Supplementary table 1 Supplementary table 2 Supplementary table
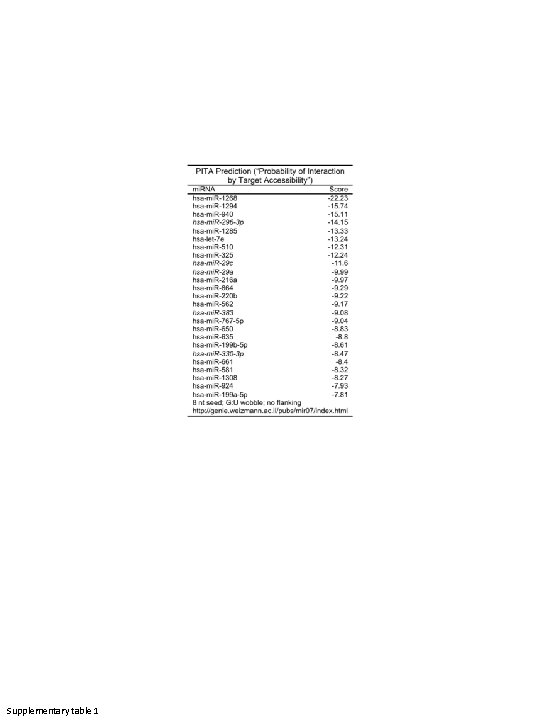
Supplementary table 1
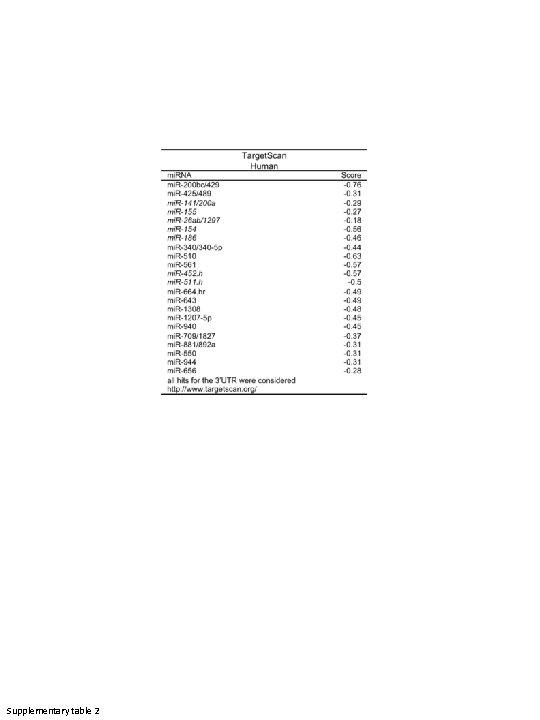
Supplementary table 2

Supplementary table 3

A. B. Supplementary table 4

VALIDATED hsa-mi. R-155 TARGETS AGTR 1 AICDA BCL 2 CSF 1 R CTLA 4 CUX 1 ETS 1 FGF 7 IKBKE IL 13 RA 1 INPP 5 D IRAK 3 JARID 2 MYD 88 RIPK 1 RNF 123 SKI SLA SOCS 1 SPI 1 WEE 1 Supplementary table 5

