Support Vector Machines Classification Venables Ripley Section 12

Support Vector Machines Classification Venables & Ripley Section 12. 5 CSU Hayward Statistics 6601 Joseph Rickert & Timothy Mc. Kusick December 1, 2004 JBR 1

Support Vector Machine What is the SVM? The SVM is a generalization of the Optimal Hyperplane Algorithm JBR Why is the SVM important? z It allows the use of more similarity measures than the OHA z Through the use of kernel methods it works with non vector data 2

Simple Linear Classifier X=Rp f(x) = w. Tx + b Each x X is classified into 2 classes labeled y {+1, -1} y = 1 if f(x) 0 and y = -1 if f(x) < 0 S = {(x 1, y 1), (x 2, y 2), . . . } Given S, the problem is to learn f (find w and b). For each f check to see if all (xi, yi) are correctly classified i. e. yif(xi) 0 Choose f so that the number of errors is minimized JBR 3

But what if the training set is not linearly separable? f(x) = w. Tx + b defines two half planes {x: f(x) 1} and {x: f(x) -1} Classify with the “Hinge” loss function: c(f, x, y) = max(0, 1 -yf(x)) c (f, x, y) as distance from correct half plane If (x, y) is correctly classified with large confidence then c(f, x, y) = 0 w. Tx+b > 1 yf(x) 1: correct with large conf 0 yf(x) < 0: correct with small conf yf(x) < 0: misclassified yf(x) JBR 1 w. Tx+b < - 1 margin = 2/||w|| 4

SVMs combine requirements of large margin and few misclassifications by solving the problem: New formulation: min 1/2||w||2 + C c(f, xi, yi) w. r. t w, x and b z C is parameter that controls tradeoff between margin and misclassification z Large C small margins but more samples correctly classified with strong confidence z Technical difficulty: hinge loss function c(f, xi, yi) is not differentiable JBR Even better formulation: use slack variables xi z min 1/2||w||2 + C xi w. r. t w, x and b under the constraint xi c(f, xi, yi) (*) z But (*) is equivalent to xi 0 for i = 1. . . n T xi - 1 + yi(w xi + b) 0 z Solve this quadratic optimization problem with Lagrange Multipliers 5
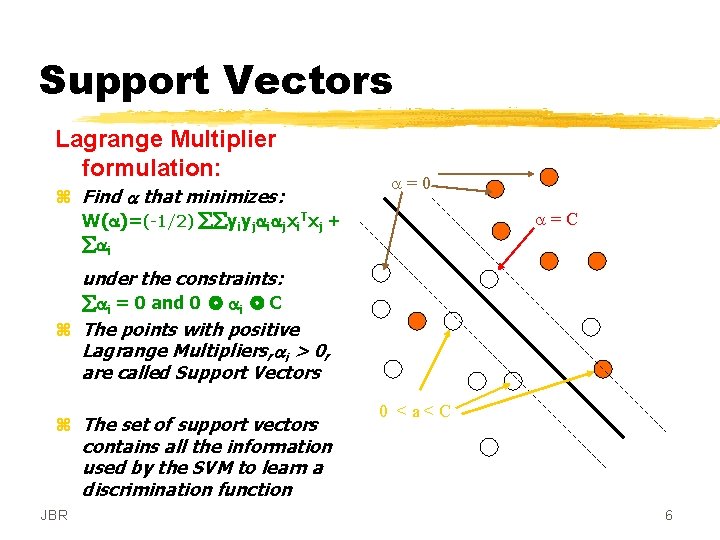
Support Vectors Lagrange Multiplier formulation: z Find a that minimizes: W(a)=(-1/2) yiyjaiajxi. Txj + ai a=0 a=C under the constraints: ai = 0 and 0 ai C z The points with positive Lagrange Multipliers, ai > 0, are called Support Vectors z The set of support vectors contains all the information used by the SVM to learn a discrimination function JBR 0 <a<C 6
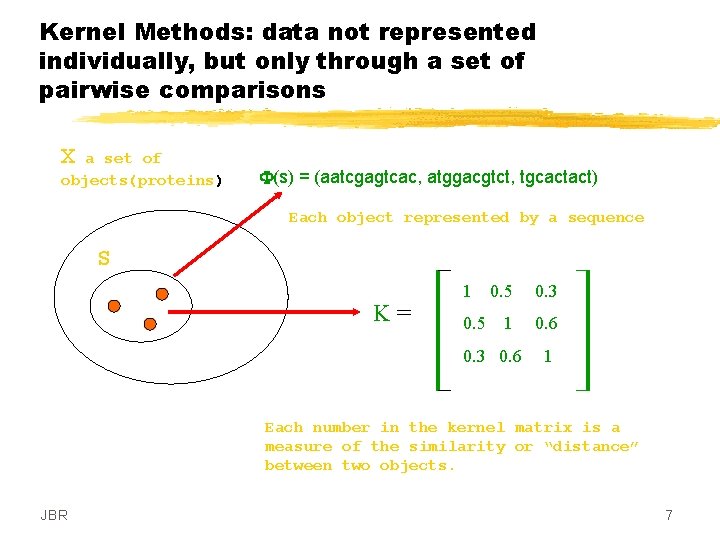
Kernel Methods: data not represented individually, but only through a set of pairwise comparisons X a set of objects(proteins) F(s) = (aatcgagtcac, atggacgtct, tgcactact) Each object represented by a sequence S K= 1 0. 5 0. 3 1 0. 6 0. 3 0. 6 1 Each number in the kernel matrix is a measure of the similarity or “distance” between two objects. JBR 7

