StructureFunction Relationship of Retinal Proteins Structure of Retinal

- Slides: 28

Structure-Function Relationship of Retinal Proteins

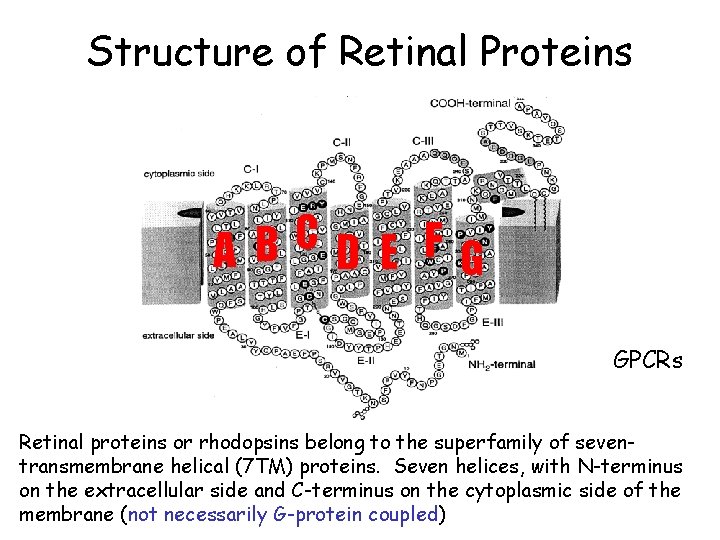
Structure of Retinal Proteins C B A D E F G GPCRs Retinal proteins or rhodopsins belong to the superfamily of seventransmembrane helical (7 TM) proteins. Seven helices, with N-terminus on the extracellular side and C-terminus on the cytoplasmic side of the membrane (not necessarily G-protein coupled)

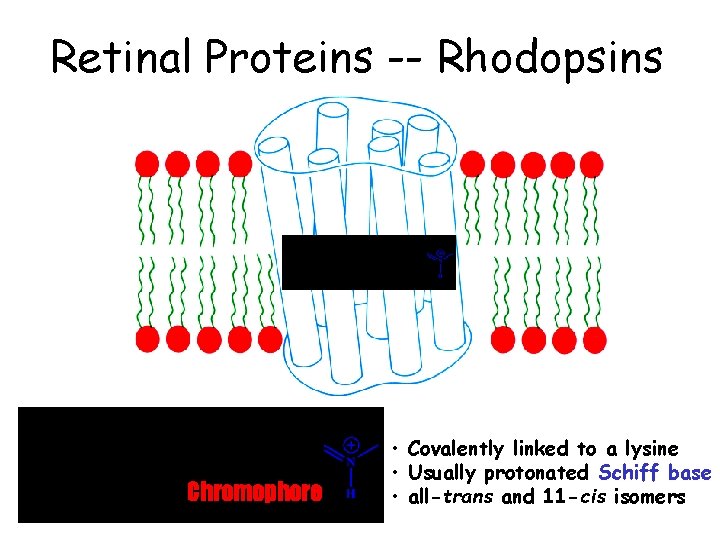
Retinal Proteins -- Rhodopsins Chromophore • Covalently linked to a lysine • Usually protonated Schiff base • all-trans and 11 -cis isomers

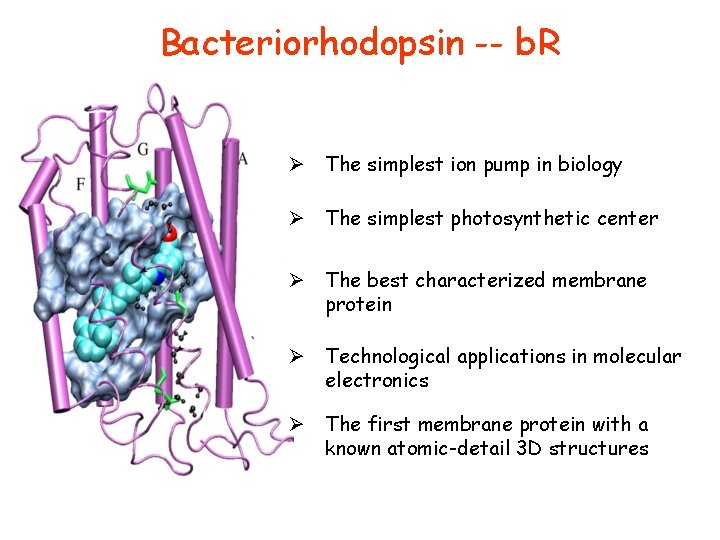
Bacteriorhodopsin -- b. R Ø The simplest ion pump in biology Ø The simplest photosynthetic center Ø The best characterized membrane protein Ø Technological applications in molecular electronics Ø The first membrane protein with a known atomic-detail 3 D structures

