Structure and Function of Proteins Ora SchuelerFurman 2009
















- Slides: 34

Structure and Function of Proteins Ora Schueler-Furman 2009 -2010 1

How to open an EKMD account ? Open this website: http: //agcc. ekmd. huji. ac. il/ekmd. asp You’ll need Ø The 1 st 8 digits of your ID number Ø 4 digit code Follow the instructions…

Website Information http: //courses. md. huji. ac. il/81817 https: //eduportal. ekmd. huji. ac. il/courses/81817 Contact us: – Ora Schueler-Furman 87094 (02 -6757094) E-mail: oraf@ekmd. huji. ac. il – Sivan Pearl 85471 E-mail: sivan. pearl@mail. huji. ac. il - Miriam Oxsman 88337 E-mail: miriam. oxsman@mail. huji. ac. il 3

PROTEINS From the Greek word proteio “Proteios” - first rank, most important Play central roles in all biological processes 4

Introduction into Protein Structure • The chemical nature of polypeptides • Forces that determine protein structure 5

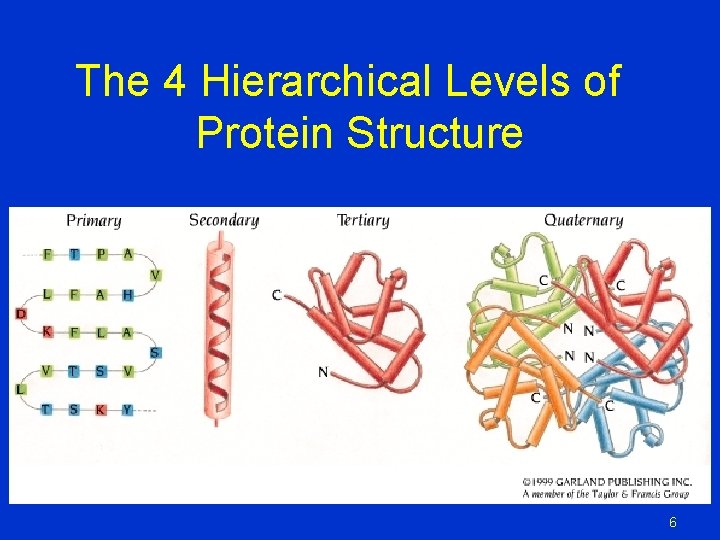
The 4 Hierarchical Levels of Protein Structure 6