Kernels Properties of Kernels z Kernels are measures of similarity: K(x, x’) large when x and x’ are similar z Kernels must be: y Positive definite y Symmetric z kernel K, a Hilbert Space F and a mapping F: X F K(x, x’) = <F(x), F(x’)> x, x’ X z Hence all kernels can be thought of as dot products in some feature space JBR Advantages of Kernels z Data of very different nature can be analyzed in a unified framework z No matter what the objects are, n objects are always represented by an n x n matrix z Many times, it is easier to compare objects than represent them numerically z Complete modularity between function to represent data and algorithm to analyze data 8

The “Kernel Trick” z Any algorithm for vector data that can be expressed in terms of dot products can be performed implicitly in the feature space associated with the kernel by replacing each dot product with the kernel representation z e. g. For some feature space F let: d(x, x’) = ||F(x) - F(x’)|| z But ||F(x)-F(x’)||2 = <F(x), F(x)> + <F(x’), F(x’)> - 2<F(x), F(x’)> z So d(x, x’) =(K(x, x)+K(x’, x’)-2 K(x, x’))1/2 JBR 9

Nonlinear Separation JBR X Nonlinear kernel: z X is a vector space z the kernel F is nonlinear z linear separation in the feature space F can be associated with non linear separation in X F F 10

