Structure and Function of Eukaryotic Transcription Activators Many







- Slides: 26

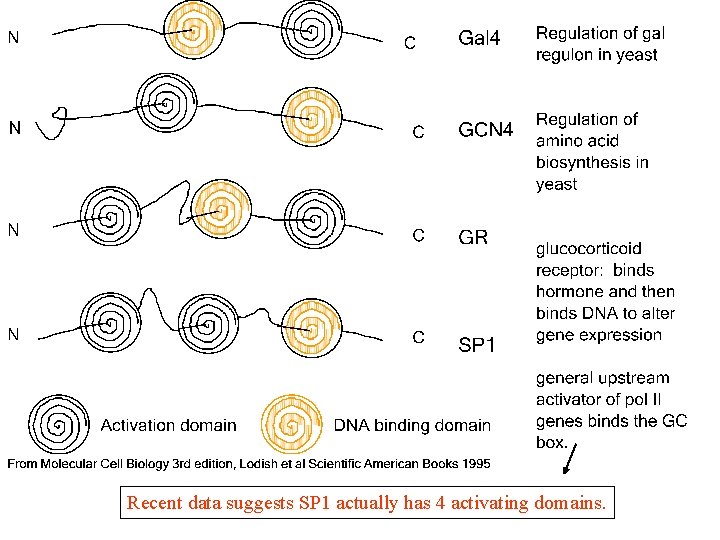
Structure and Function of Eukaryotic Transcription Activators • Many have modular structure: 1. DNA-binding domain 2. Transcription activating domain • • Proteins can have > 1 of each, and they can be in different positions in protein. Many also have a dimerization domain

Recent data suggests SP 1 actually has 4 activating domains.

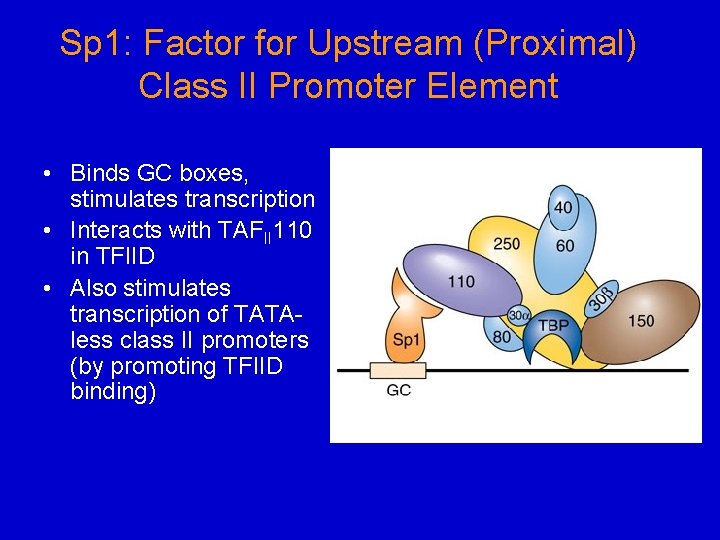
Sp 1: Factor for Upstream (Proximal) Class II Promoter Element • Binds GC boxes, stimulates transcription • Interacts with TAFII 110 in TFIID • Also stimulates transcription of TATAless class II promoters (by promoting TFIID binding)

Activation Domains 1. Acidic (e. g. , GAL 4, 49 aa domain – 11 acidic aa) 2. Glutamine-rich (e. g. , 2 in Sp 1, ~25% gln) 3. Proline-rich (e. g. , CTF, 84 aa domain – 19 are proline)

DNA-binding domains 1. Zinc–containing motifs – Zinc fingers (Sp 1 and TFIIIA) – Zinc modules (GR and other nuclear receptors) – Modules with 2 Zinc ions and 6 cysteines (GAL 4) 2. Homeodomains - 60 -aa domains originally found in homeotic mutants 3. b. ZIP and b. HLH motifs - a highly basic DNA -binding domain and a dimerization domain (leucine zipper or helix-loop-helix)