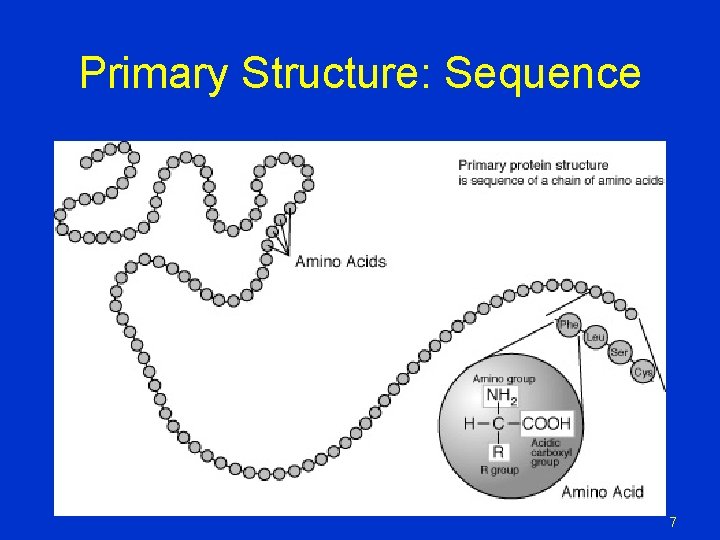
Primary Structure: Sequence 7

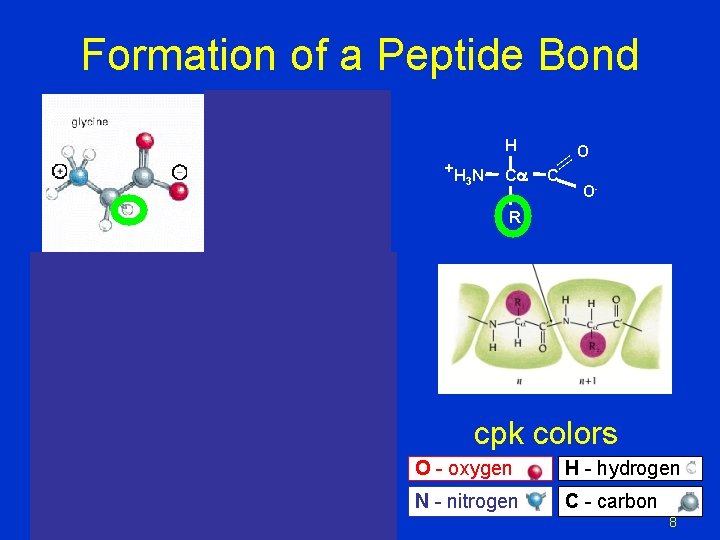
Formation of a Peptide Bond H Ca || +H N 3 O C O- R cpk colors O - oxygen H - hydrogen N - nitrogen C - carbon 8

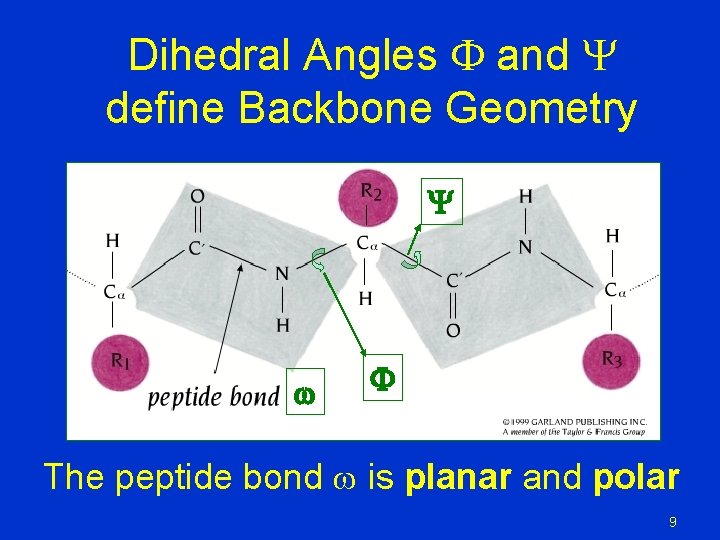
Dihedral Angles F and define Backbone Geometry w F The peptide bond w is planar and polar 9

Basic Facts Polypeptide chain: But also >20000 aa: 50 -10’ 000 aa 34’ 350 aa (Titin) Average MW of aa : 110 daltons Average MW of a protein of length n aa: n X 110 Distance (Cai – Cai+1) <=3. 63Ǻ aa – amino acid 10

Basic Terms Main chain, backbone, side chain, residue bb sc Peptide: a small # of connected aa Polypeptide: a longer chain of aa Protein: polypeptide chain with defined aa sequence & conformation 11

The Protein Alphabet: 20 letters A C D E F G H I K L Ala Cys Asp Glu Phe Gly His Ile Lys Leu Alanine Cysteine Aspartate Glutamate Phenylalanine Glycine Histidine Isoleucine Lysine Leucine M N P Q R S T V W Y Met Asn Pro Gln Arg Ser Thr Val Trp Tyr Methionine Aspargine Proline Glutamine Arginine Serine Threonine Valine Tryptophane Tyrosine amino acids vary in: volume, shape, chemical nature (charge, hydrogen bonding capability, etc. ) 12

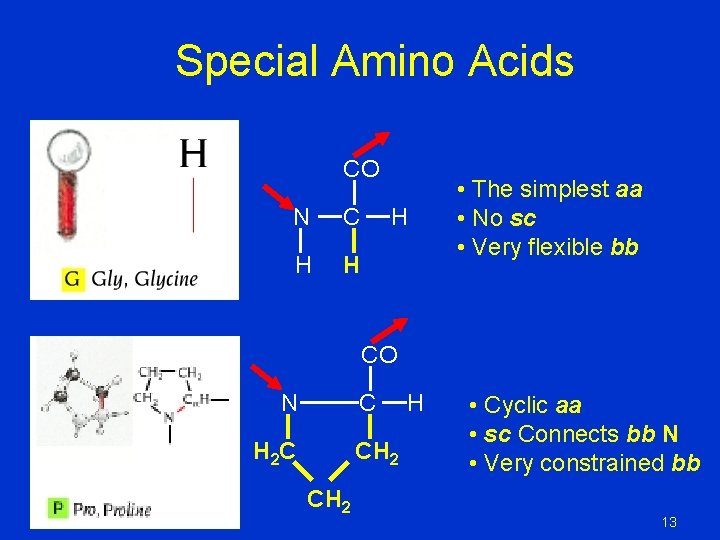
Special Amino Acids CO N C H H H • The simplest aa • No sc • Very flexible bb CO N C H 2 C CH 2 H • Cyclic aa • sc Connects bb N • Very constrained bb 13

