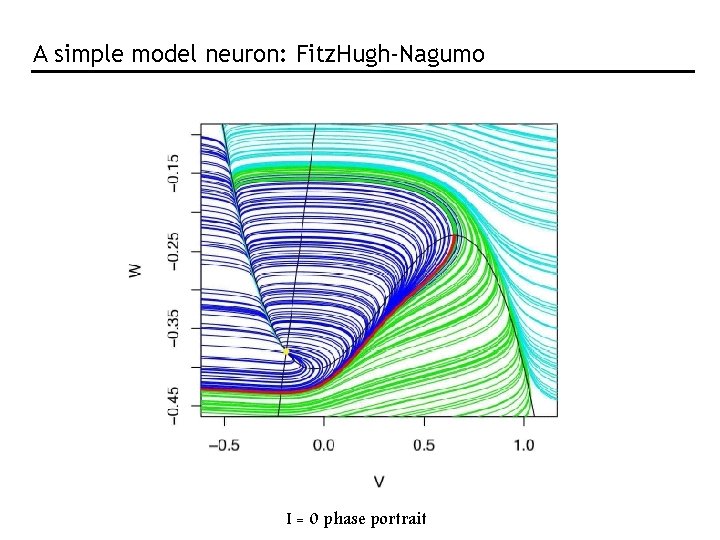
A simple model neuron: Fitz. Hugh-Nagumo I = 0 phase portrait

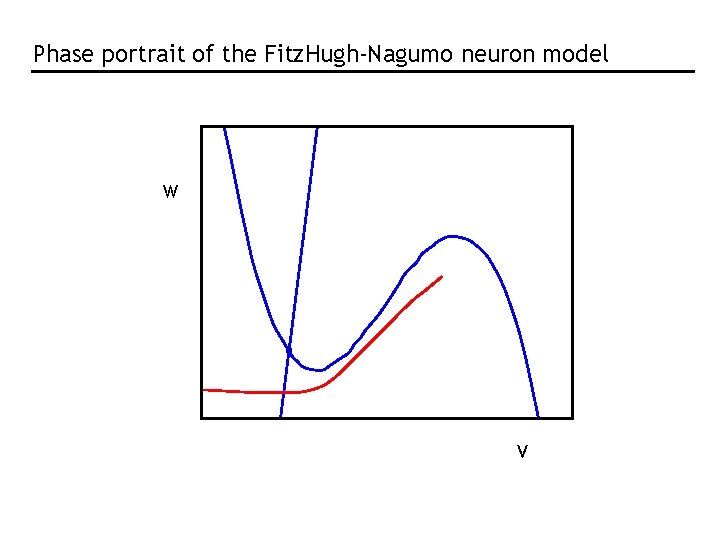
Phase portrait of the Fitz. Hugh-Nagumo neuron model W V

Reduced dynamical model for neurons

Population coding • Population code formulation • Methods for decoding: population vector Bayesian inference maximum a posteriori maximum likelihood • Fisher information

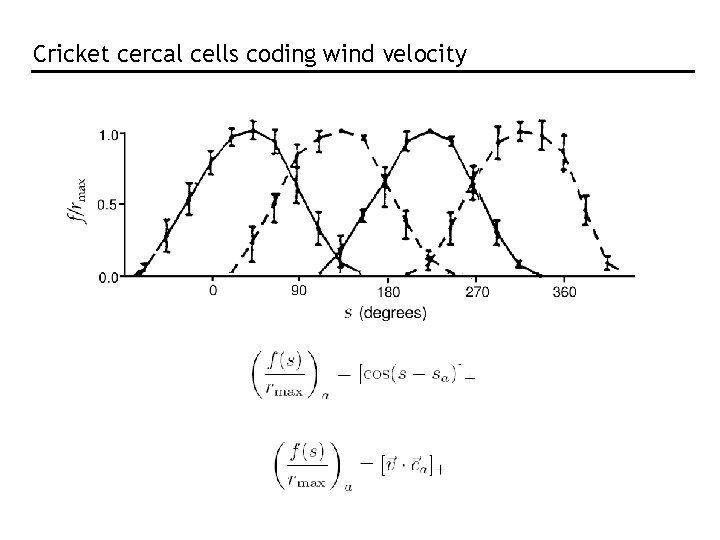
Cricket cercal cells coding wind velocity

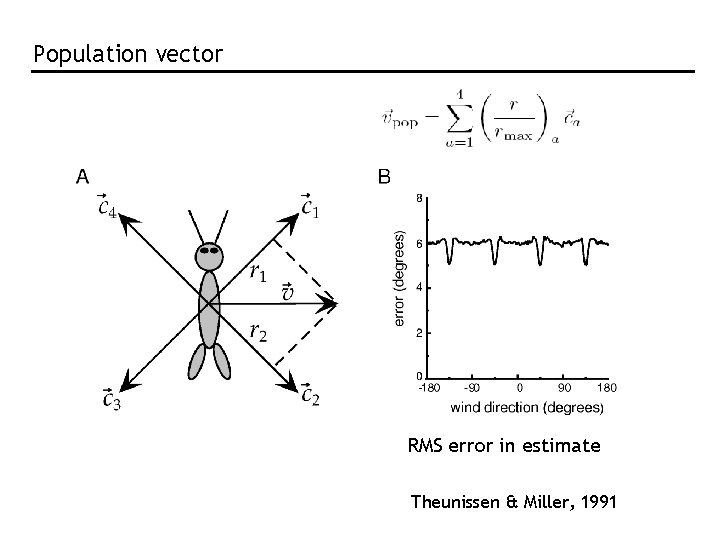
Population vector RMS error in estimate Theunissen & Miller, 1991