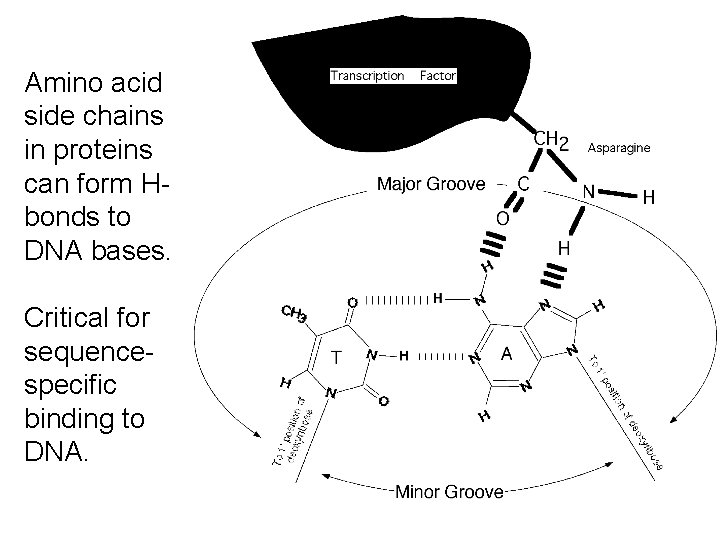
Amino acid side chains in proteins can form Hbonds to DNA bases. Critical for sequencespecific binding to DNA.

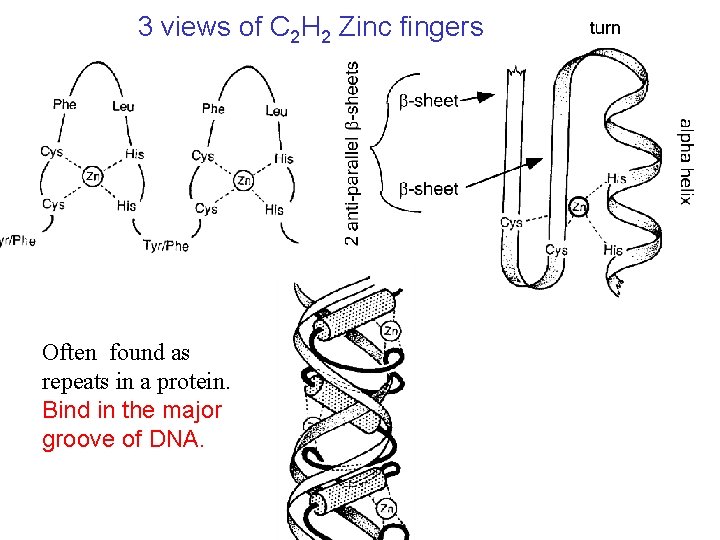
3 views of C 2 H 2 Zinc fingers Often found as repeats in a protein. Bind in the major groove of DNA.

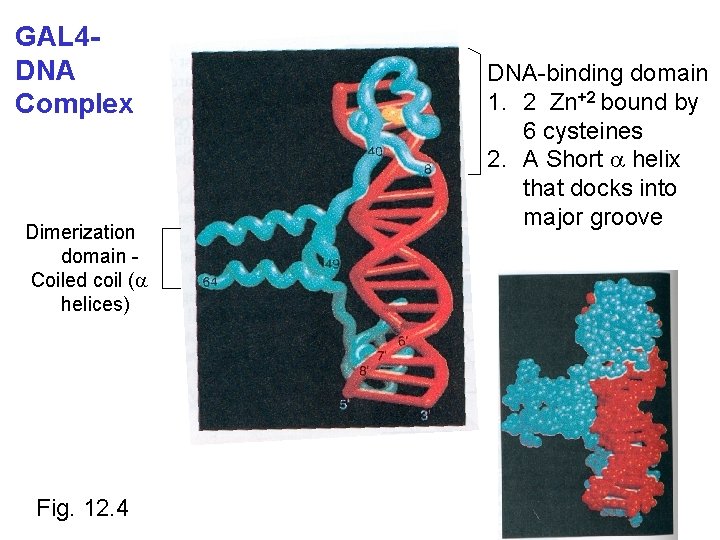
GAL 4 DNA Complex Dimerization domain Coiled coil (a helices) Fig. 12. 4 DNA-binding domain 1. 2 Zn+2 bound by 6 cysteines 2. A Short a helix that docks into major groove