Supplementary table 6
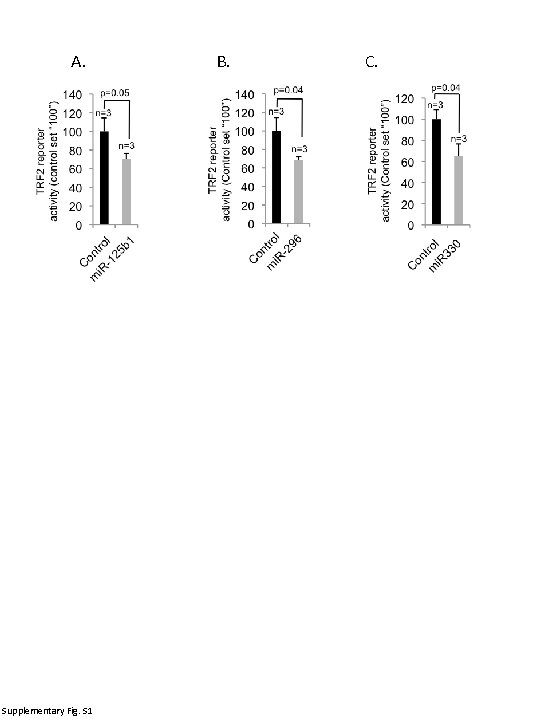
A. Supplementary Fig. S 1 B. C.
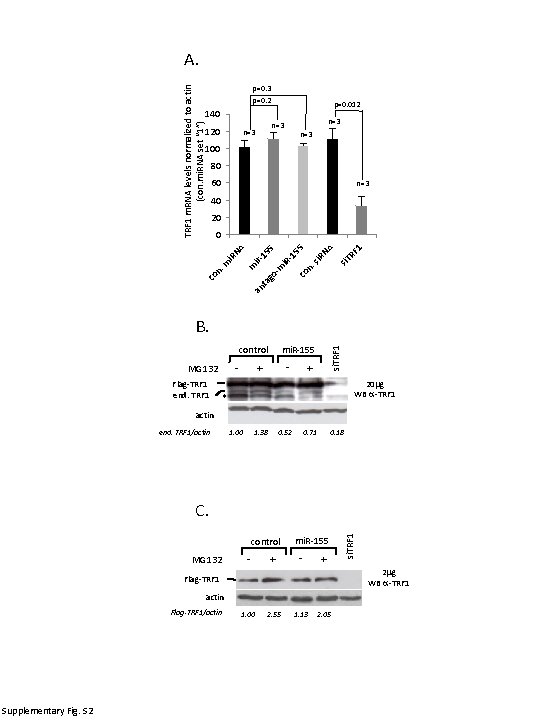
A. TRF 1 m. RNA levels normalized to actin (con. mi. RNA set “ 1”) p=0. 3 p=0. 2 140 120 p=0. 012 n=3 n=3 100 80 60 n=3 40 20 n. RF 1 co si. T -1 i. R m si. R NA 55 5 ir 15 an ta co go - n. m m i. R NA 0 B. - MG 132 Flag-TRF 1 end. TRF 1 - + si. TRF 1 mi. R-155 control + 20 mg WB a-TRF 1 * actin end. TRF 1/actin 1. 00 1. 38 0. 52 0. 71 0. 18 control MG 132 - + mi. R-155 - + 2 mg WB a-TRF 1 Flag-TRF 1 actin Flag-TRF 1/actin Supplementary Fig. S 2 si. TRF 1 C. 1. 00 2. 55 1. 13 2. 05
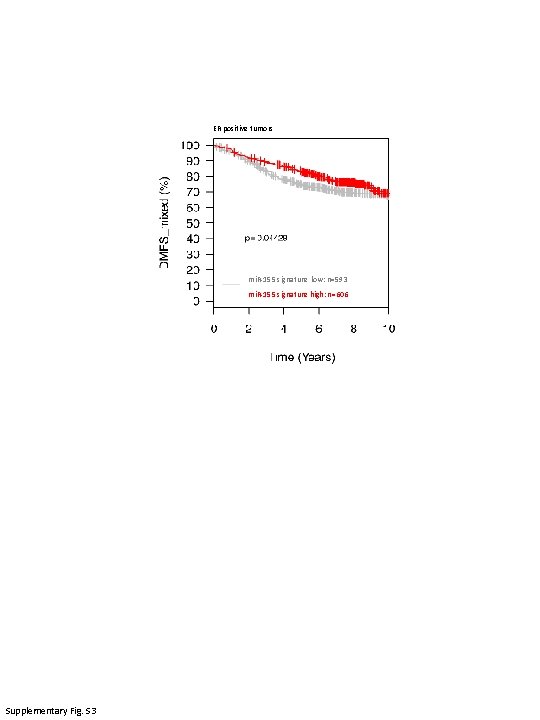
ER positive tumors mi. R-155 signature low: n=593 mi. R-155 signature high: n=606 Supplementary Fig. S 3
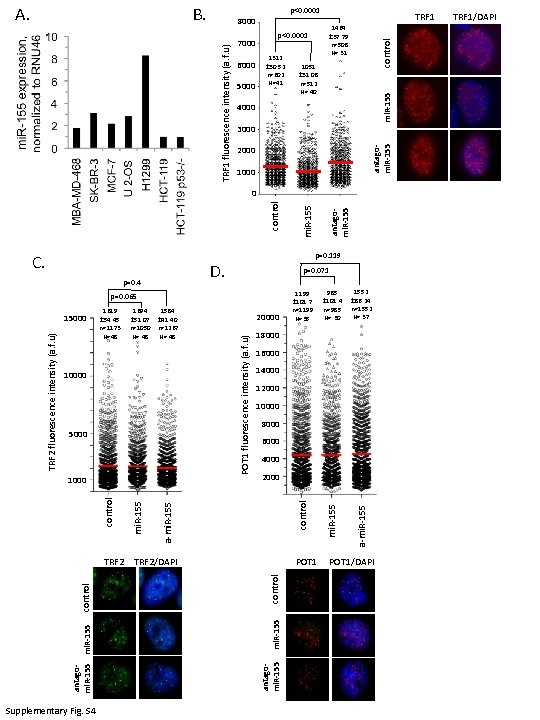
B. p<0. 0001 8000 5000 1051 ± 31. 08 n=512 N= 40 mi. R-155 1312 ± 30. 52 n=602 N=41 6000 control p<0. 0001 7000 TRF 1 1464 ± 37. 79 n=508 N= 31 4000 3000 antagomi. R-155 TRF 1 fluorescence intensity (a. f. u) U 2 -OS MBA-MD-468 A. 2000 1000 D. 1694 1584 ± 51. 07 ± 41. 