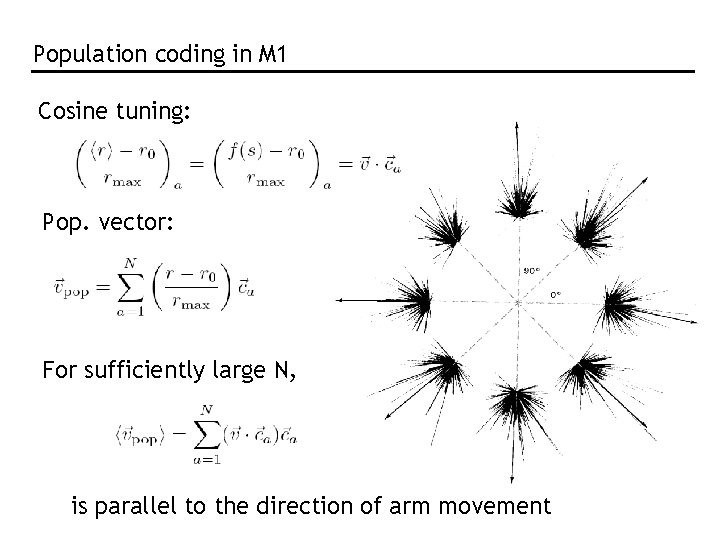
Population coding in M 1 Cosine tuning: Pop. vector: For sufficiently large N, is parallel to the direction of arm movement

The population vector is neither general nor optimal. “Optimal”: Bayesian inference and MAP

Bayesian inference By Bayes’ law, Introduce a cost function, L(s, s. Bayes); minimise mean cost. For least squares, L(s, s. Bayes) = (s – s. Bayes)2 ; solution is the conditional mean.
![MAP and ML MAP s which maximizes psr ML s which maximizes prs Difference MAP and ML MAP: s* which maximizes p[s|r] ML: s* which maximizes p[r|s] Difference](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-14.jpg)
MAP and ML MAP: s* which maximizes p[s|r] ML: s* which maximizes p[r|s] Difference is the role of the prior: differ by factor p[s]/p[r] For cercal data:

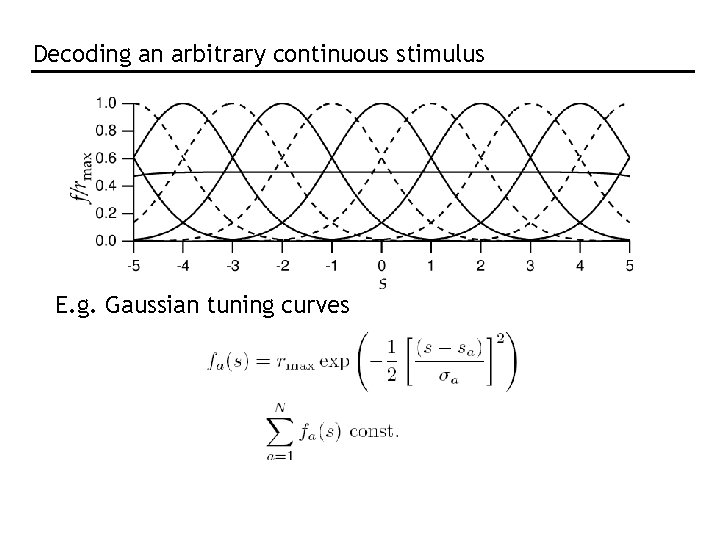
Decoding an arbitrary continuous stimulus E. g. Gaussian tuning curves
![Need to know full Prs Assume Poisson Assume independent Population response of 11 cells Need to know full P[r|s] Assume Poisson: Assume independent: Population response of 11 cells](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-16.jpg)
Need to know full P[r|s] Assume Poisson: Assume independent: Population response of 11 cells with Gaussian tuning curves
![Apply ML maximise Prs with respect to s Set derivative to zero use sum Apply ML: maximise P[r|s] with respect to s Set derivative to zero, use sum](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-17.jpg)
Apply ML: maximise P[r|s] with respect to s Set derivative to zero, use sum = constant From Gaussianity of tuning curves, If all s same
![Apply MAP maximise psr with respect to s Set derivative to zero use sum Apply MAP: maximise p[s|r] with respect to s Set derivative to zero, use sum](https://slidetodoc.com/presentation_image_h2/ecec867498c72e2dcd7c9873cf9c6d86/image-18.jpg)
Apply MAP: maximise p[s|r] with respect to s Set derivative to zero, use sum = constant From Gaussianity of tuning curves,