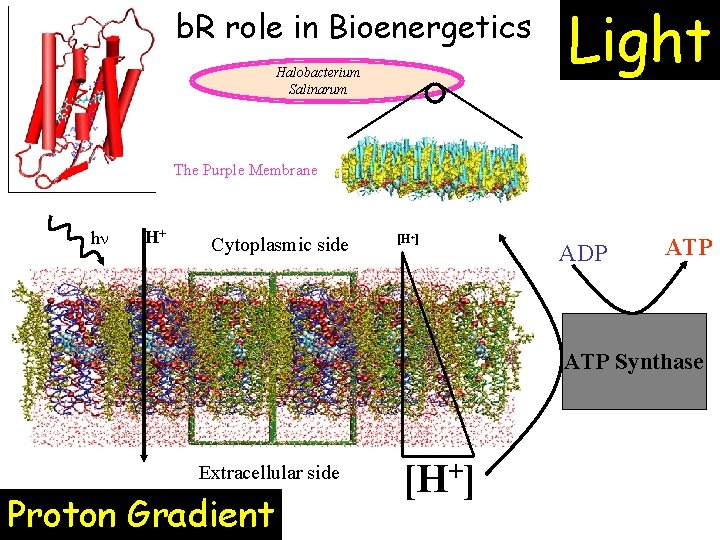
b. R role in Bioenergetics Halobacterium Salinarum Light The Purple Membrane hn H+ Cytoplasmic side [H+] ADP ATP Synthase Extracellular side Proton Gradient [H+]

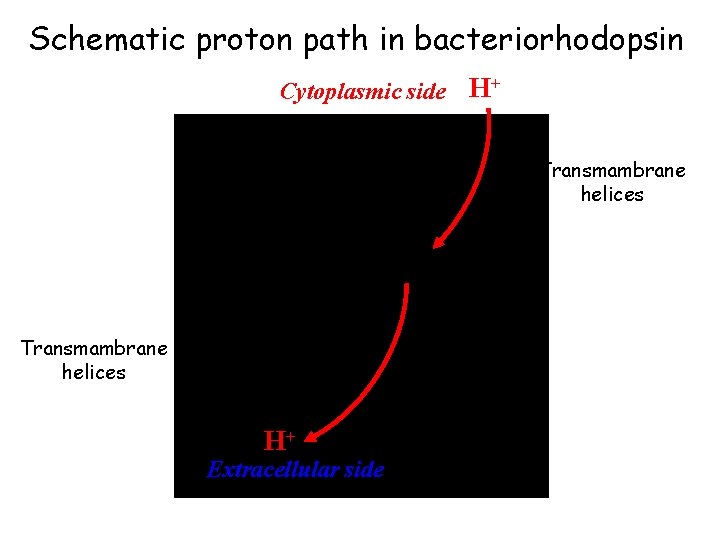
Schematic proton path in bacteriorhodopsin Cytoplasmic side H+ Transmambrane helices H+ Extracellular side

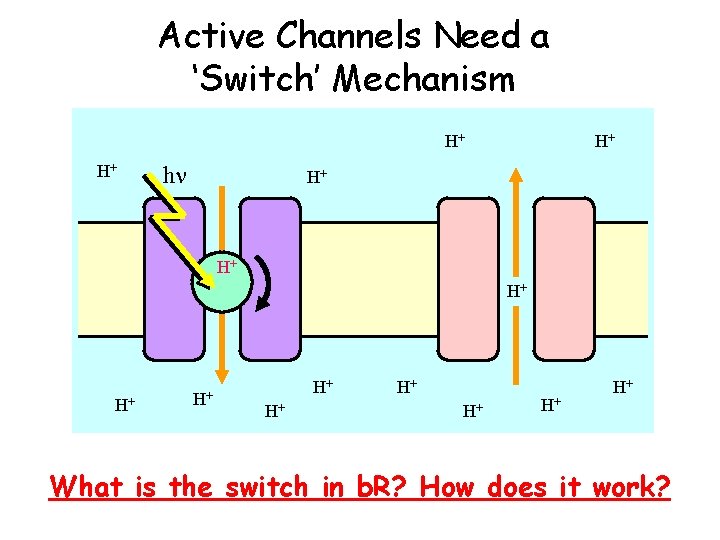
Active Channels Need a ‘Switch’ Mechanism H+ H+ hn H+ H+ H+ What is the switch in b. R? How does it work?