SVM with Kernel Final formulation: z Find a that minimizes: W(a)=(-1/2) yiyjaiajxi. Txj + ai under the constraints: ai = 0 and 0 ai C z Find an index i, 0 < ai < C and set: b = yi - yjajk(xixj) z The classification of a new object x X is then determined by the sign of the function f(x) = yiaik(xix)+ b JBR 11
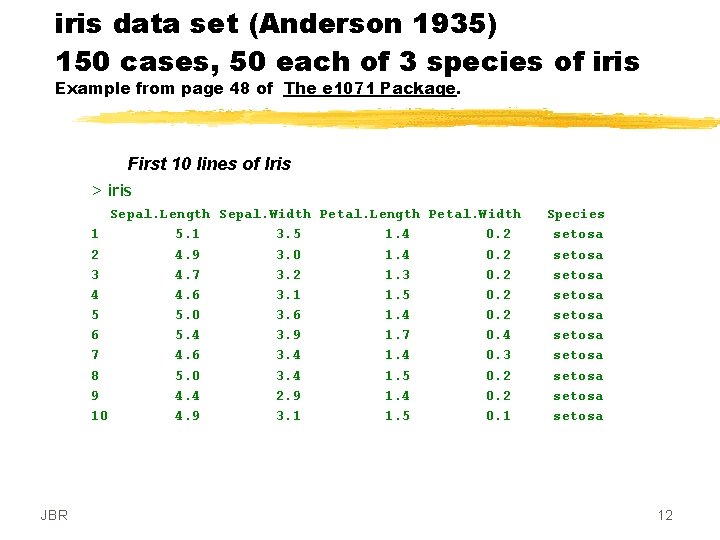
iris data set (Anderson 1935) 150 cases, 50 each of 3 species of iris Example from page 48 of The e 1071 Package. First 10 lines of Iris > iris Sepal. Length Sepal. Width Petal. Length Petal. Width 1 5. 1 3. 5 1. 4 0. 2 2 4. 9 3. 0 1. 4 0. 2 3 4. 7 3. 2 1. 3 0. 2 4 4. 6 3. 1 1. 5 0. 2 5 5. 0 3. 6 1. 4 0. 2 6 5. 4 3. 9 1. 7 0. 4 7 4. 6 3. 4 1. 4 0. 3 8 5. 0 3. 4 1. 5 0. 2 9 4. 4 2. 9 1. 4 0. 2 10 4. 9 3. 1 1. 5 0. 1 JBR Species setosa setosa setosa 12
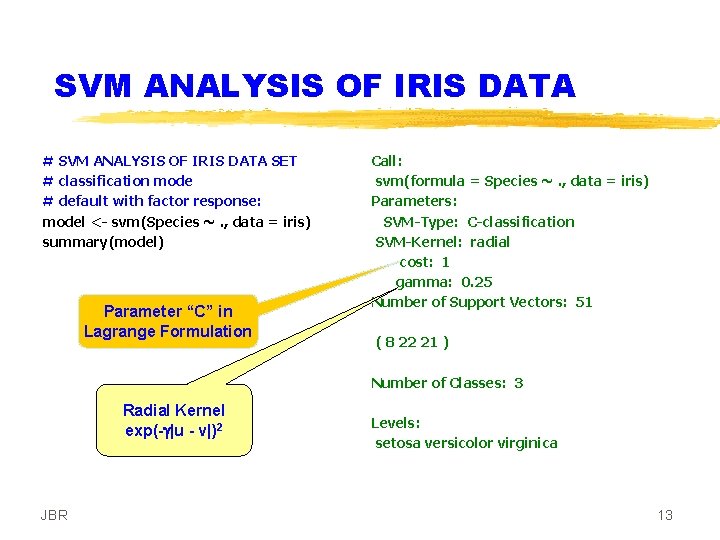
SVM ANALYSIS OF IRIS DATA # SVM ANALYSIS OF IRIS DATA SET # classification mode # default with factor response: model <- svm(Species ~. , data = iris) summary(model) Parameter “C” in Lagrange Formulation Call: svm(formula = Species ~. , data = iris) Parameters: SVM-Type: C-classification SVM-Kernel: radial cost: 1 gamma: 0. 25 Number of Support Vectors: 51 ( 8 22 21 ) Number of Classes: 3 Radial Kernel exp(-g|u - v|)2 JBR Levels: setosa versicolor virginica 13

Exploring the SVM Model # test with training data x <- subset(iris, select = -Species) y <- Species pred <- predict(model, x) # Check accuracy: table(pred, y) # compute decision values: pred <- predict(model, x, decision. values = TRUE) attr(pred, "decision. values")[1: 4, ] JBR y pred setosa versicolor virginica setosa 50 0 0 versicolor 0 48 2 virginica 0 2 48 setosa/versicolor setosa/virginica versicolor/virginica [1, ] 1. 196000 1. 091667 0. 6706543 [2, ] 1. 064868 1. 055877 0. 8482041 [3, ] 1. 181229 1. 074370 0. 6438237 [4, ] 1. 111282 1. 052820 0. 6780645 14
![Visualize classes with MDS # visualize (classes by color, SV by crosses): plot(cmdscale(dist(iris[, -5])), Visualize classes with MDS # visualize (classes by color, SV by crosses): plot(cmdscale(dist(iris[, -5])),](http://slidetodoc.com/presentation_image/28b510db9074321e0418aa101cdbdb74/image-15.jpg)
Visualize classes with MDS # visualize (classes by color, SV by crosses): plot(cmdscale(dist(iris[, -5])), col = as. integer(iris[, 5]), ch = c("o", "+")[1: 150 %in% model$index + 1]) cmdscale : multidimensional scaling or principal coordinates analysis black: sertosa red: versicolor green: virginica JBR 15

