Structural genomics includes the genetic mapping physical mapping









- Slides: 28

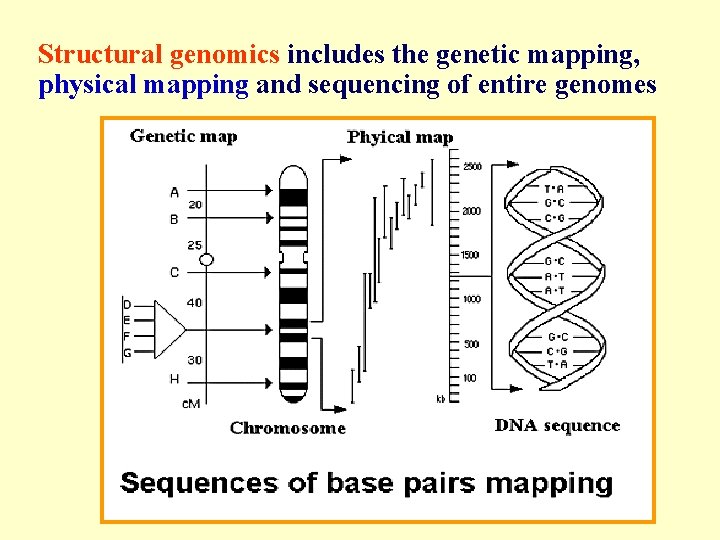
Structural genomics includes the genetic mapping, physical mapping and sequencing of entire genomes

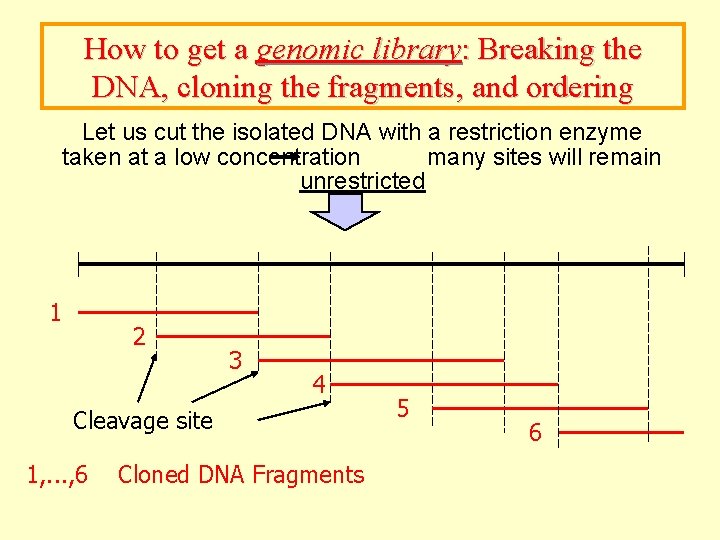
How to get a genomic library: Breaking the DNA, cloning the fragments, and ordering Let us cut the isolated DNA with a restriction enzyme taken at a low concentration many sites will remain unrestricted 1 2 3 4 Cleavage site 1, . . . , 6 Cloned DNA Fragments 5 6

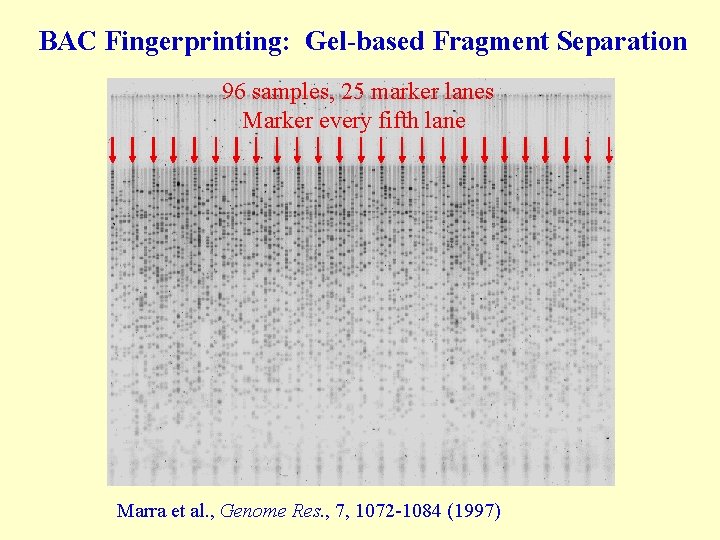
BAC Fingerprinting: Gel-based Fragment Separation 96 samples, 25 marker lanes Marker every fifth lane Marra et al. , Genome Res. , 7, 1072 -1084 (1997)