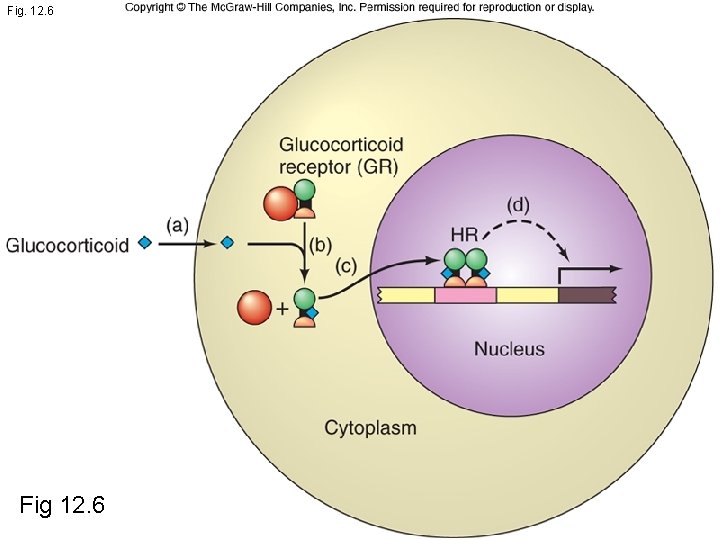
Fig. 12. 6 Fig 12. 6

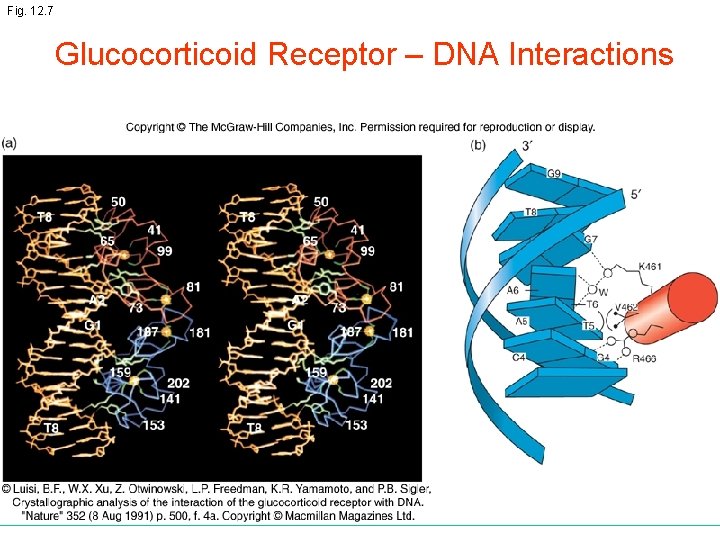
Fig. 12. 7 Glucocorticoid Receptor – DNA Interactions

- Homeotic mutants have wrong organs (organ-identity mutants) - Occur in animals and plants - Important regulatory genes Wild-type “Here’s looking at you” antennapedia

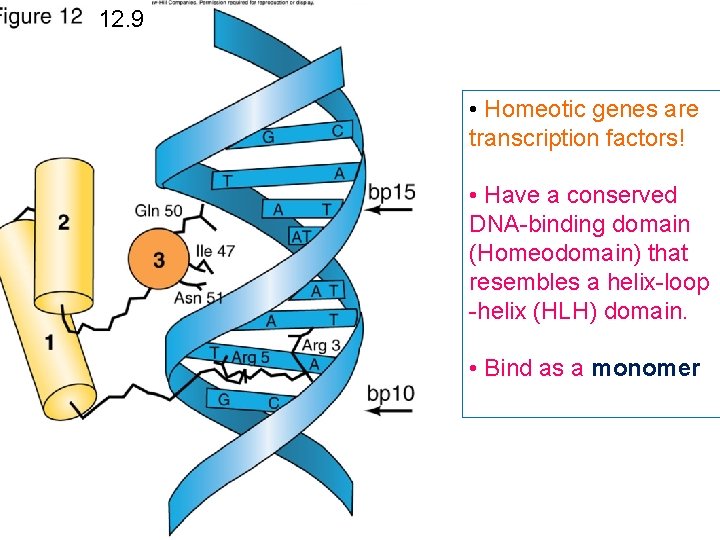
12. 9 • Homeotic genes are transcription factors! • Have a conserved DNA-binding domain (Homeodomain) that resembles a helix-loop -helix (HLH) domain. • Bind as a monomer

