Surfacebased Group Analysis in Free Surfer 1 Outline

Surface-based Group Analysis in Free. Surfer 1

Outline • Processing Stages • Command-line Stream • Assemble Data • Design/Contrast (GLM Theory) • Analyze • Visualize • Interactive/Automated GUI (QDEC) • Correction for multiple comparisons 2
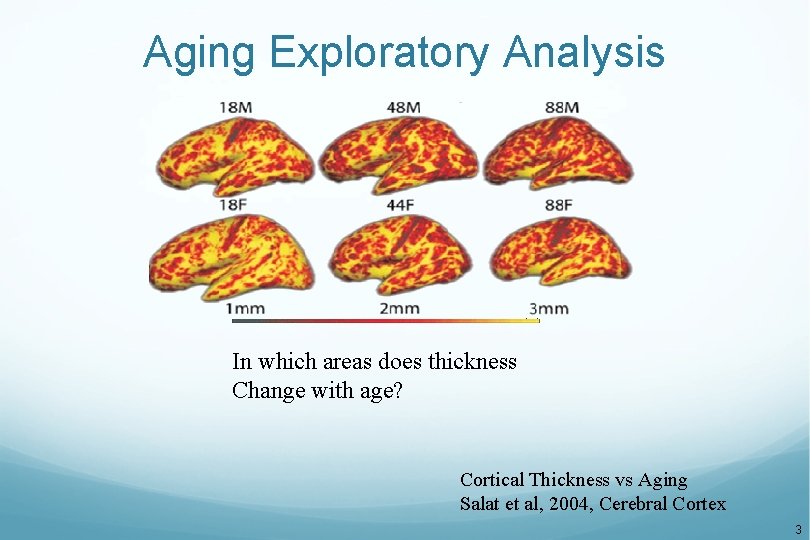
Aging Exploratory Analysis In which areas does thickness Change with age? Cortical Thickness vs Aging Salat et al, 2004, Cerebral Cortex 3

Aging Thickness Study N=40 (all in fsaverage space) Negative Age Correlation p<. 01 Positive Age Correlation 4

Surface-based Measures Morphometric (e. g. , thickness) Functional PET MEG/EEG Diffusion (? ) sampled just under the surface 5

Processing Stages Specify Subjects and Surface measures Assemble Data: Resample into Common Space Smooth Concatenate into one file Model and Contrasts (GLM) Fit Model (Estimate) Correct for multiple comparisons Visualize 6

