SWISSPROT The SWISSPROT database consists of sequence entries

SWISS-PROT • The SWISS-PROT database consists of sequence entries. It contains high-quality annotation, is non-redundant and crossreferenced to many other databases. • Release 39. 0 of SWISS-PROT contains 86, 593 sequence entries. • SWISS-PROT is accompanied by Tr. EMBL, a computerannotated supplement to SWISS-PROT. Tr. EMBL contains the translations of all coding sequences (CDS) present in the EMBL Nucleotide Sequence Database, which are not yet integrated into SWISS-PROT. •

Tr. EMBL • Tr. EMBL release 17 (June 2001) was created from the EMBL Nucleotide Sequence Database release 66 and updates up to 01. 05. 01 and contains 540, 195 sequence entries, comprising 155, 771, 315 amino acids. • Tr. EMBL is split into two main sections; SP-Tr. EMBL and REMTr. EMBL. SP-Tr. EMBL (SWISS-PROT Tr. EMBL) contains the entries which should eventually be incorporated into SWISSPROT and can be considered as a preliminary section of SWISS-PROT as all SP-Tr. EMBL entries have been assigned SWISS-PROT accession numbers. REM-Tr. EMBL (REMaining Tr. EMBL) contains the entries that we do not want to include in SWISS-PROT. REM-Tr. EMBL entries have no accession numbers.

Protein Information Resource (PIR) The Protein Information Resource (PIR), in collaboration with MIPS and JIPID, produces the PIR-International Protein Sequence Database (PIR-PSD) -- a comprehensive, non-redundant, expertly annotated, fully classified and extensively cross-referenced protein sequence database in the public domain. The PIR-PSD, i. Pro. Class and other PIR auxiliary databases provide an integration of sequences, functional, and structural information to support genomics and proteomics research.

Nyhet: Uni. Prot (Universal Protein Resource) is the world's most comprehensive catalogue of information on proteins. It is a central repository of protein sequence and function created by joining the information contained in Swiss-Prot, Tr. EMBL, and PIR. Uni. Prot is comprised of three components, each optimised for different uses. The Uni. Prot Knowledgebase (Uni. Prot) is the central access point for extensive curated protein information, including function, classification, and cross-reference. The Uni. Prot Non-redundant Reference (Uni. Ref) databases combine closely related sequences into a single record to speed searches. The Uni. Prot Archive (Uni. Parc) is a comprehensive repository, reflecting the history of all protein sequences. The sequences and information in Uni. Prot is accessible via text search, BLAST similarity search, and FTP.

Entrez at NCBI Entrez is a retrieval system for searching several linked databases. It provides access to: Pub. Med: The biomedical literature (Pub. Med) Nucleotide sequence database (Genbank) Protein sequence database Structure: three-dimensional macromolecular structures Genome: complete genome assemblies Pop. Set: population study data sets OMIM: Online Mendelian Inheritance in Man Taxonomy: organisms in Gen. Bank Books: online books Probe. Set: Gene Expression Omnibus (GEO) 3 D Domains: domains from Entrez Structure • Go to NCBI
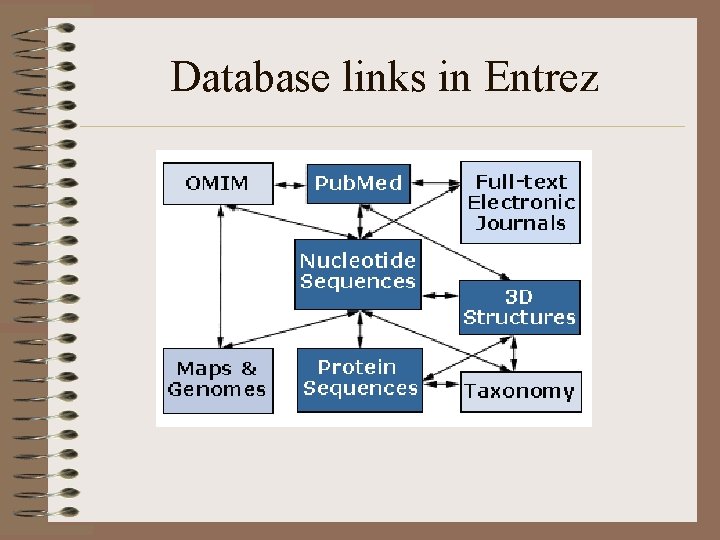
Database links in Entrez

