Supplementary Figure Legends Table 1 List of primers

Supplementary Figure Legends: Table 1. List of primers used for Real Time – PCR Supp Fig 1. S 100 A 7 -overexpressing MCF 7 cells show decreased formation of migratory structures compared to vector control. Phase contrast image of cultured MCF 7/Vec and MCF 7/S 100 A 7 cells taken at 10 x magnification. Inset image shows the representative zoomed area. Supp Fig 2. S 100 A 7 -overexpression in MCF 7 cells down-regulate genes which are involved in cancer pathways. Total RNA from MCF 7/Vec and MCF 7/S 100 A 7 cells were analyzed by Affymetrix Human Genome U 133 chip. (A) Gene ontology studies using IPA analysis of microarray data. (B) Heat-map of differentially expressed genes in MCF 7/Vec and MCF 7/S 100 A 7 cells showing reduced expression of β-catenin/TCF 4 pathway genes. Supp Fig 3. Expression of estrogen receptor alpha (ERα) in S 100 A 7 -overexpressing and vector control MCF 7 and T 47 D cells. 50 μg cell lysates of MCF 7/Vec, MCF 7/S 100 A 7, T 47 D/Vec and T 47 D/S 100 A 7 cells were subjected to Western blotting with ERα antibody. GAPDH was used as a loading control in all the blots.