The General Linear Model (GLM) 7
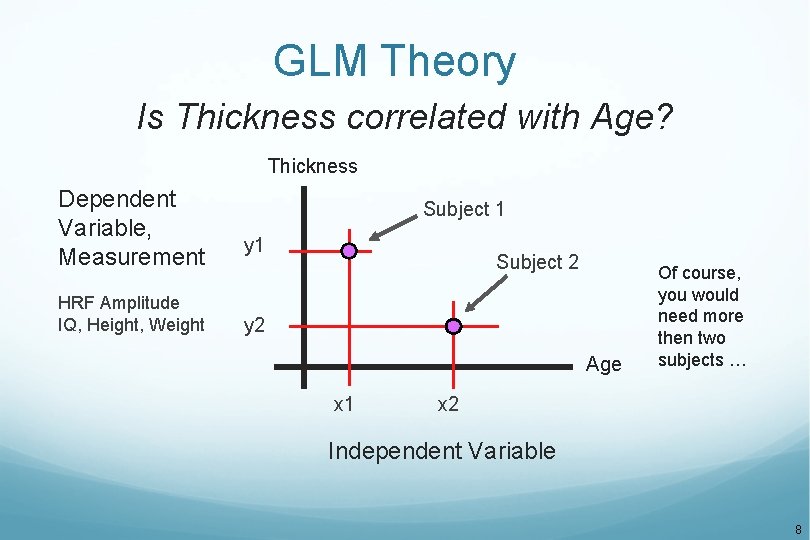
GLM Theory Is Thickness correlated with Age? Thickness Dependent Variable, Measurement HRF Amplitude IQ, Height, Weight Subject 1 y 1 Subject 2 y 2 Age x 1 Of course, you would need more then two subjects … x 2 Independent Variable 8
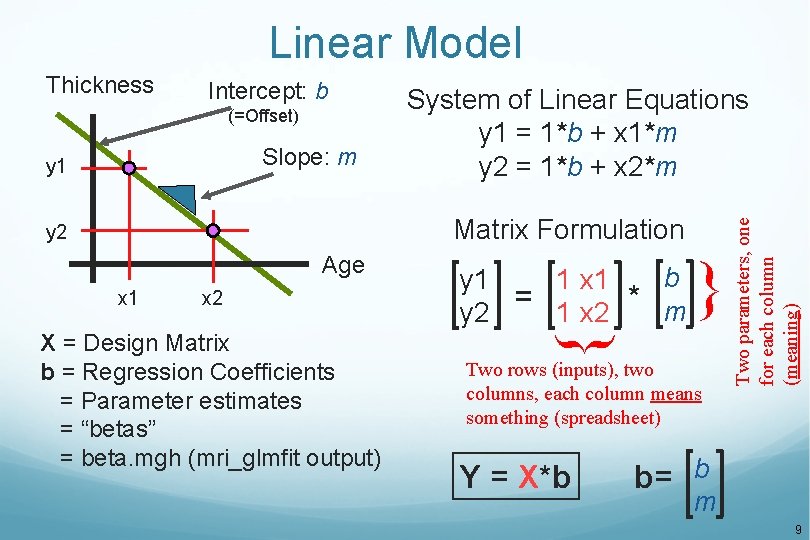
Linear Model Intercept: b (=Offset) Slope: m y 1 System of Linear Equations y 1 = 1*b + x 1*m y 2 = 1*b + x 2*m Matrix Formulation y 2 Age x 1 x 2 } } X = Design Matrix b = Regression Coefficients = Parameter estimates = “betas” = beta. mgh (mri_glmfit output) b 1 x 1 y 1 = * m 1 x 2 y 2 Two rows (inputs), two columns, each column means something (spreadsheet) Y = X*b Two parameters, one for each column (meaning) Thickness b= b m 9
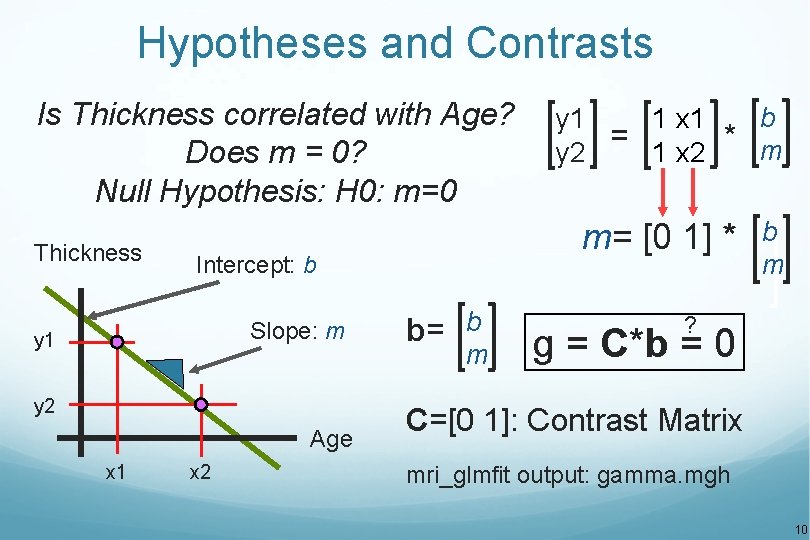
Hypotheses and Contrasts Is Thickness correlated with Age? Does m = 0? Null Hypothesis: H 0: m=0 Thickness y 1 y 2 Age x 1 m= [0 1] * Intercept: b Slope: m x 2 b 1 x 1 y 1 = * m 1 x 2 y 2 b= b m ? g = C*b = 0 C=[0 1]: Contrast Matrix mri_glmfit output: gamma. mgh 10