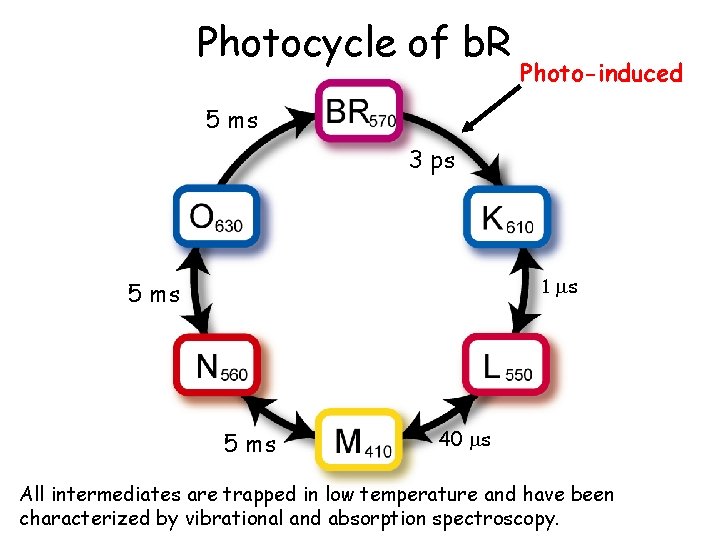
Photocycle of b. R Photo-induced 5 ms 3 ps 1 ms 5 ms 40 ms All intermediates are trapped in low temperature and have been characterized by vibrational and absorption spectroscopy.

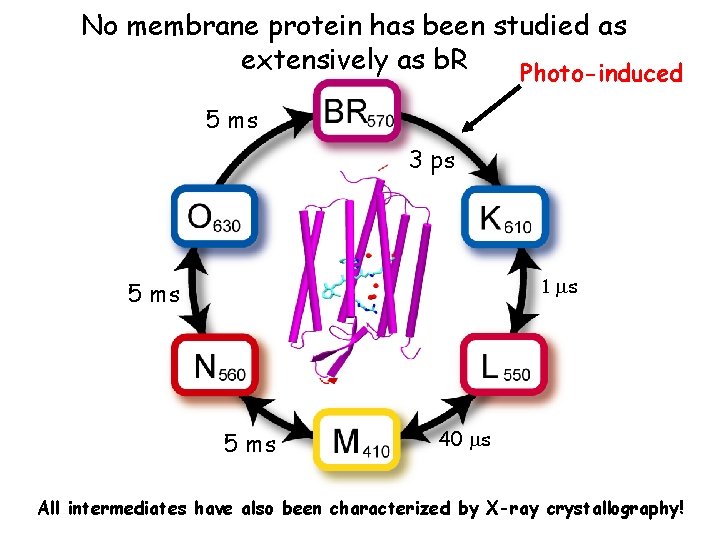
No membrane protein has been studied as extensively as b. R Photo-induced 5 ms 3 ps 1 ms 5 ms 40 ms All intermediates have also been characterized by X-ray crystallography!

Schematic proton path in bacteriorhodopsin Cytoplasmic side Transmambrane helices Extracellular side

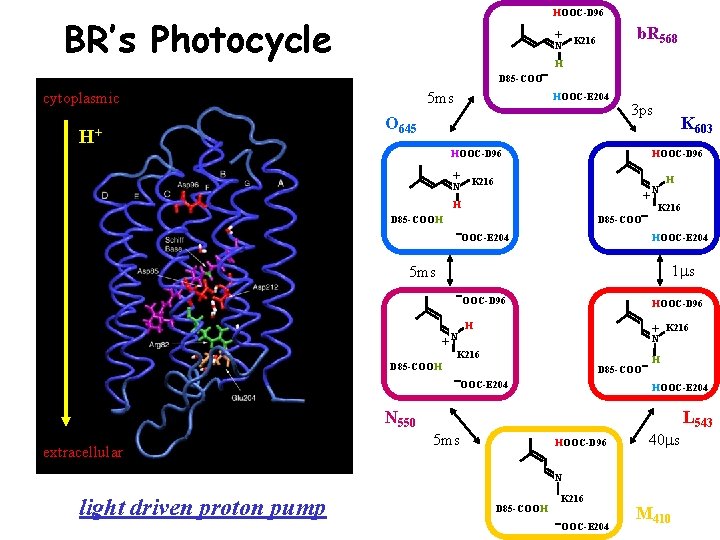
HOOC-D 96 BR’s Photocycle b. R 568 + K 216 N H D 85 -COO cytoplasmic H+ 5 ms HOOC-E 204 O 645 3 ps K 603 HOOC-D 96 + K 216 N N + H H K 216 D 85 -COOH D 85 -COO OOC-E 204 HOOC-E 204 1 ms 5 ms + N OOC-D 96 H + K 216 N K 216 D 85 -COOH D 85 -COO OOC-E 204 H HOOC-E 204 N 550 extracellular L 543 5 ms HOOC-D 96 40 ms N light driven proton pump D 85 -COOH K 216 OOC-E 204 M 410