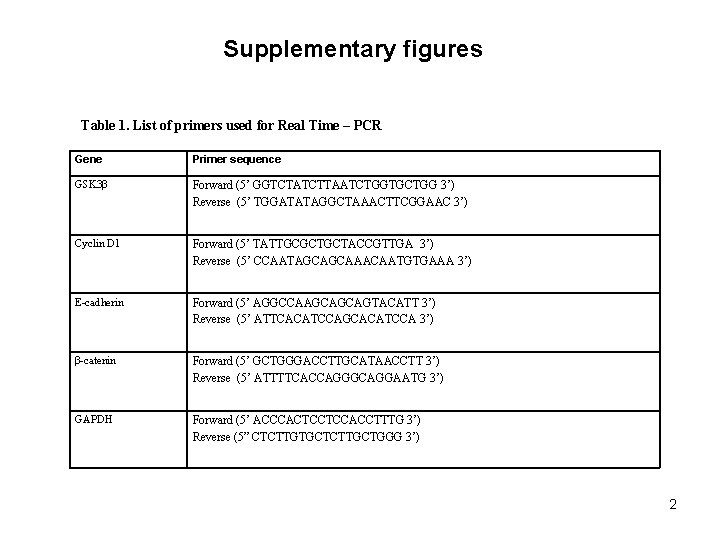
Supplementary figures Table 1. List of primers used for Real Time – PCR Gene Primer sequence GSK 3β Forward (5’ GGTCTATCTTAATCTGGTGCTGG 3’) Reverse (5’ TGGATATAGGCTAAACTTCGGAAC 3’) Cyclin D 1 Forward (5’ TATTGCGCTGCTACCGTTGA 3’) Reverse (5’ CCAATAGCAGCAAACAATGTGAAA 3’) E-cadherin Forward (5’ AGGCCAAGCAGCAGTACATT 3’) Reverse (5’ ATTCACATCCAGCACATCCA 3’) β-catenin Forward (5’ GCTGGGACCTTGCATAACCTT 3’) Reverse (5’ ATTTTCACCAGGGCAGGAATG 3’) GAPDH Forward (5’ ACCCACTCCTCCACCTTTG 3’) Reverse (5” CTCTTGTGCTCTTGCTGGG 3’) 2
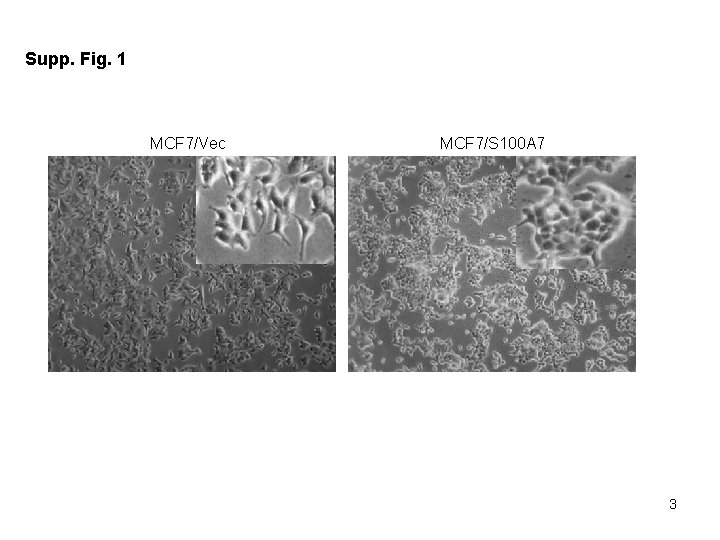
Supp. Fig. 1 MCF 7/Vec MCF 7/S 100 A 7 3
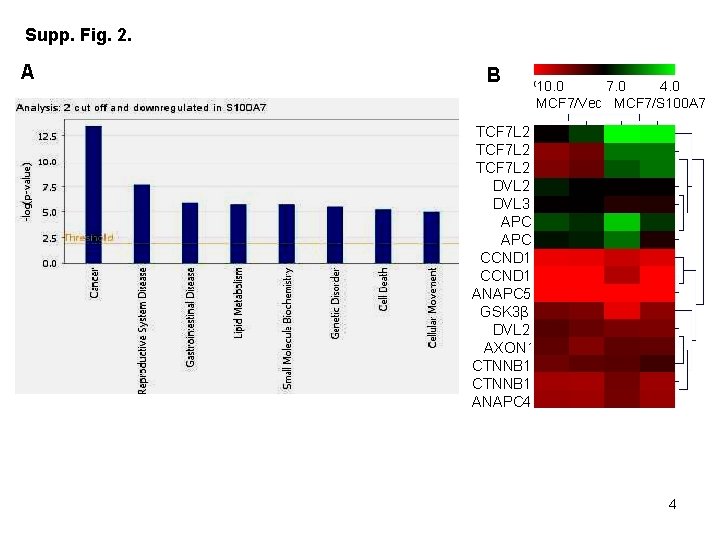
Supp. Fig. 2. A B 10. 0 7. 0 4. 0 MCF 7/Vec MCF 7/S 100 A 7 TCF 7 L 2 DVL 3 APC CCND 1 ANAPC 5 GSK 3β DVL 2 AXON 1 CTNNB 1 ANAPC 4 4
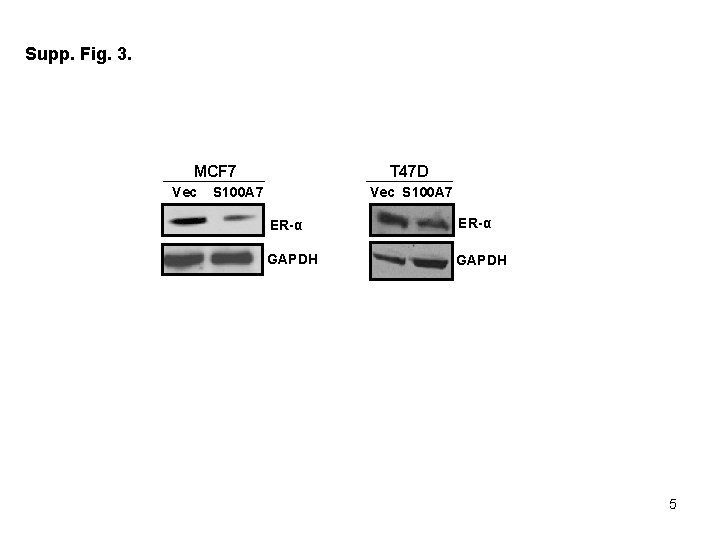
Supp. Fig. 3. MCF 7 Vec T 47 D S 100 A 7 Vec S 100 A 7 ER-α GAPDH 5
- Slides: 5