SRS at EBI SRS is a powerful data integration platform, providing rapid, easy and user friendly access to the large volumes of diverse and heterogeneous Life Science data stored in more than 400 internal and public domain databases. SRS enables the querying of diverse biological and Life Science data through only one interface, SRS facilitates the rapid development of applications and algorithms, as well as bioinformatics portals for the Inter- or Intranet, making the data efficiently available to entire organizations. Today, SRS is answering the most demanding requirements of modern Life Science companies and will truly add value to their research programs.

SRS enables: ØFast access to diverse life science data - genetic, protein, cellular, molecular, and clinical - for researchers and bioinformaticians ØIntegration of public and proprietary data through one interface ØUnique ability to perform cross-database queries ØRapid string search of large volumes of data ØScalability to the customer's specific requirements EBI, the European Bioinformatics Institute (EMBL Outstation, Hinxton, UK)

Forskjellige sekvensformater Her er en sekvens i GCG-format EXTRACTPEPTIDE of frames: C from: caupol. map (Linear) MAP of: caupol. raw check: 2457 from: 1 to: 3957 Frame C from: 1 to: 1318 caupol. pep Length: 941 August 27, 1995 16: 35 Type: P Check: 9501. . 1 MAYPLLVLVD GHALAYRAFF ALRESGLRSS RGEPTYAVFG FAQILLTALA 51 EYRPDYAAVA FDVGRTFRDD LYAEYKAGRA ETPEEFYPQF ERIKQLVQAL 101 NIPIYTAEGY EADDVIGTLA RQATERGVDT IILTGDSDVL QLVNDHVRVA 151 LANPYGGKTS VTLYDLEQVR KRYDGLEPDQ LADLRGLKGD TSDNIPGVRG Her er en annen i FASTA-format >ECPOLA V 00317 E. coli gene pol. A coding for DNA polymerase I. 9/93 CACCGGGCAACGGCGGCAGAAGTGTTTGCCACTGGAAACCGTCACCAGCGAGCAA CGCCGTAGCGCGAAAGCGATCAACTTTGGTCTGATTTATGGCATGAGTGCTTTCGGTCTG GCGCGGCAATTGAACATTCCACGTAAAGAAGCGCAGAAGTACATGGACCTTTACTTCGAA CGCTACCCTGGCGTGCTGGAGTATATGGAACGCACCCGTGCTCAGGCGAAAGAGCAGGGC TACGTTGAAACGCTGGACGCCGTCTGTATCTGCCGGATATCAAATCCAGCAATGGT GCTCGTCGTGCAGCGGCTGAACGTGCAGCCATTAACGCGCCAATGCAGGGAACCGCCGCC GA Mens dette er et eksempel på en ren tekstfil CGCCGTAGCGCGAAAGCGATCAACTTTGGTCTGATTTATGGCATGAGTGCTTTCGGTCTG GCGCGGCAATTGAACATTCCACGTAAAGAAGCGCAGAAGTACATGGACCTTTACTTCGAA CGCTACCCTGGCGTGCTGGAGTATATGGAACGCACCCGTGCTCAGGCGAAAGAGCAGGGC TACGTTGAAACGCTGGACGCCGTCTGTATCTGCCGGATATCAAATCCAGCAATGGT GCTCGTCGTGCAGCGGCTGAACGTGCAGCCATTAACGCGCCAATGCAGGGAACCGCCGCC GA
Hvordan oversette fra et format til et annet? Read. Seq http: //www. ebi. ac. uk/cgi-bin/readseq. cgi Read. Seq kan oversette fra og til 21 forskjellige sekvensformater
- Slides: 10