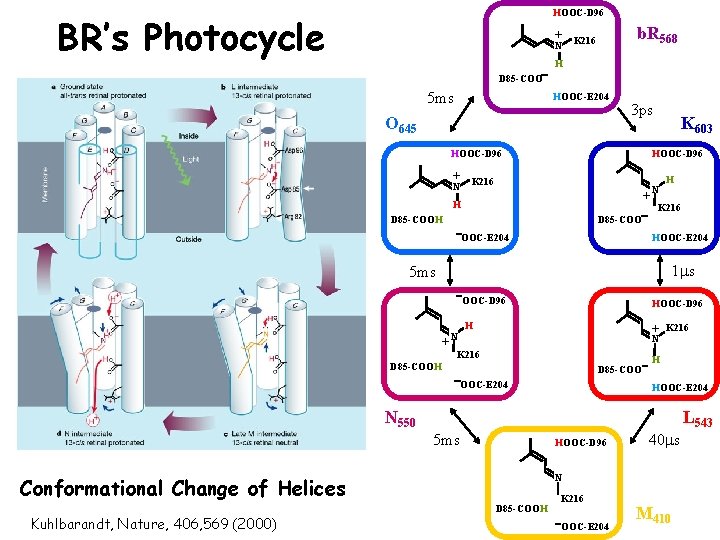
HOOC-D 96 BR’s Photocycle b. R 568 + K 216 N H D 85 -COO 5 ms HOOC-E 204 O 645 3 ps K 603 HOOC-D 96 + K 216 N N + H H K 216 D 85 -COOH D 85 -COO OOC-E 204 HOOC-E 204 1 ms 5 ms + N OOC-D 96 H + K 216 N K 216 D 85 -COOH D 85 -COO OOC-E 204 H HOOC-E 204 N 550 L 543 5 ms HOOC-D 96 N Conformational Change of Helices Kuhlbarandt, Nature, 406, 569 (2000) 40 ms D 85 -COOH K 216 OOC-E 204 M 410

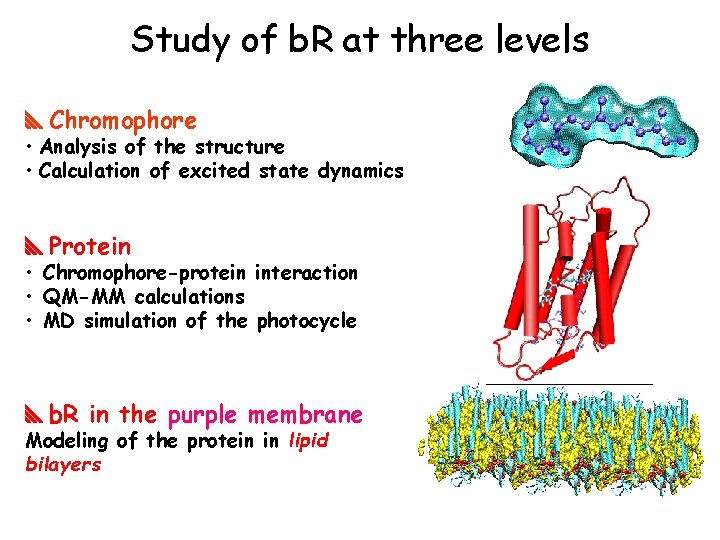
Study of b. R at three levels Chromophore • Analysis of the structure • Calculation of excited state dynamics Protein • Chromophore-protein interaction • QM-MM calculations • MD simulation of the photocycle b. R in the purple membrane Modeling of the protein in lipid bilayers

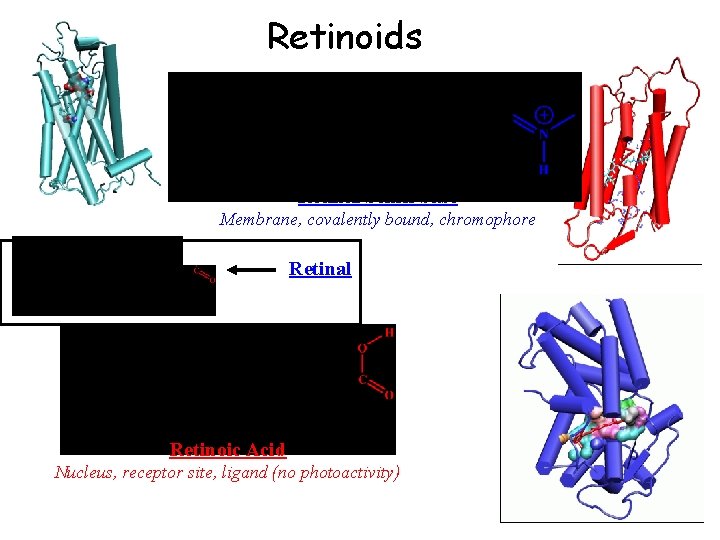
Retinoids Retinal Schiff base Membrane, covalently bound, chromophore Retinal Retinoic Acid Nucleus, receptor site, ligand (no photoactivity)