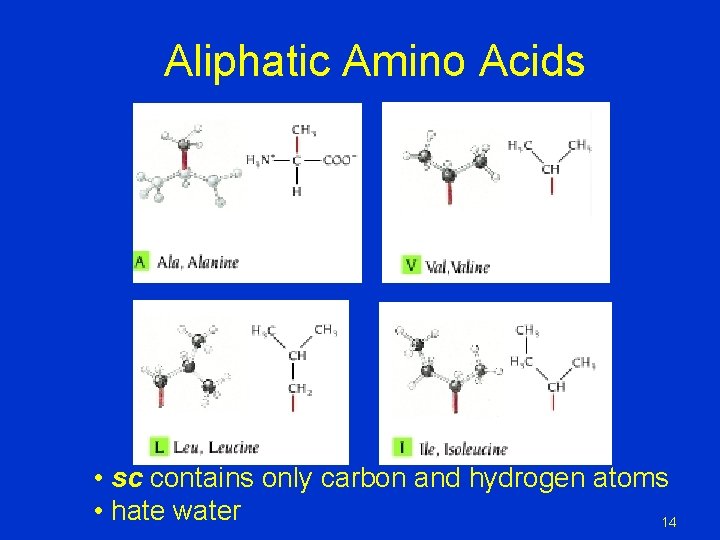
Aliphatic Amino Acids • sc contains only carbon and hydrogen atoms • hate water 14

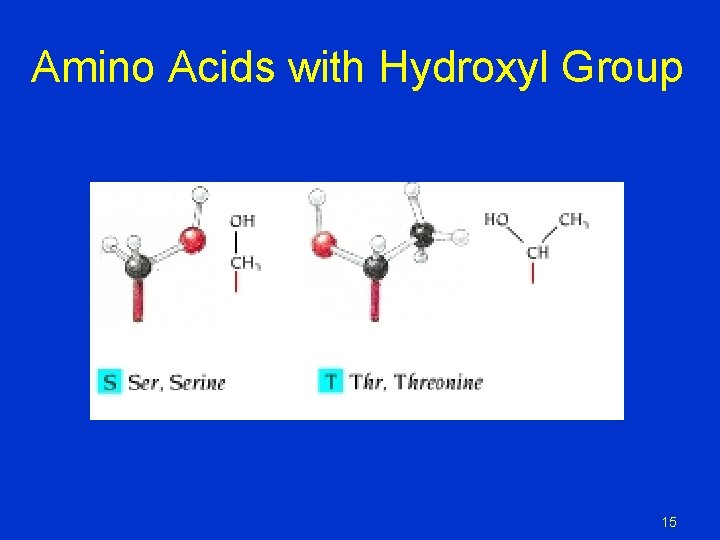
Amino Acids with Hydroxyl Group 15

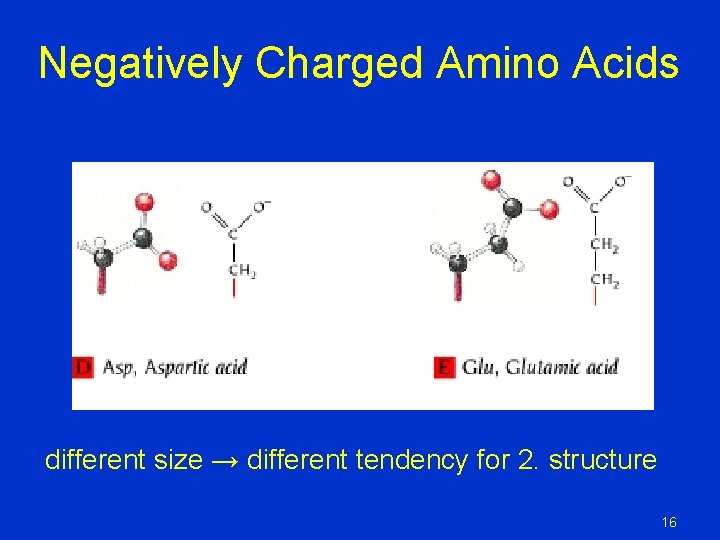
Negatively Charged Amino Acids different size → different tendency for 2. structure 16