iris split into training and test sets first 25 of each case training set ## SECOND SVM ANALYSIS OF IRIS DATA SET ## classification mode # default with factor response # Train with iris. train. data model. 2 <- svm(f. S. TR ~. , data = iris. train) # output from summary(model. 2) JBR Call: svm(formula = f. S. TR ~. , data = iris. train) Parameters: SVM-Type: C-classification SVM-Kernel: radial cost: 1 gamma: 0. 25 Number of Support Vectors: 32 ( 7 13 12 ) Number of Classes: 3 Levels: setosa veriscolor virginica 16

iris test results # test with iris. test. data x. 2 <- subset(iris. test, select = f. S. TE) y. 2 <- f. S. TE pred. 2 <- predict(model. 2, x. 2) # Check accuracy: table(pred. 2, y. 2) # compute decision values and probabilities: pred. 2 <- predict(model. 2, x. 2, decision. values = TRUE) attr(pred. 2, "decision. values")[1: 4, ] JBR y. 2 pred. 2 setosa veriscolor virginica setosa 25 0 0 veriscolor 0 25 0 virginica 0 0 25 setosa/veriscolor setosa/virginica veriscolor/virginica [1, ] 1. 253378 1. 086341 0. 6065033 [2, ] 1. 000251 1. 021445 0. 8012664 [3, ] 1. 247326 1. 104700 0. 6068924 [4, ] 1. 164226 1. 078913 0. 6311566 17
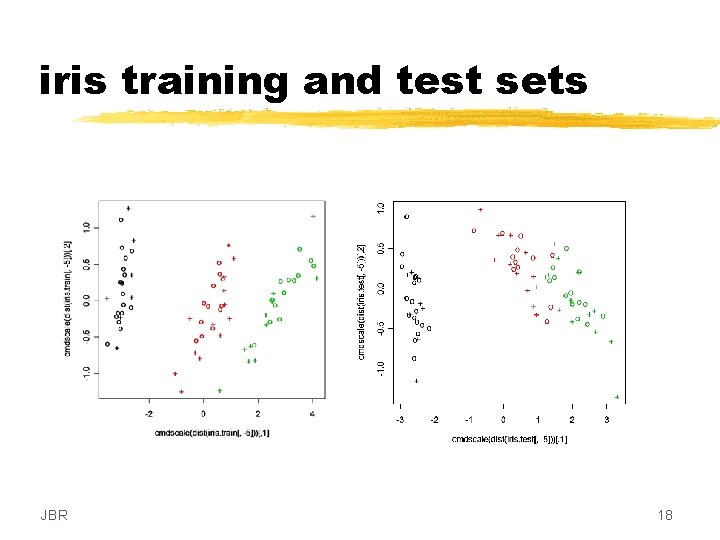
iris training and test sets JBR 18
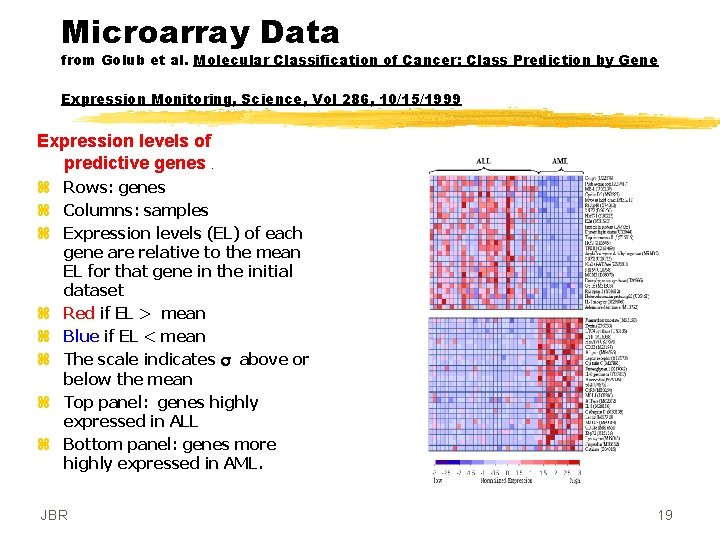