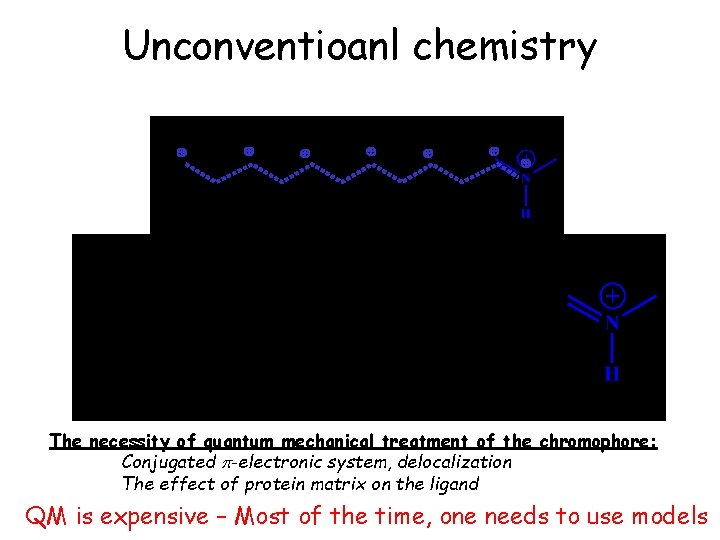
Unconventioanl chemistry 7 9 11 13 15 The necessity of quantum mechanical treatment of the chromophore: Conjugated p-electronic system, delocalization The effect of protein matrix on the ligand QM is expensive – Most of the time, one needs to use models

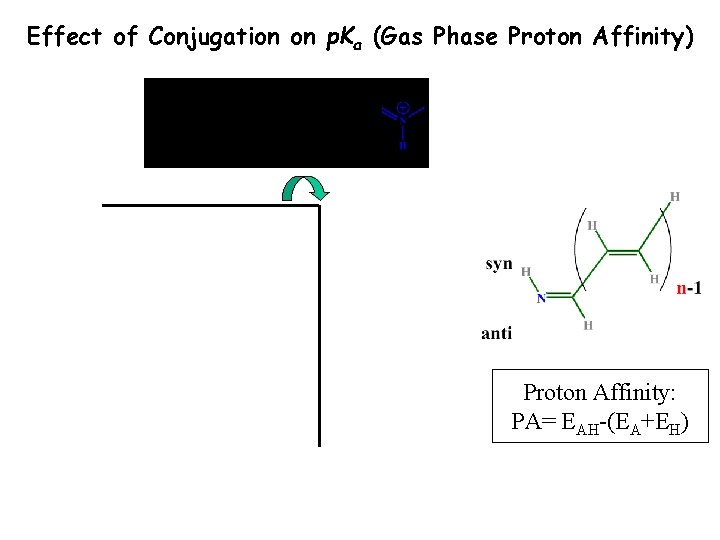
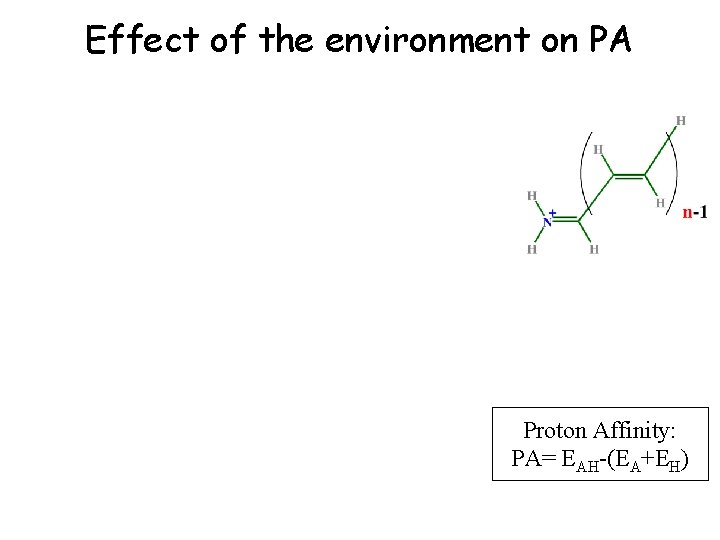
Effect of Conjugation on p. Ka (Gas Phase Proton Affinity) Proton Affinity: PA= EAH-(EA+EH)