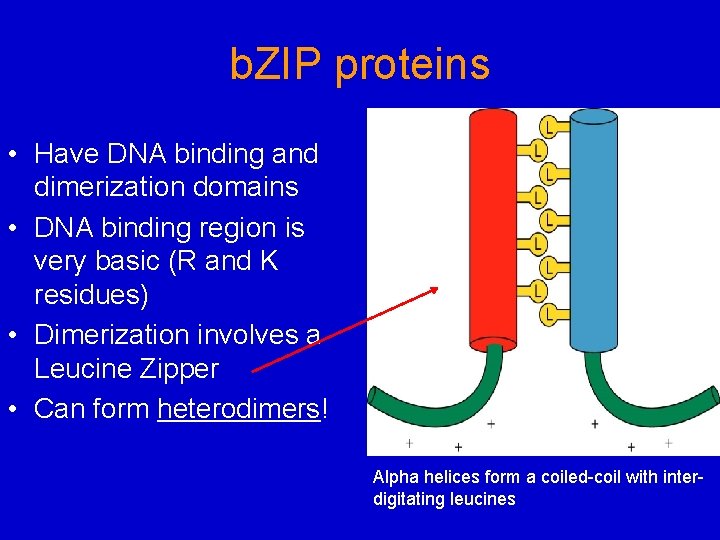
b. ZIP proteins • Have DNA binding and dimerization domains • DNA binding region is very basic (R and K residues) • Dimerization involves a Leucine Zipper • Can form heterodimers! Alpha helices form a coiled-coil with interdigitating leucines

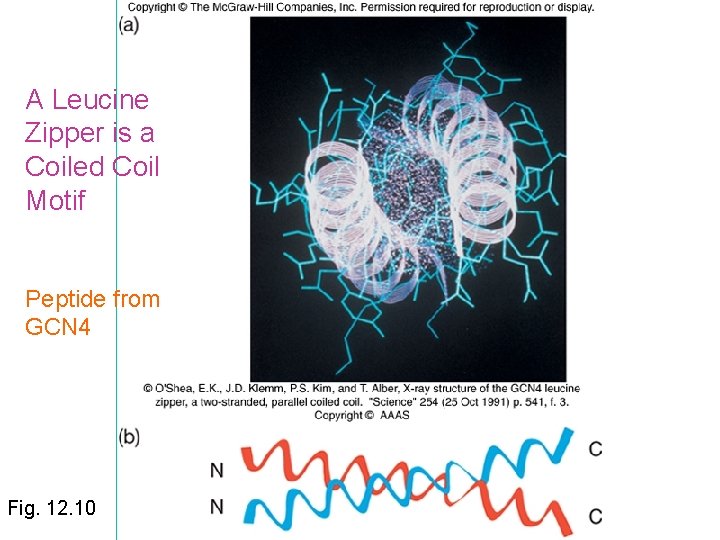
A Leucine Zipper is a Coiled Coil Motif Peptide from GCN 4 Fig. 12. 10