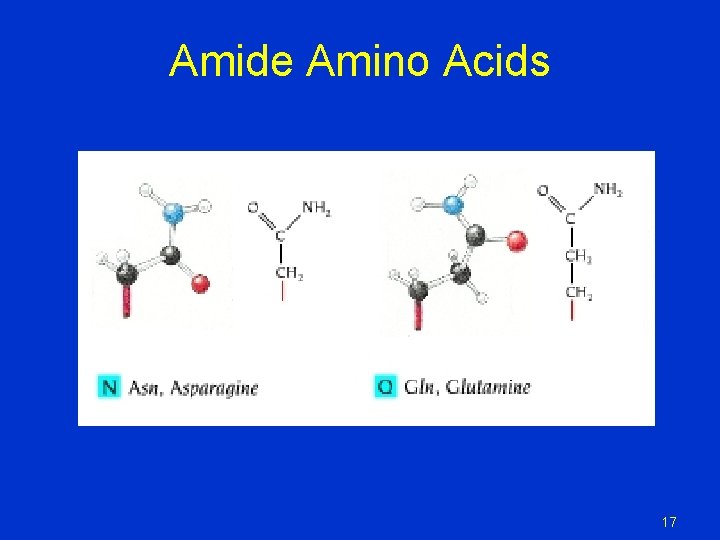
Amide Amino Acids 17

Positively Charged Amino Acids • p. Ka 11. 1 • p. Ka 12 • large sc 18

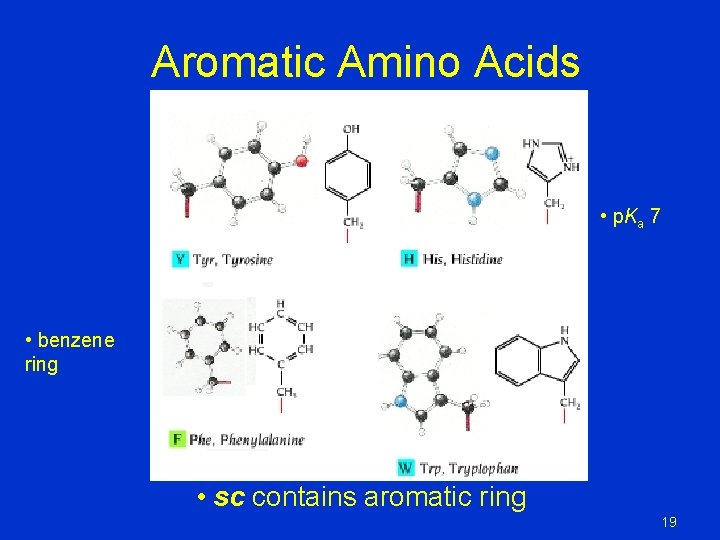
Aromatic Amino Acids • p. Ka 7 • benzene ring • sc contains aromatic ring 19