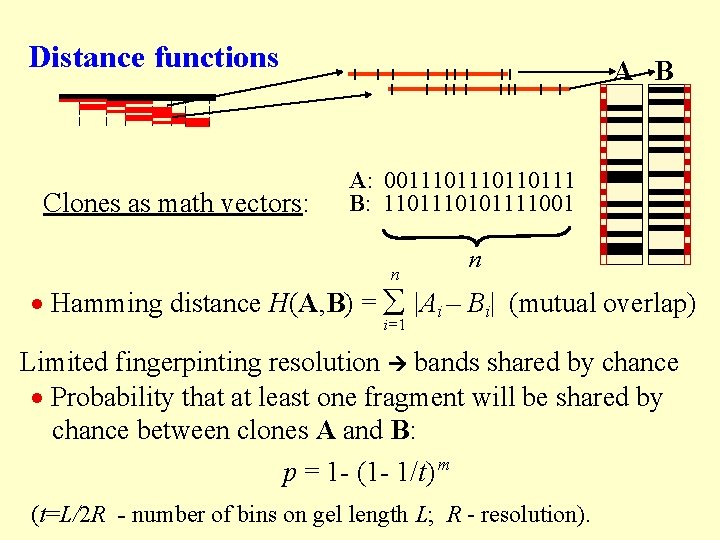
Distance functions Clones as math vectors: A B A: 001110110111 B: 110101111001 n n Hamming distance H(A, B) = |Ai – Bi| (mutual overlap) i=1 Limited fingerpinting resolution bands shared by chance Probability that at least one fragment will be shared by chance between clones A and B: p = 1 - (1 - 1/t) m (t=L/2 R - number of bins on gel length L; R - resolution).

Genome physical mapping problems are computationally challenging “… We have been looking at the assemblies of large genomes … and for every ‘draft’ genome we look at, we find hundreds - and sometimes thousands - of mis-assemblies”. Salzberg & Yorke (2005) Beware of mis-assembled genomes. Bioinformatics, 21: 4320 -4322

Which factors may affect the quality of physical map ? Bioinformatics and Human Factors Reading the scores Clustering (contig assembly) Ordering the clusters Merging contigs Anchoring (getting genetic and physical maps together) Verification of mapping results (at each stage) Where bioinformatics can help ?

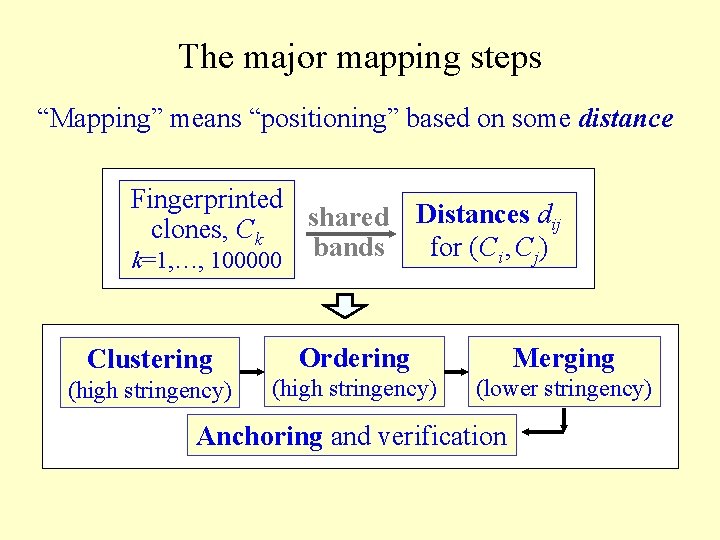
The major mapping steps “Mapping” means “positioning” based on some distance Fingerprinted Distances dij shared clones, Ck for (Ci , Cj) k=1, …, 100000 bands Clustering Ordering Merging (high stringency) (lower stringency) Anchoring and verification