Microarray Data from Golub et al. Molecular Classification of Cancer: Class Prediction by Gene Expression Monitoring, Science, Vol 286, 10/15/1999 Expression levels of predictive genes. z Rows: genes z Columns: samples z Expression levels (EL) of each gene are relative to the mean EL for that gene in the initial dataset z Red if EL > mean z Blue if EL < mean z The scale indicates s above or below the mean z Top panel: genes highly expressed in ALL z Bottom panel: genes more highly expressed in AML. JBR 19
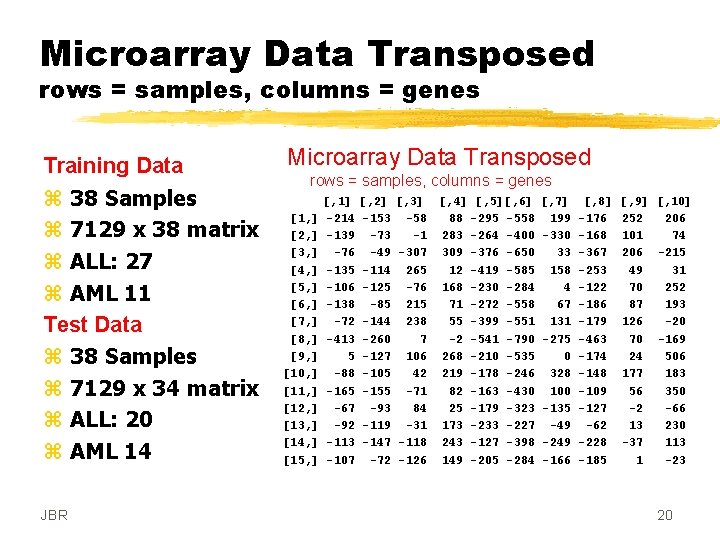
Microarray Data Transposed rows = samples, columns = genes Training Data z 38 Samples z 7129 x 38 matrix z ALL: 27 z AML 11 Test Data z 38 Samples z 7129 x 34 matrix z ALL: 20 z AML 14 JBR Microarray Data Transposed rows = samples, columns = genes [1, ] [2, ] [3, ] [4, ] [5, ] [6, ] [7, ] [8, ] [9, ] [10, ] [11, ] [12, ] [13, ] [14, ] [15, ] [, 1] -214 -139 -76 -135 -106 -138 -72 -413 5 -88 -165 -67 -92 -113 -107 [, 2] -153 -73 -49 -114 -125 -85 -144 -260 -127 -105 -155 -93 -119 -147 -72 [, 3] [, 4] [, 5][, 6] [, 7] [, 8] -58 88 -295 -558 199 -176 -1 283 -264 -400 -330 -168 -307 309 -376 -650 33 -367 265 12 -419 -585 158 -253 -76 168 -230 -284 4 -122 215 71 -272 -558 67 -186 238 55 -399 -551 131 -179 7 -2 -541 -790 -275 -463 106 268 -210 -535 0 -174 42 219 -178 -246 328 -148 -71 82 -163 -430 100 -109 84 25 -179 -323 -135 -127 -31 173 -233 -227 -49 -62 -118 243 -127 -398 -249 -228 -126 149 -205 -284 -166 -185 [, 9] [, 10] 252 206 101 74 206 -215 49 31 70 252 87 193 126 -20 70 -169 24 506 177 183 56 350 -2 -66 13 230 -37 113 1 -23 20