More than Two Data Points Thickness Intercept: b Slope: m Age y 1 = 1*b + x 1*m + n 1 y 2 = 1*b + x 2*m + n 2 y 3 = 1*b + x 3*m + n 3 y 4 = 1*b + x 4*m + n 4 b + n 1 y 1 1 x 1 = 1 x 2 * m n 2 y 2 n 3 y 3 1 x 3 n 4 y 4 1 x 4 Y = X*b+n • Model Error • Noise • Residuals • eres. mgh 11

t-Test and p-values Y = X*b+n g = C*b p-value/significance • value between 0 and 1 • closer to 0 means more significant Free. Surfer stores p-values as –log 10(p): • 0. 1=10 -1 sig=1, 0. 01=10 -2 sig=2 • sig. mgh files • Signed by sign of g • p-value is for an unsigned test 12
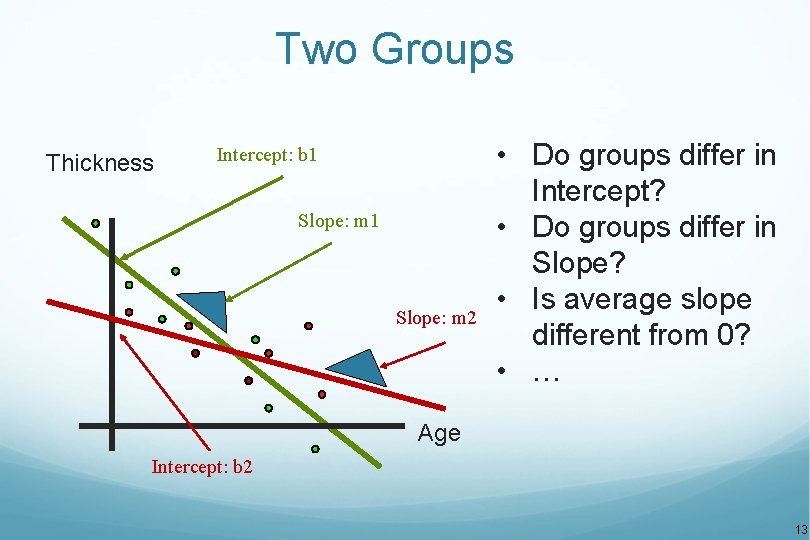
Two Groups Thickness Intercept: b 1 Slope: m 2 • Do groups differ in Intercept? • Do groups differ in Slope? • Is average slope different from 0? • … Age Intercept: b 2 13