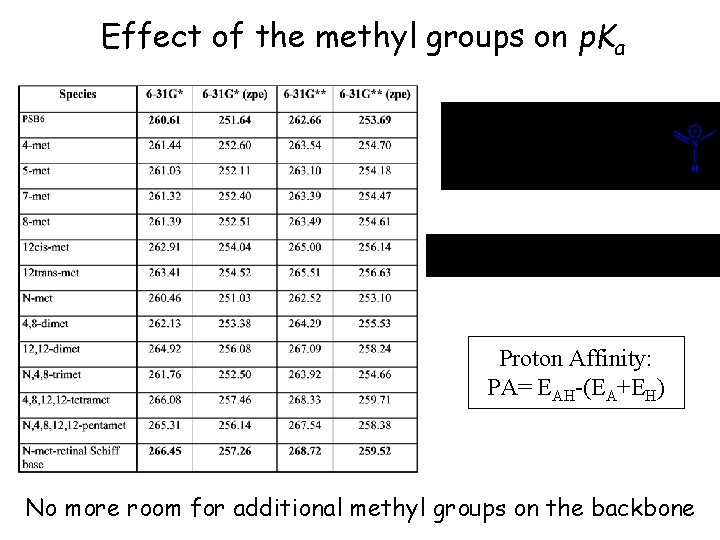
Effect of the methyl groups on p. Ka Proton Affinity: PA= EAH-(EA+EH) No more room for additional methyl groups on the backbone

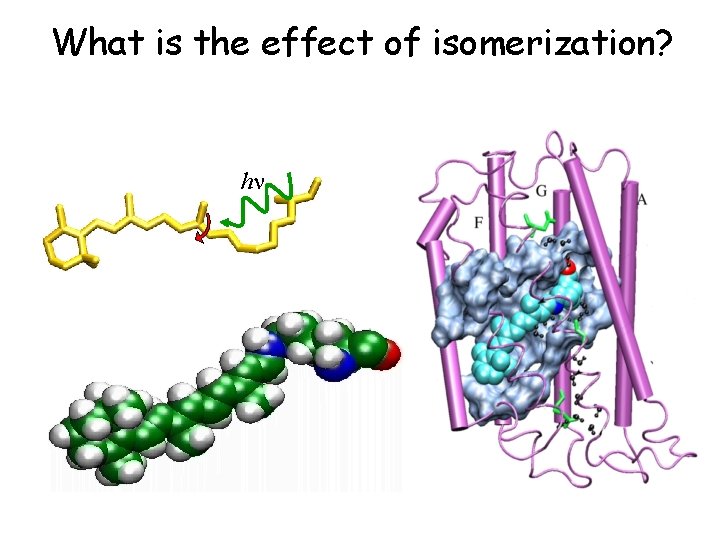
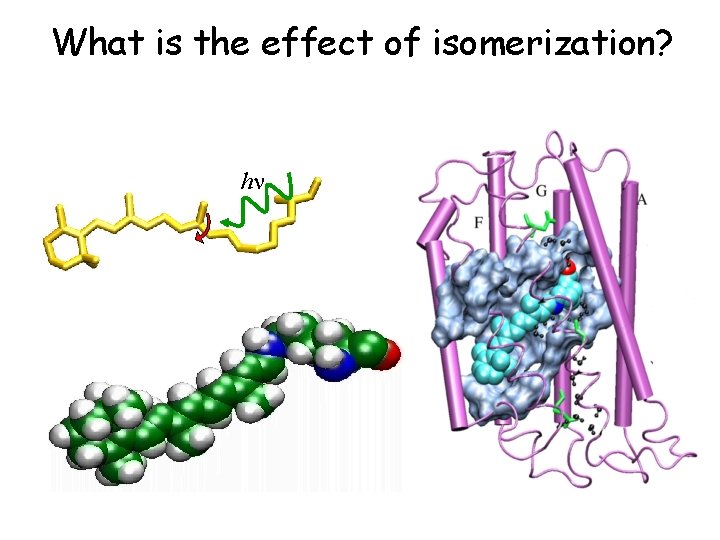
What is the effect of isomerization? hn

Isomerization State and Proton Affinity: cc: B 2, B 3 -di-cis isomer tc: B 2 -strans, B 3 -cis isomer ct: B 2 -s-cis, B 3 -trans isomer tt: all-trans isomer. PA= EAH-(EA+EH) Isomerization does not have a strong impact on PA!