SVM ANALYSIS OF MICROARRAY DATA classification mode # default with factor response y <-c(rep(0, 27), rep(1, 11)) fy <-factor(y, levels=0: 1) levels(fy) <-c("ALL", "AML") # compute svm on first 3000 genes only because of memory overflow problems model. ma <- svm(fy ~. , data = fmat. train[, 1: 3000]) JBR Call: svm(formula = fy ~. , data = fmat. train[, 1: 3000]) Parameters: SVM-Type: C-classification SVM-Kernel: radial cost: 1 gamma: 0. 0003333333 Number of Support Vectors: 37 ( 26 11 ) Number of Classes: 2 Levels: ALL AML 21
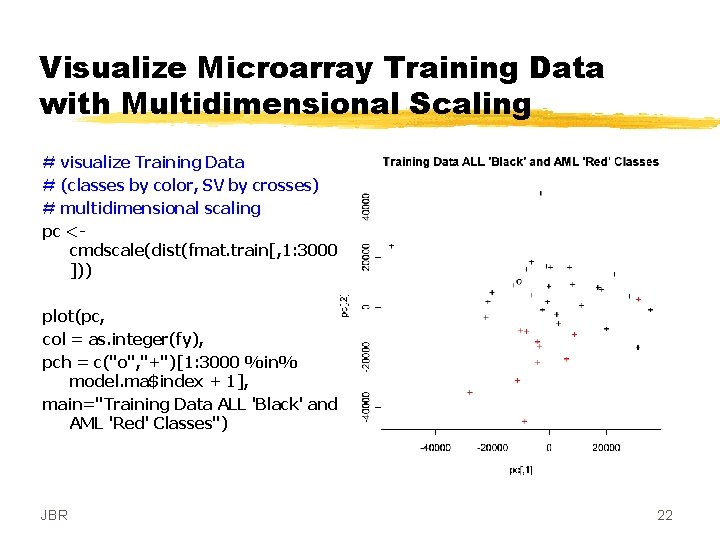
Visualize Microarray Training Data with Multidimensional Scaling # visualize Training Data # (classes by color, SV by crosses) # multidimensional scaling pc <cmdscale(dist(fmat. train[, 1: 3000 ])) plot(pc, col = as. integer(fy), pch = c("o", "+")[1: 3000 %in% model. ma$index + 1], main="Training Data ALL 'Black' and AML 'Red' Classes") JBR 22
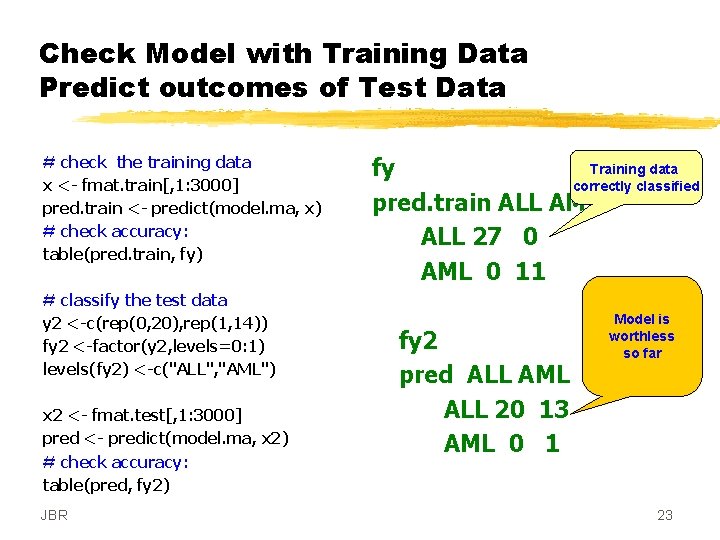
Check Model with Training Data Predict outcomes of Test Data # check the training data x <- fmat. train[, 1: 3000] pred. train <- predict(model. ma, x) # check accuracy: table(pred. train, fy) # classify the test data y 2 <-c(rep(0, 20), rep(1, 14)) fy 2 <-factor(y 2, levels=0: 1) levels(fy 2) <-c("ALL", "AML") x 2 <- fmat. test[, 1: 3000] pred <- predict(model. ma, x 2) # check accuracy: table(pred, fy 2) JBR Training data fy correctly classified pred. train ALL AML ALL 27 0 AML 0 11 fy 2 pred ALL AML ALL 20 13 AML 0 1 Model is worthless so far 23

Conclusion: z. The SVM appears to be a powerful classifier applicable to many different kinds of data But z. Kernel design is a full time job z. Selecting model parameters is far from obvious z. The math is formidable JBR 24
- Slides: 24