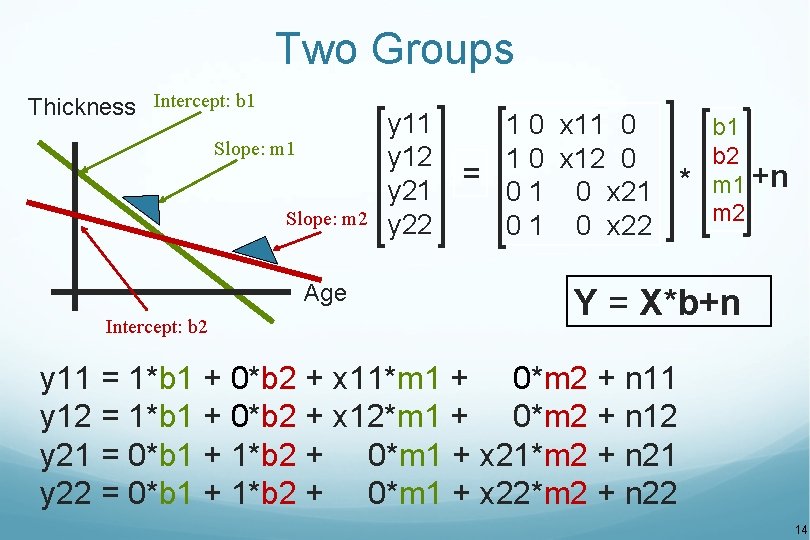
Two Groups Thickness Intercept: b 1 Slope: m 2 Age Intercept: b 2 y 11 y 12 y 21 y 22 1 0 x 11 0 b 1 b 2 1 0 x 12 0 = +n * m 1 0 x 21 m 2 0 1 0 x 22 Y = X*b+n y 11 = 1*b 1 + 0*b 2 + x 11*m 1 + 0*m 2 + n 11 y 12 = 1*b 1 + 0*b 2 + x 12*m 1 + 0*m 2 + n 12 y 21 = 0*b 1 + 1*b 2 + 0*m 1 + x 21*m 2 + n 21 y 22 = 0*b 1 + 1*b 2 + 0*m 1 + x 22*m 2 + n 22 14
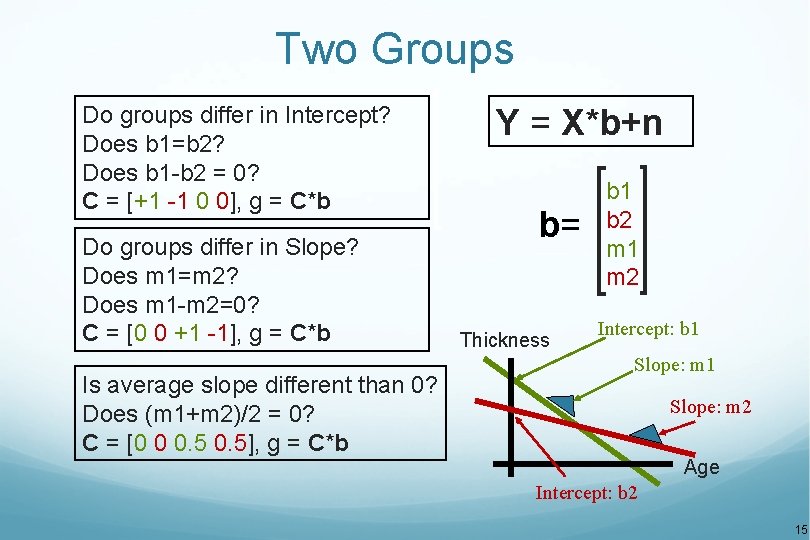
Two Groups Do groups differ in Intercept? Does b 1=b 2? Does b 1 -b 2 = 0? C = [+1 -1 0 0], g = C*b Do groups differ in Slope? Does m 1=m 2? Does m 1 -m 2=0? C = [0 0 +1 -1], g = C*b Is average slope different than 0? Does (m 1+m 2)/2 = 0? C = [0 0 0. 5], g = C*b Y = X*b+n b= Thickness b 1 b 2 m 1 m 2 Intercept: b 1 Slope: m 2 Age Intercept: b 2 15