40 n=1050 n=1267 N= 48 10000 5000 Supplementary Fig. S 4 mi. R-155 TRF 2/DAPI 1552 ± 86. 24 n=1552 N= 37 16000 14000 12000 10000 8000 6000 4000 2000 a-mi. R-155 control TRF 2 985 ± 101. 4 n=985 N= 30 18000 mi. R-155 control POT 1 antagomi. R-155 control 1000 20000 POT 1 fluorescence intensity (a. f. u) TRF 2 fluorescence intensity (a. f. u) 1619 ± 54. 45 n=1173 N=48 1199 ± 101. 7 n=1199 N=35 a-mi. R-155 p=0. 065 15000 antago- p=0. 071 mi. R-155 p=0. 4 a-mi. R-155 p=0. 119 control C. mi. R-155 control 0 POT 1/DAPI TRF 1/DAPI
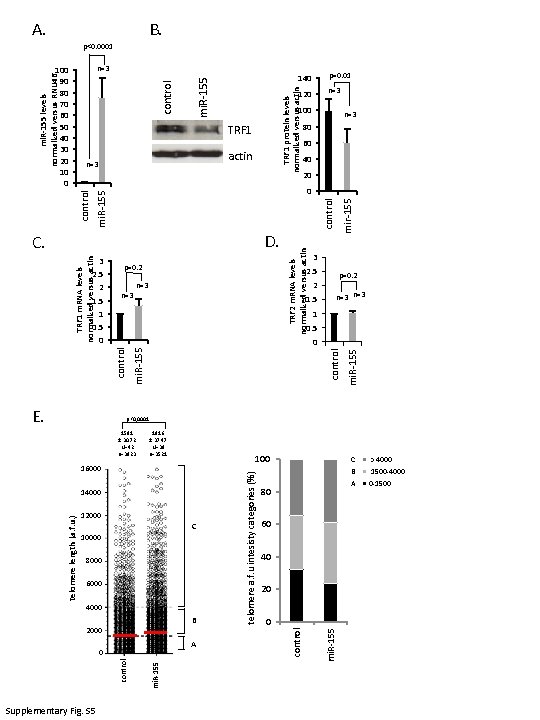
1591 ± 30. 72 N=42 n=3823 Supplementary Fig. S 5 1916 ± 37. 47 N=39 n=3521 16000 14000 12000 10000 C 8000 6000 4000 2000 B 0 A C. D. p=0. 2 n=3 2 1. 5 mir-155 actin control 100 2. 5 100 80 60 40 20 0 mi. R-155 control 120 TRF 1 protein levels normalized versus actin TRF 1 0. 5 TRF 2 m. RNA levels normalized versus actin 140 mi. R-155 E. mi. R-155 control n=3 control mi. R-155 control n=3 telomere a. f. u intesisty categories (%) 3 2. 5 2 1. 5 1 0. 5 0 mi. R-155 control TRF 1 m. RNA levels normalized versus actin 100 90 80 70 60 50 40 30 20 10 0 mi. R-155 control telomere length (a. f. u. ) mi. R-155 levels normalized versus RNU 46 A. B. p<0. 