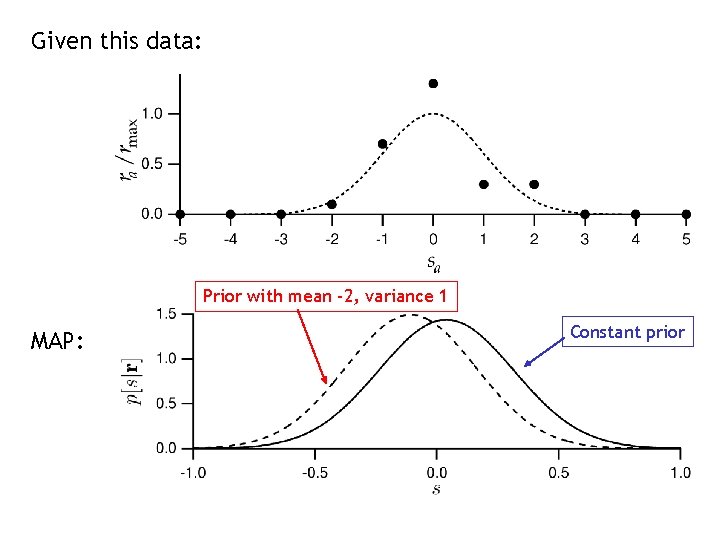
Given this data: Prior with mean -2, variance 1 MAP: Constant prior

How good is our estimate? For stimulus s, have estimated sest Bias: Variance: Mean square error: Cramer-Rao bound: Fisher information

Fisher information Alternatively: For the Gaussian tuning curves w/Poisson statistics:

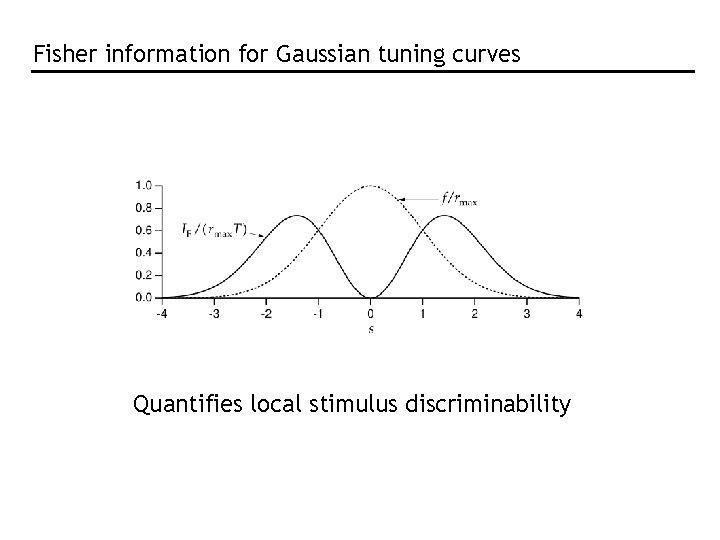
Fisher information for Gaussian tuning curves Quantifies local stimulus discriminability

Do narrow or broad tuning curves produce better encodings? Approximate: Thus, Narrow tuning curves are better But not in higher dimensions!

Fisher information and discrimination Recall d' = mean difference/standard deviation Can also decode and discriminate using decoded values. Trying to discriminate s and s+Ds: Difference in estimate is Ds (unbiased) variance in estimate is 1/IF(s).

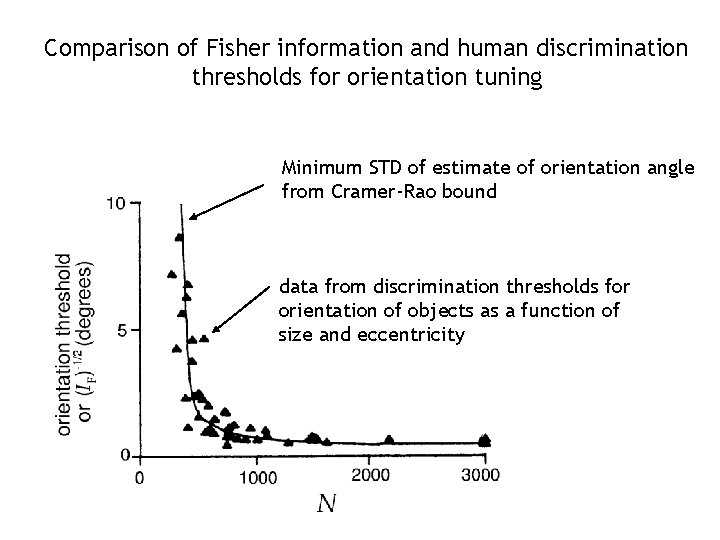
Comparison of Fisher information and human discrimination thresholds for orientation tuning Minimum STD of estimate of orientation angle from Cramer-Rao bound data from discrimination thresholds for orientation of objects as a function of size and eccentricity