Surface-based Group Analysis in Free. Surfer Create your own design and contrast matrices Create an FSGD File Free. Surfer creates design matrix You still have to specify contrasts QDEC Limited to 2 discrete variables, 2 levels max Limited to 2 continuous variables 16

Command-line Processing Stages Assemble Data (mris_preproc) } Resample into Common Space recon-all -qcache Smooth Concatenate into one file Model and Contrasts (GLM) (FSGD) Fit Model (Estimate) (mri_glmfit) Correct for multiple comparisons Visualize (tksurfer) Run after all editing is done. 17
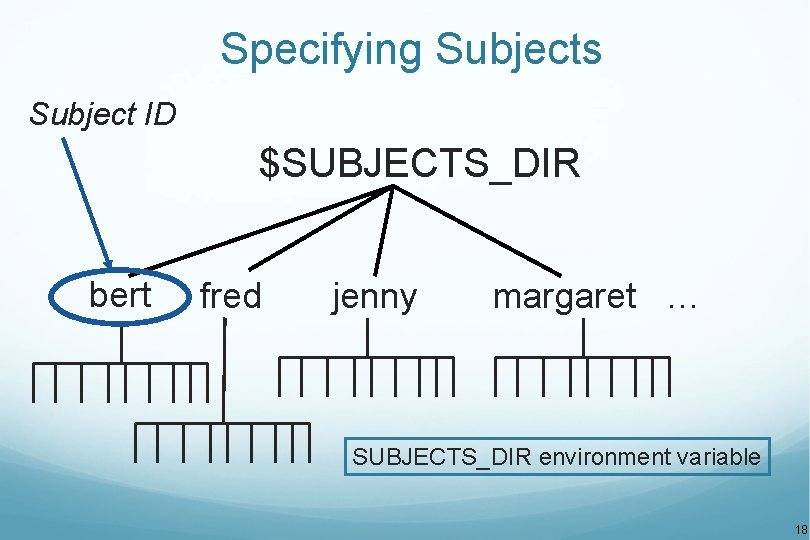
Specifying Subjects Subject ID $SUBJECTS_DIR bert fred jenny margaret … SUBJECTS_DIR environment variable 18
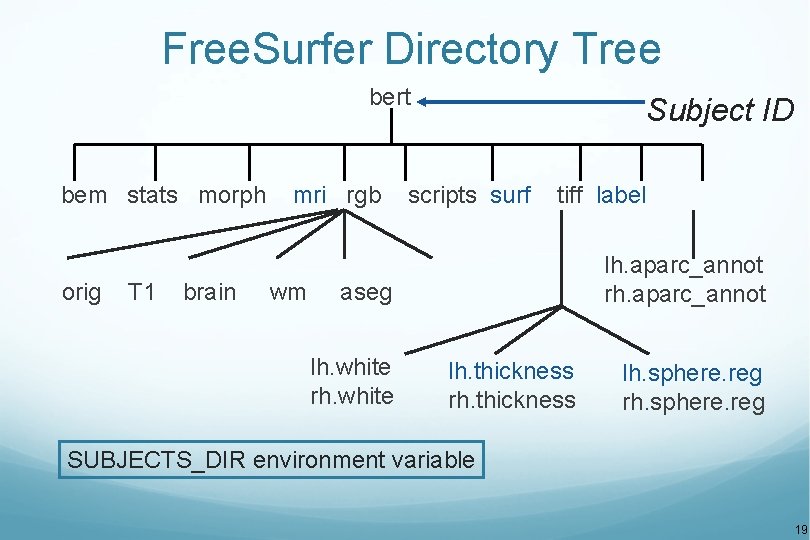
Free. Surfer Directory Tree bert bem stats morph orig T 1 brain mri rgb wm Subject ID scripts surf tiff label lh. aparc_annot rh. aparc_annot aseg lh. white rh. white lh. thickness rh. thickness lh. sphere. reg rh. sphere. reg SUBJECTS_DIR environment variable 19

Example: Thickness Study 1. $SUBJECTS_DIR/bert/surf/lh. thickness 2. $SUBJECTS_DIR/fred/surf/lh. thickness 3. $SUBJECTS_DIR/jenny/surf/lh. thickness 4. $SUBJECTS_DIR/margaret/surf/lh. thickness 5. … 20

Free. Surfer Group Descriptor (FSGD) File Simple text file List of all subjects in the study Accompanying demographics Automatic design matrix creation You must still specify the contrast matrices Integrated with tksurfer Note: Can specify design matrix explicitly with --design 21

FSGD Format Group. Descriptor. File 1 Class Male Class Female Variables Input bert Male 10 Input fred Male 15 Input jenny Female Input margaret Female 100 150 20 25 Age Weight 1000 1500 2000 2500 IQ • One Discrete Factor (Gender) with Two Levels (M&F) • Three Continuous Variables: Age, Weight, IQ Class = Group Note: Can specify design matrix explicitly with --design 22

FSGDF X (Automatic) Male Age Female Group Male Group X= 10 10 01 01 10 0 15 0 0 25 100 0 150 0 0 200 0 250 Age Weight [-1 1 0 } C= Female Age 0 0 0 1000 0 1500 0 0 2000 0 2500 IQ 0 0 ] Tests for the difference in intercept/offset between groups [ 0 0 -1 1 } C= 0 0] Tests for the difference in age slope between groups DODS – Different Offset, Different Slope 23

Another FSGD Example • Two Discrete Factors – Gender: Two Levels (M&F) – Handedness: Two Levels (L&R) • One Continuous Variable: Age Group. Descriptor. File 1 Class Male. Right Class Male. Left Class Female. Right Class Female. Left Variables Input bert Male. Left Input fred Male. Right Input jenny Female. Right 20 Input margaret Female. Left Class = Group 10 15 Age 25 24