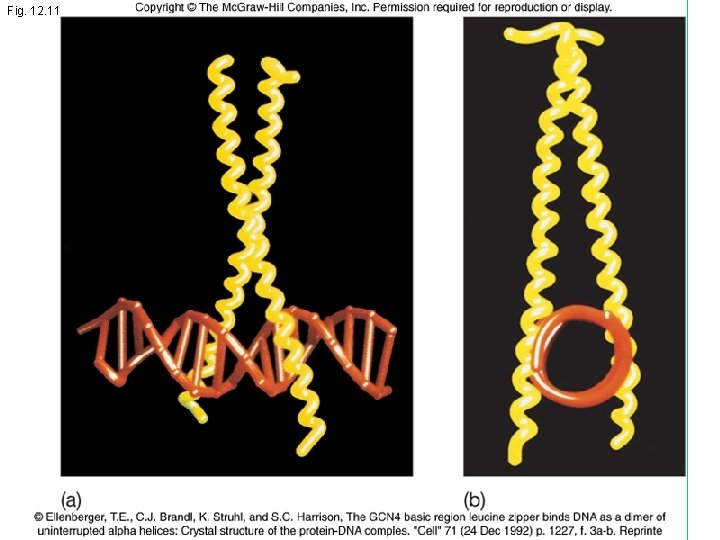
Fig. 12. 11

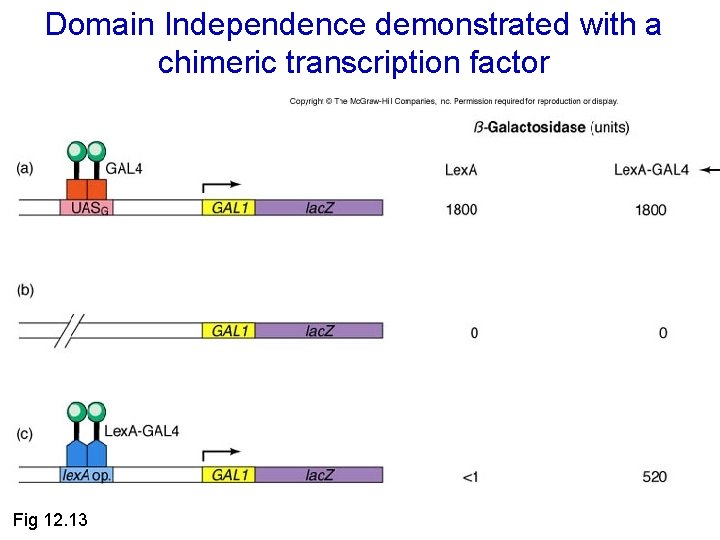
Domain Independence demonstrated with a chimeric transcription factor Fig 12. 13

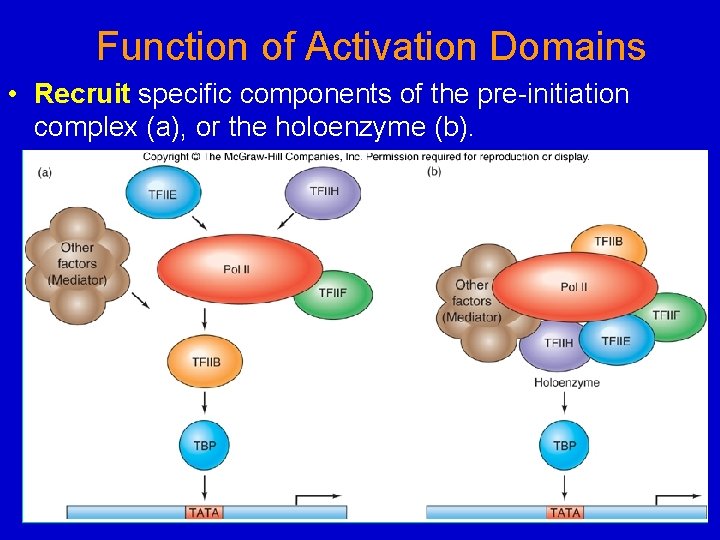
Function of Activation Domains • Recruit specific components of the pre-initiation complex (a), or the holoenzyme (b).

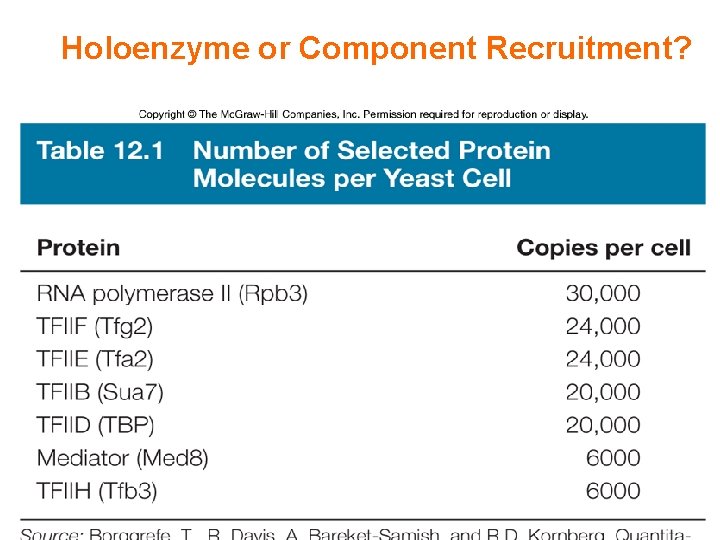
Holoenzyme or Component Recruitment?

GAL 4 (which binds to an upstream element) 1. Promotes binding of TFIIB, which promotes recruitment of the other factors and RNAP. – Probably binds directly to TFIIB (i. e. , it doesn’t work by stimulating TFIID to bind TFIIB tighter) 2. GAL 4 also promotes assembly of downstream basal factors, TFIIE and/or TFIIF+RNAP II.