What is the effect of isomerization? hn

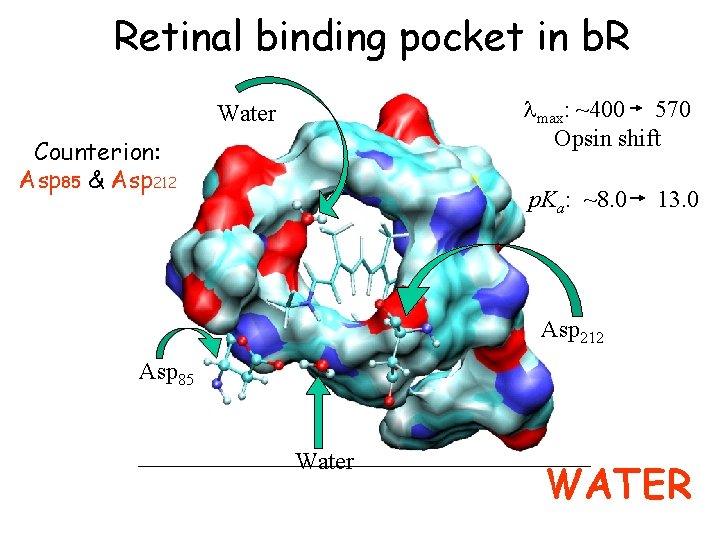
Retinal binding pocket in b. R lmax: ~400 570 Opsin shift Water Counterion: Asp 85 & Asp 212 p. Ka: ~8. 0 13. 0 Asp 212 Asp 85 Water WATER

Effect of the environment on PA Proton Affinity: PA= EAH-(EA+EH)

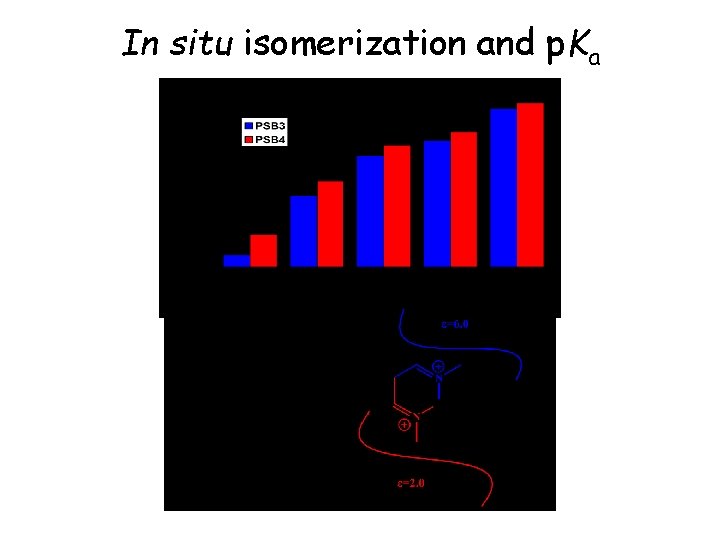
In situ isomerization and p. Ka

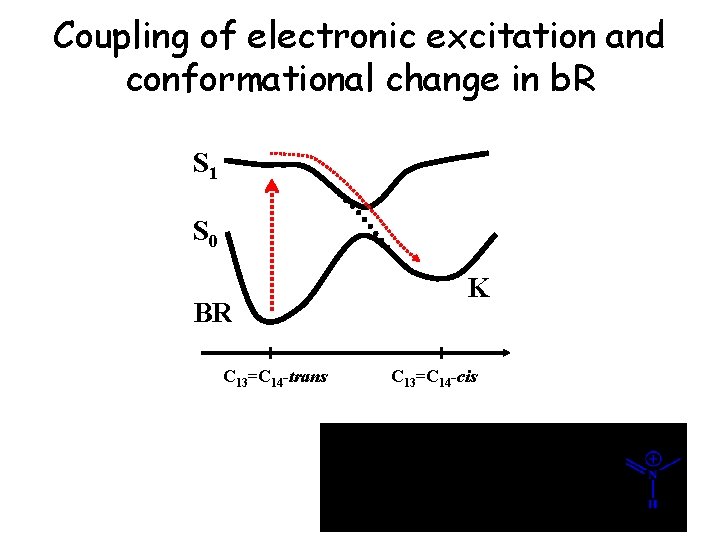
Coupling of electronic excitation and conformational change in b. R S 1 S 0 K BR C 13=C 14 -trans C 13=C 14 -cis 13 7 9 11 15

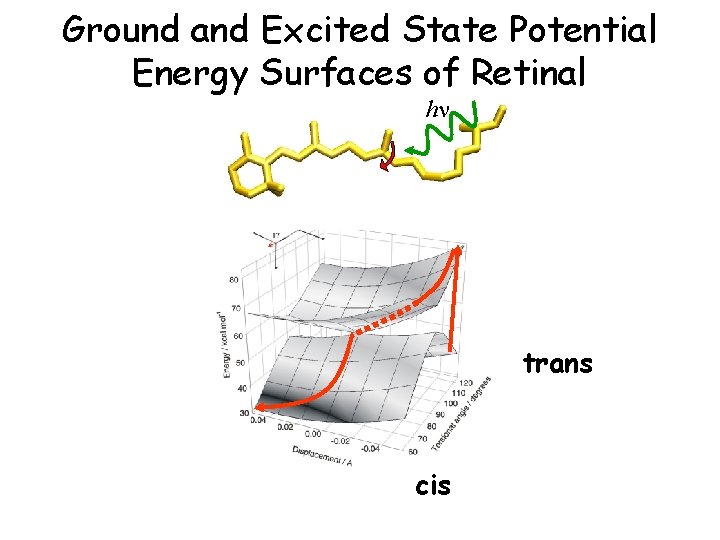
Ground and Excited State Potential Energy Surfaces of Retinal hn trans cis