0001 p=0. 01 n=3 80 n=3 60 40 20 0 3 p=0. 2 n=3 1 0 p<0, 0001 B C > 4000 1500 -4000 A 0 -1500
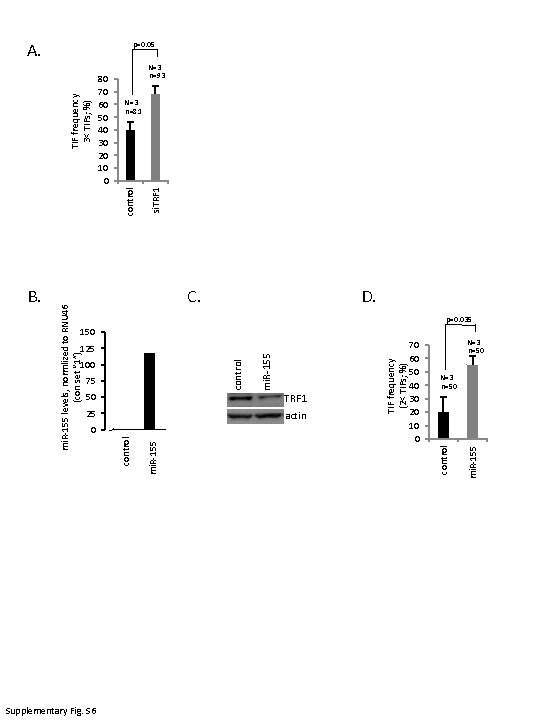
B. 150 125 100 Supplementary Fig. S 6 50 25 actin 0 mi. R-155 TIF frequency (2< TIFs; %) TRF 1 70 60 50 40 30 20 10 0 mi. R-155 75 control C. control mi. R-155 si. TRF 1 control TIF frequency 3< TIFs; %) 80 70 60 50 40 30 20 10 0 control mi. R-155 levels, normlized to RNU 46 (con set “ 1”) A. p=0. 05 N=3 n=93 N=3 n=81 D. p=0. 035 N=3 n=50
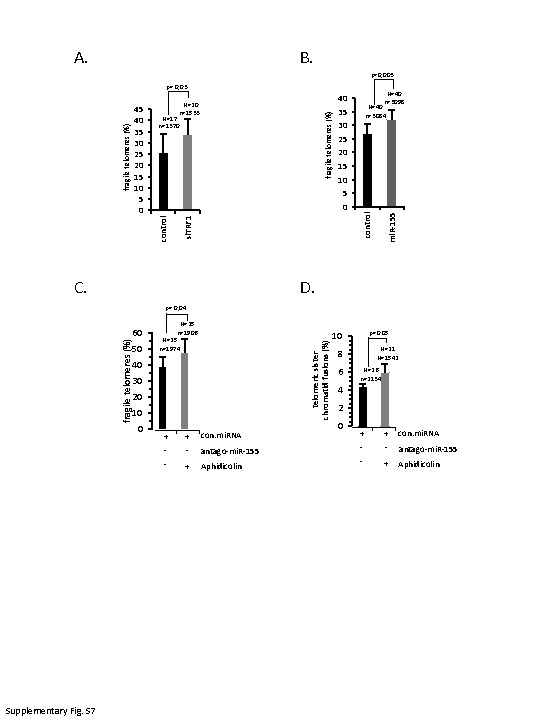
A. B. p=0, 003 p= 0, 03 fragile telomeres (%) N=17 n=1370 N=40 n=3084 35 30 25 20 15 10 5 si. TRF 1 control mi. R-155 0 control fragile telomeres (%) 45 40 35 30 25 20 15 10 5 0 N=40 n=3098 40 N=20 n=1533 C. D. p= 0, 04 50 N=25 n=1906 N=25 n=1974 telomeric sister chromatid fusions (%) fragile telomeres (%) 60 40 30 20 10 Supplementary Fig. S 7 0 + + con. mi. RNA - - antago-mi. R-155 - + Aphidicolin p=0. 05 10 N=21 N=1542 8 6 N=26 n=2134 4 2 0 + + con. mi. RNA - - antago-mi. R-155 - + Aphidicolin
- Slides: 13