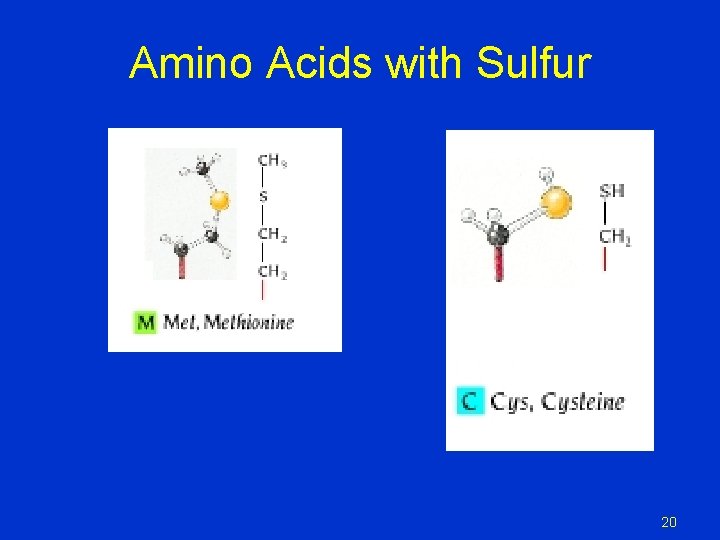
Amino Acids with Sulfur 20

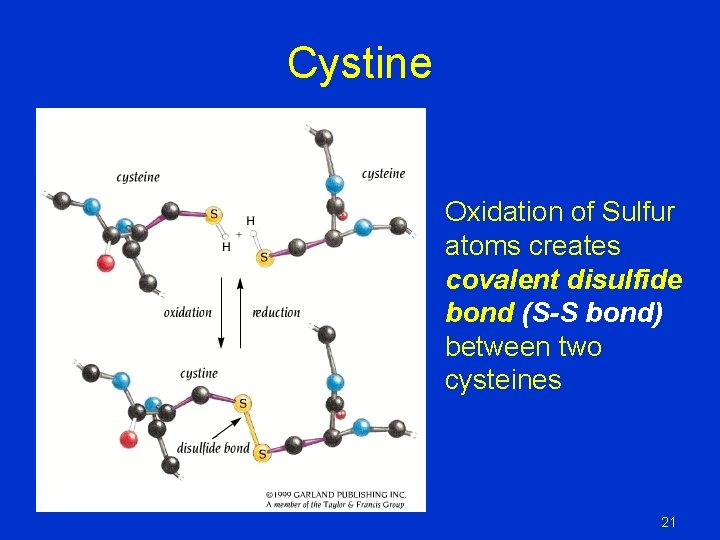
Cystine Oxidation of Sulfur atoms creates covalent disulfide bond (S-S bond) between two cysteines 21

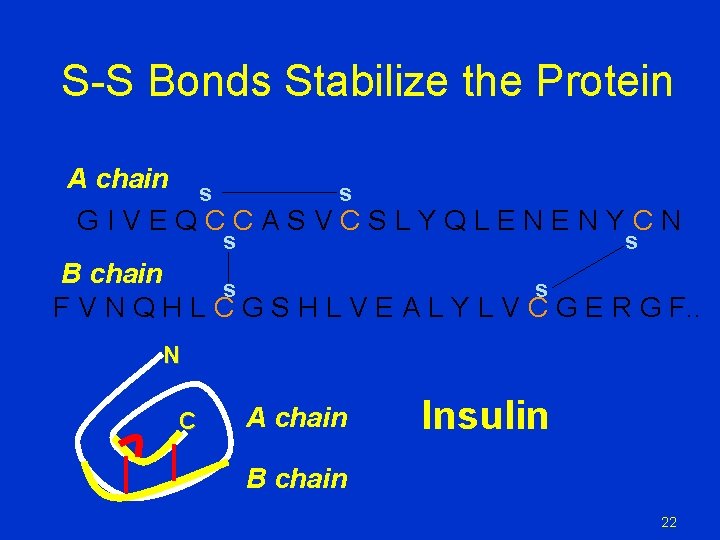
S-S Bonds Stabilize the Protein A chain s s GIVEQCCASVCSLYQLENENYCN s s B chain s s F V N Q H L C G S H L V E A L Y L V C G E R G F. . N C A chain Insulin B chain 22

Post-Translational Modifications • Processing (proinsulin/insulin) – control of protein activity • Glycosylation – protein trafficking • Phosphorylation (Tyr, Ser, Thr) – regulation of signaling • Methylation, Acetylation – histone tagging 23

