Swarms and Bundles Bioinformatics and Biostatistics on Biowulf

Swarms and Bundles: Bioinformatics and Biostatistics on Biowulf David Hoover Scientific Computing Branch, Division of Computer System Services CIT, NIH

Embarrassingly Parallel Problems • • • GWAS, with huge numbers of SNPs Sequence analysis, assembly, and mapping Testing and validating statistical models Protein folding and threading Molecular docking and compound screening Tomographic reconstruction
Characterization of Surface Protein 3 from Malaria Parasite P. Falciparum Protein folding calculations with Rosetta++ 100, 000 cpu hours Tsai et al. , Mol. Biochem. Parasitology, online preprint 2008
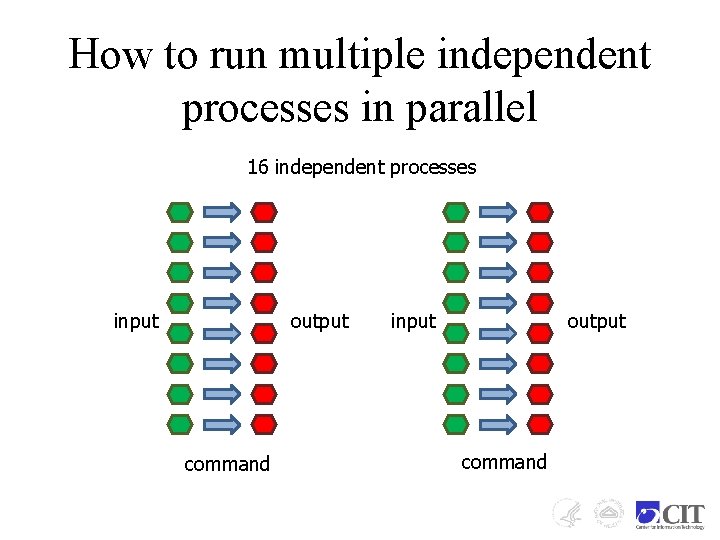
How to run multiple independent processes in parallel 16 independent processes input output command
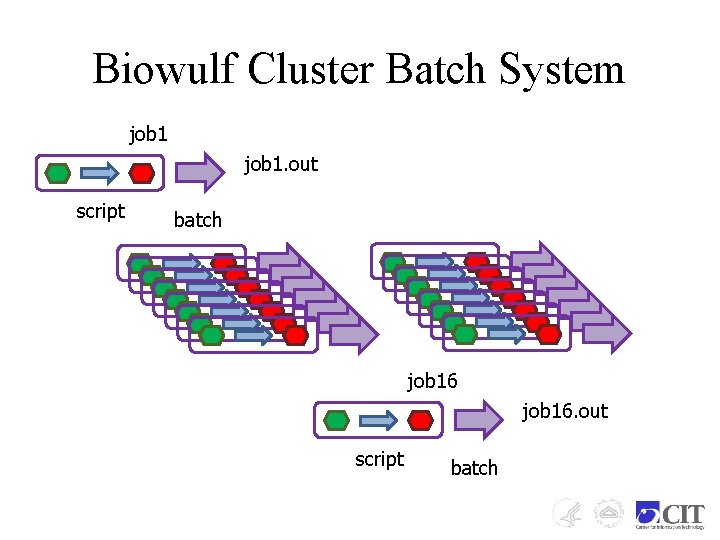
Biowulf Cluster Batch System job 1. out script batch job 16. out script batch

Swarm biowulf% swarm -f file job 1 job 2 Node 1 job 1. out job 3 Node 2 job 2. out job 4 Node 3 job 3. out Node 4 job 4. out
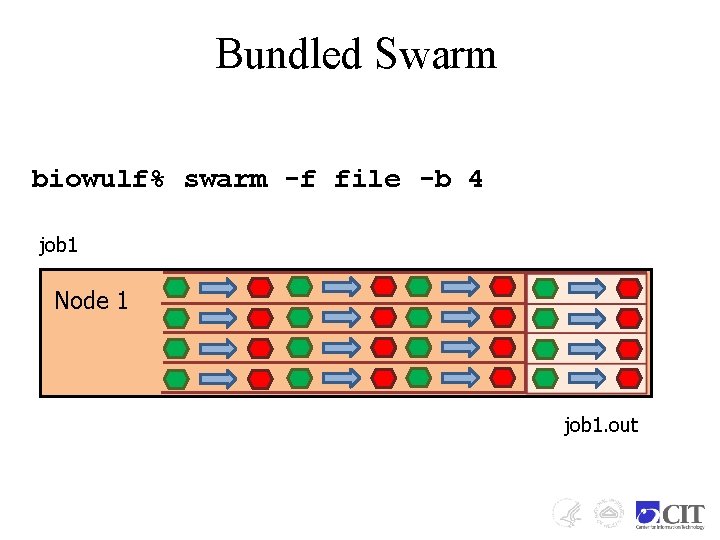
Bundled Swarm biowulf% swarm -f file -b 4 job 1 Node 1 job 1. out

Swarm Facts • Written and maintained by Helix Systems Staff • swarm introduced in late 2000 • 82% of all batch jobs run on the cluster since 2002 are swarm jobs • ~60% of all wall time spent on swarm jobs • swarm has been shared with clusters around the world

Swarm World Records • Largest swarm: 683, 445 commands • Largest bundle: 24, 000 commands per CPU

Future Challenges • How to deal with larger multicore nodes? Node 1 Node 2 Node 3
- Slides: 10