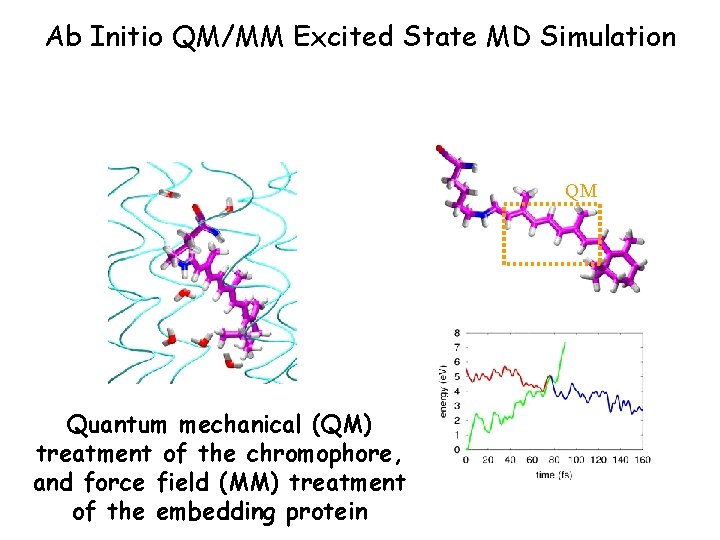
Ab Initio QM/MM Excited State MD Simulation QM Quantum mechanical (QM) treatment of the chromophore, and force field (MM) treatment of the embedding protein

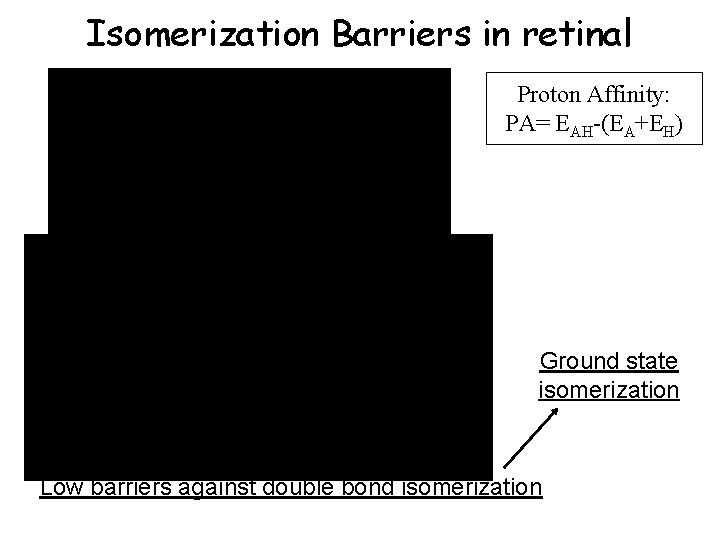
Isomerization Barriers in retinal Proton Affinity: PA= EAH-(EA+EH) Ground state isomerization Low barriers against double bond isomerization

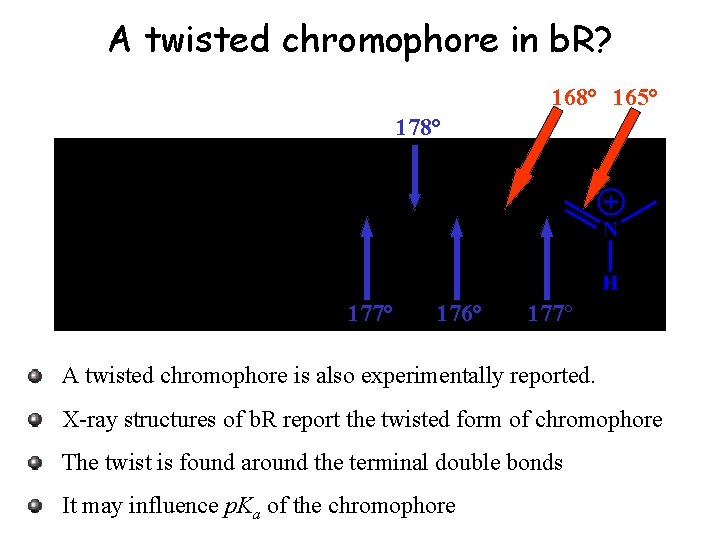
A twisted chromophore in b. R? 168° 165° 178° 177° 176° 177° A twisted chromophore is also experimentally reported. X-ray structures of b. R report the twisted form of chromophore The twist is found around the terminal double bonds It may influence p. Ka of the chromophore