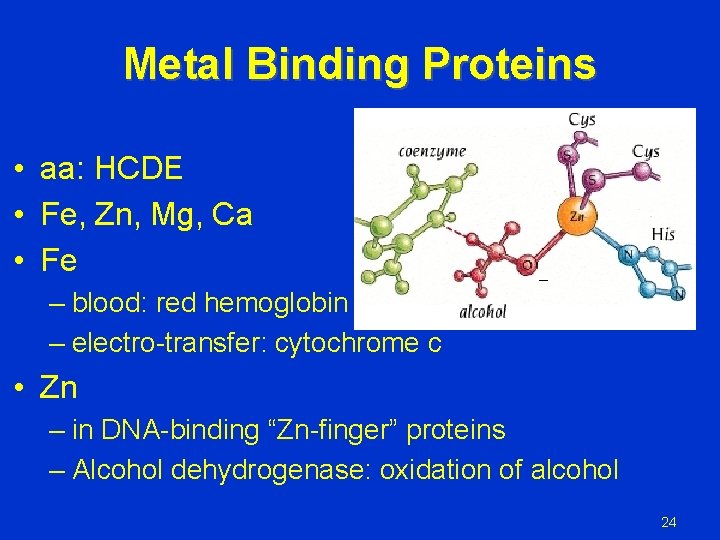
Metal Binding Proteins • aa: HCDE • Fe, Zn, Mg, Ca • Fe – blood: red hemoglobin – electro-transfer: cytochrome c • Zn – in DNA-binding “Zn-finger” proteins – Alcohol dehydrogenase: oxidation of alcohol 24

Forces that Determine Protein Structure 25

Non-Covalent Forces Add up Each bond is weak Large number of bonds From: the Molecular Biology of the Cell, 4 th ed. 26

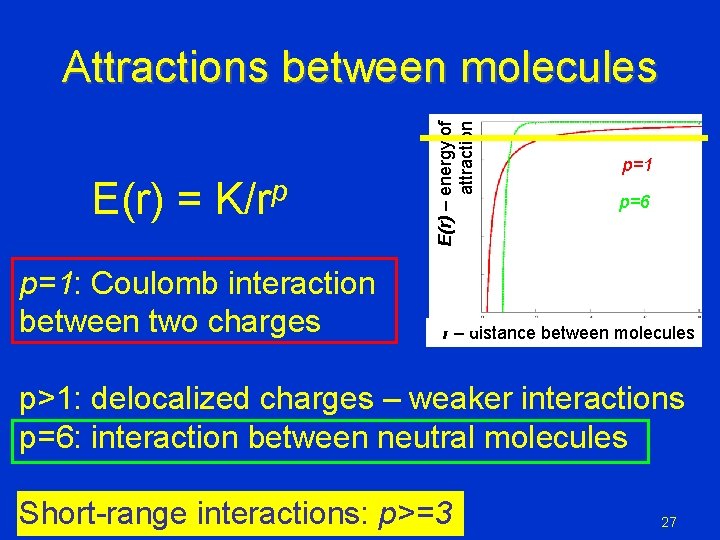
E(r) = K/rp p=1: Coulomb interaction between two charges E(r) – energy of attraction Attractions between molecules p=1 p=6 r – distance between molecules p>1: delocalized charges – weaker interactions p=6: interaction between neutral molecules Short-range interactions: p>=3 27

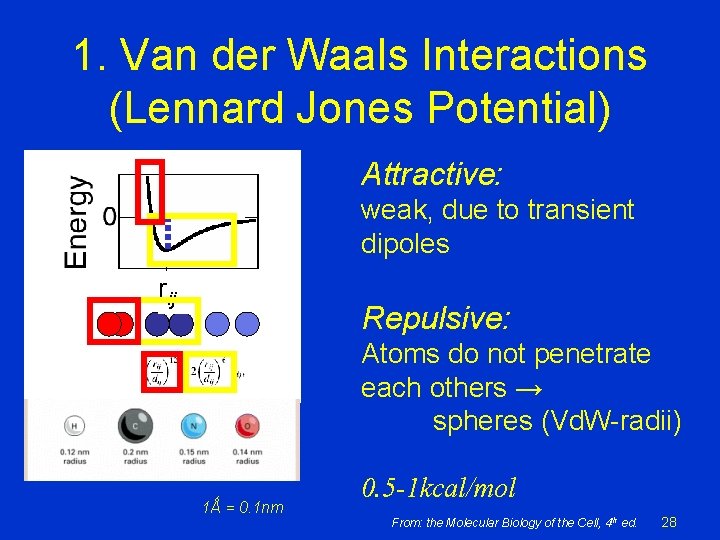
1. Van der Waals Interactions (Lennard Jones Potential) Attractive: weak, due to transient dipoles rij Repulsive: Atoms do not penetrate each others → spheres (Vd. W-radii) 1Ǻ = 0. 1 nm 0. 5 -1 kcal/mol From: the Molecular Biology of the Cell, 4 th ed. 28