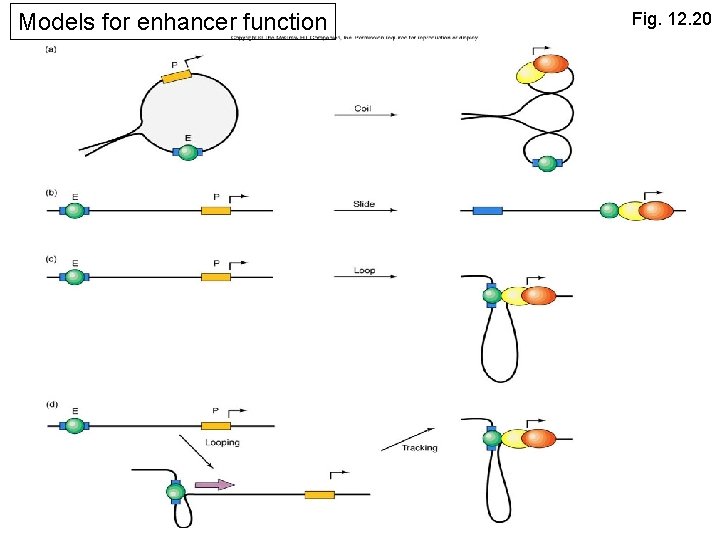
Activation from a Distance: Enhancers • There at least 4 possible models Factor binding to the enhancer induces: 1. supercoiling 2. sliding 3. Looping 4. Tracking

Models for enhancer function Fig. 12. 20

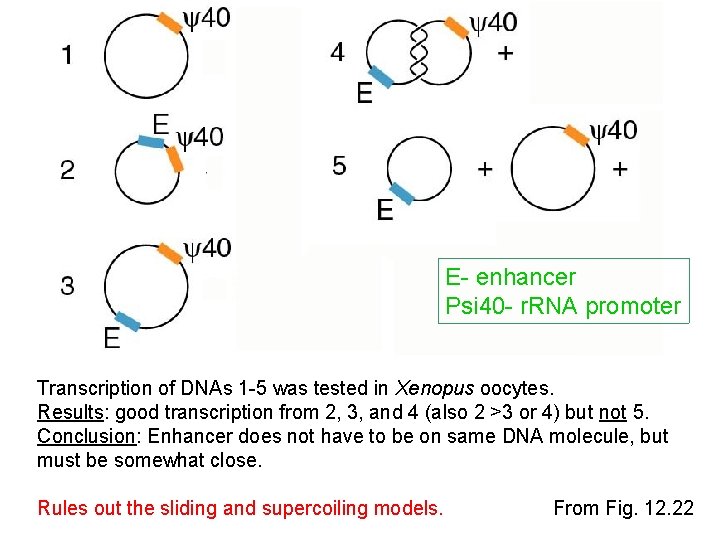
E- enhancer Psi 40 - r. RNA promoter Transcription of DNAs 1 -5 was tested in Xenopus oocytes. Results: good transcription from 2, 3, and 4 (also 2 >3 or 4) but not 5. Conclusion: Enhancer does not have to be on same DNA molecule, but must be somewhat close. Rules out the sliding and supercoiling models. From Fig. 12. 22

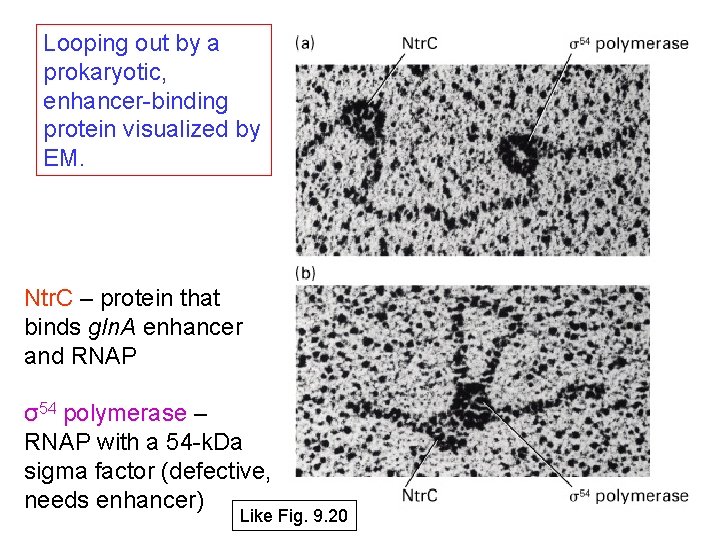
Looping out by a prokaryotic, enhancer-binding protein visualized by EM. Ntr. C – protein that binds gln. A enhancer and RNAP σ54 polymerase – RNAP with a 54 -k. Da sigma factor (defective, needs enhancer) Like Fig. 9. 20