Interaction Contrast • Two Discrete Factors (no continuous, for now) – Gender: Two Levels (M&F) – Handedness: Two Levels (L&R) b 4 L D 2 b 2 • Four Regressors (Offsets) – MR (b 1), ML (b 2), FR (b 3), FL (b 4) Group. Descriptor. File 1 Class Male. Right Class Male. Left Class Female. Right Class Female. Left Input bert Male. Left Input fred Male. Right Input jenny Female. Left Input margaret Female. Right R b 3 D 1 b 1 M F ? g = D 1 -D 2=0 g = (b 3 -b 1)- (b 4 -b 2) = -b 1+b 2+ b 3 -b 4 C = [-1 +1 +1 -1] 25

Factors, Levels, Groups Usually each Group/Class: Has its own Intercept Has its own Slope (for each continuous variable) NRegressors = NClasses * (NVariables+1) DODS Nregressors = NClasses + Nvariables DOSS Why is this important? Because you will need to create contrast matrices, and the contrast matrix must have Nregressor elements. 26

Factors, Levels, Groups, Classes Continuous Variables/Factors: Age, IQ, Volume, etc. Discrete Variables/Factors: Gender, Handedness, Diagnosis Levels of Discrete Variables: • Handedness: Left and Right • Gender: Male and Female • Diagnosis: Normal, MCI, AD Group or Class: Specification of All Discrete Factors • Left-handed Male MCI • Right-handed Female Normal 27

Assemble Data: mris_preproc --help --fsgd FSGDFile --hemi lh --meas thickness --target fsaverage --o lh. thickness. mgh : Specify subjects thru FSGD File : Process left hemisphere : subjectid/surf/hemi. thickness : common space is subject fsaverage : output “volume-encoded surface file” Lots of other options! Output: lh. thickness. mgh – file with stacked thickness maps for all subjects Input to Smoother or GLM 28

Surface Smoothing mri_surf 2 surf --help Loads stacked lh. thickness. mgh 2 D surface-based smoothing Specify FWHM (eg, fwhm = 10 mm) Saves stacked lh. thickness. sm 10. mgh Can be slow (~10 -60 min) recon-all –qcache (computes for each subject, run after you are finished editing subject) 29

mri_glmfit Reads in FSGD File and constructs X Reads in your contrasts (C 1, C 2, etc. ) Loads data (lh. thickness. sm 10. mgh) Fits GLM (ie, computes b) Computes contrasts (g=C*b) t or F ratios, significances Significance -log 10(p) (. 01 2, . 001 3) 30

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir mri_glmfit --help 31

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir • Input file (output from smoothing). • Stack of subjects, one frame per subject. 32

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir • Free. Surfer Group Descriptor File (FSGD) • Group membership • Covariates 33

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir • Contrast Matrices • Simple text/ASCII files • Test hypotheses 34

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir • Perform analysis on left hemisphere of fsaverage subject • Masks by fsaverage cortex. label • Computes FWHM in 2 D 35

mri_glmfit --y lh. thickness. sm 10. mgh --fsgd gender_age. txt --C age. mtx –C gender. mtx --surf fsaverage lh --cortex --glmdir lh. gender_age. glmdir Output directory: lh. gender_age. glmdir/ beta. mgh – parameter estimates rvar. mgh – residual error variance etc … age/ sig. mgh – -log 10(p), uncorrected gamma. mgh, F. mgh gender/ sig. mgh – -log 10(p), uncorrected gamma. mgh, F. mgh 36

Visualization with tksurfer Threshold: -log 10(p), Eg, 2=. 01 Saturation: -log 10(p), Eg, 5=. 00001 False Dicovery Rate Eg, . 01 View->Configure->Overlay File->Load. Overlay 37

Tutorial Command-line Stream • Create an FSGD File for a thickness study • Age and Gender • Run mris_preproc mri_surf 2 surf mri_glmfit 38
- Slides: 38