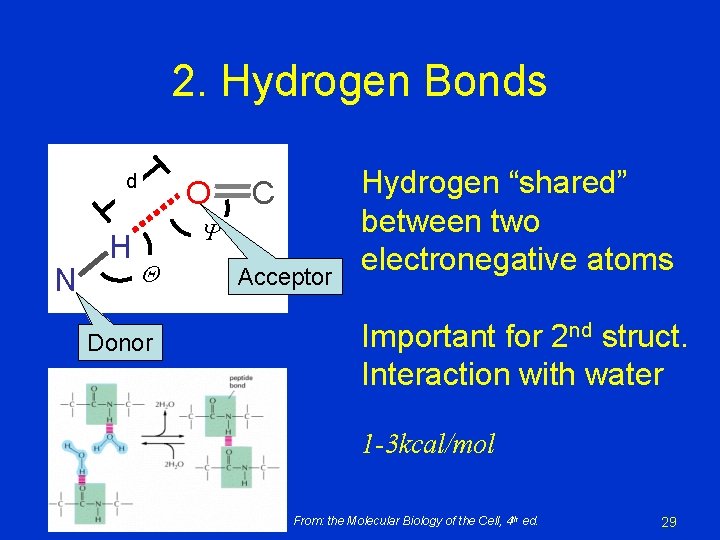
2. Hydrogen Bonds d N H O C Donor Acceptor Hydrogen “shared” between two electronegative atoms Important for 2 nd struct. Interaction with water 1 -3 kcal/mol From: the Molecular Biology of the Cell, 4 th ed. 29

3. Electrostatic Forces (Salt Bridges) Coulomb’s law: E = kq. Aq. B/Dr q. A, q. B : point charges r: distance k=332 (for units of kcal/mol) D: dielectric constant (water: 80; protein: ~4) Solvent screening: D’ = Dr. AB q. A q. B r 3Ǻ: ~1. 4 kcal/mol 30

4. Aromatic Rings: p-p and Cation-p Interactions Aromatic ring: cloud of p electrons: negative charge From: wikipedia. Can interact with • positive charge of Lys, Arg, His • edge of other aromatic ring 31

5. Water • polar • cohesive • competes with interactions in the protein (Hb, SB) • high D – reduces electrostatic forces • hydrophobic effect • explicit modeling difficult From: the Molecular Biology of the Cell, 4 th ed. 32

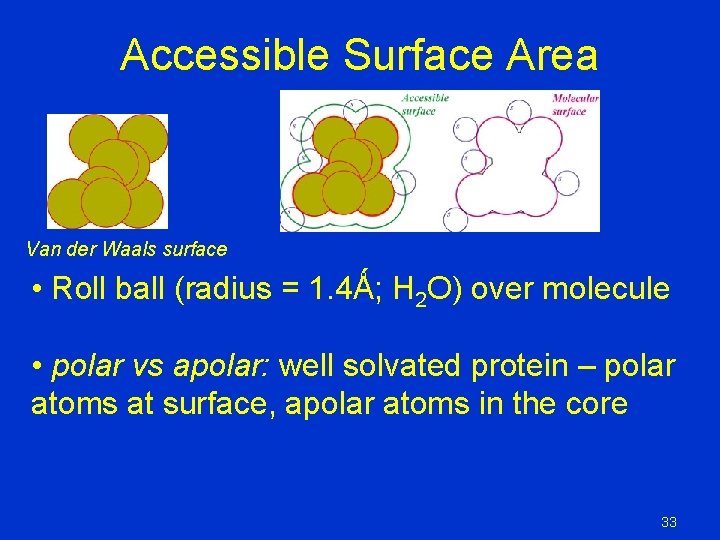
Accessible Surface Area Van der Waals surface • Roll ball (radius = 1. 4Ǻ; H 2 O) over molecule • polar vs apolar: well solvated protein – polar atoms at surface, apolar atoms in the core 33

Non-Covalent Forces: Summary • Vd. W: Many small contributions sum up to significant energy • Salt bridges and hydrogen bonds: polar interactions are reduced by competing water at the surface • Surface of protein: hydrophobic atoms should be all buried 34