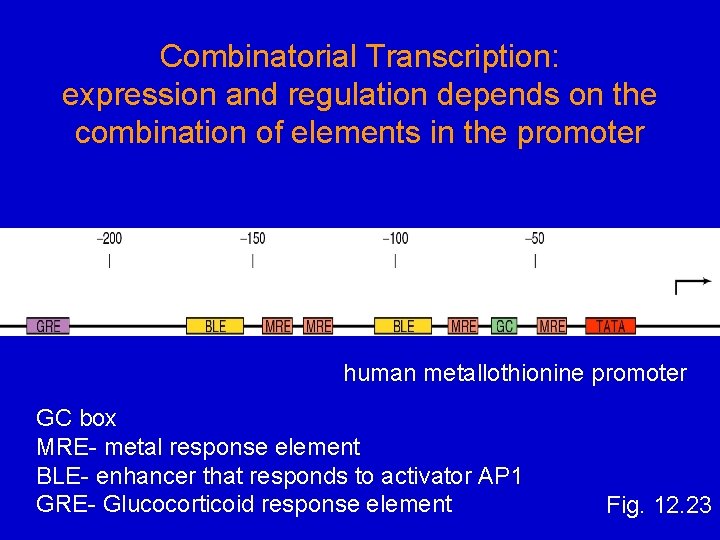
Combinatorial Transcription: expression and regulation depends on the combination of elements in the promoter human metallothionine promoter GC box MRE- metal response element BLE- enhancer that responds to activator AP 1 GRE- Glucocorticoid response element Fig. 12. 23

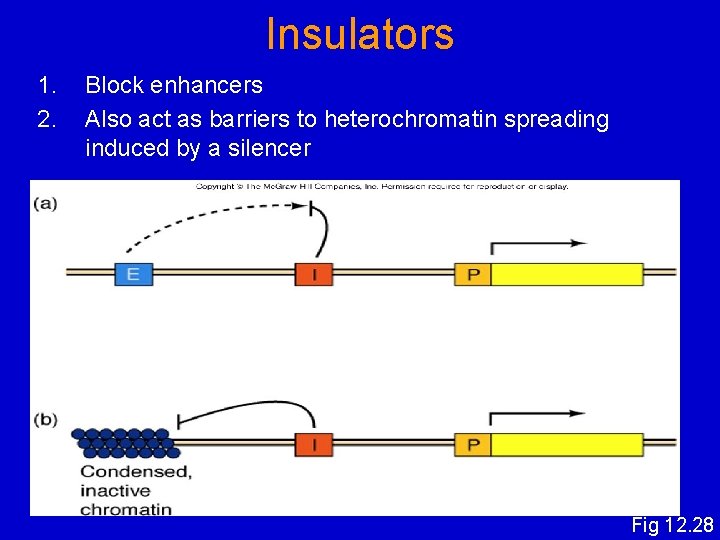
Insulators 1. 2. Block enhancers Also act as barriers to heterochromatin spreading induced by a silencer Fig 12. 28

Regulation of Transcription factors or “Regulating the Regulators” A lot of post-translational regulation: Why? - Quicker response time - Avoid silencing by keeping the transcription factor gene on (? ) Some of the mechanisms: 1. Coactivators or mediators 2. Phosphorylation-dephosphorylation: can be + or 3. Ubiquitination (deubiquitination): covalent attachment of ubiquitin (small protein) to lysines can modulate activity or trigger destruction 4. Sumoylation: covalent attachment of SUMO (small ubiquitin-like modifier) peptide to lysines, factor is inactivated but not destroyed 5. Acetylation: histone acetyltransferases (HATs) acetylate lysines on histone and non-histone proteins, can be + or -