P-value of clone overlaps Sulston score (Sulston et al. , 1988): p = 1 -(1 -1/N)n(c 2) is the probability of random incidence of two bands; n(c) – number of bands in clone c; N – total number of distinguishable bands

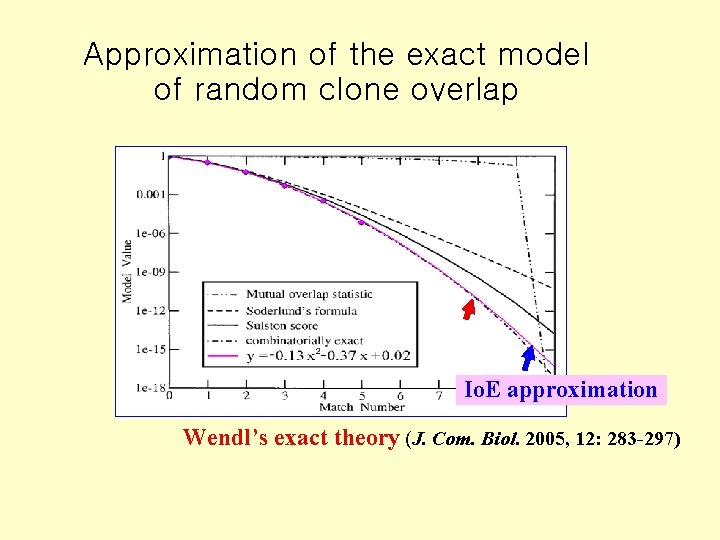
Approximation of the exact model of random clone overlap Io. E approximation Wendl’s exact theory (J. Com. Biol. 2005, 12: 283 -297)

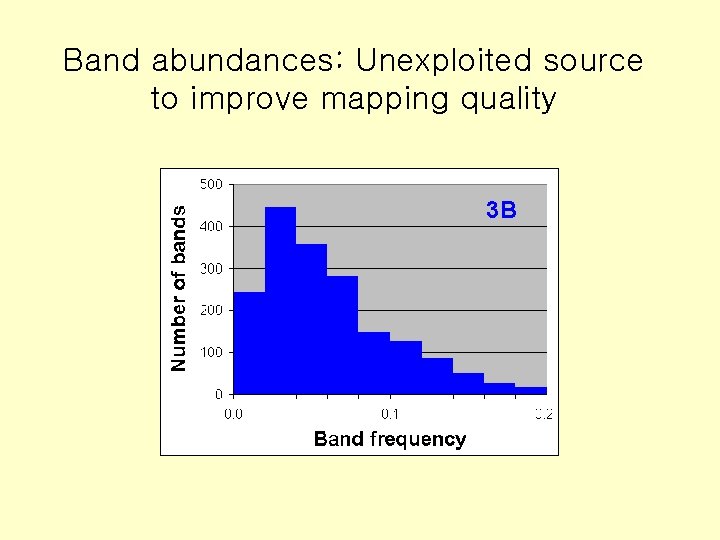
Band abundances: Unexploited source to improve mapping quality 3 B

Adaptive Clustering Varying cutoff: increasing rather than decreasing stringency 1100 t c e t o pr s r e t s u ed cl 244

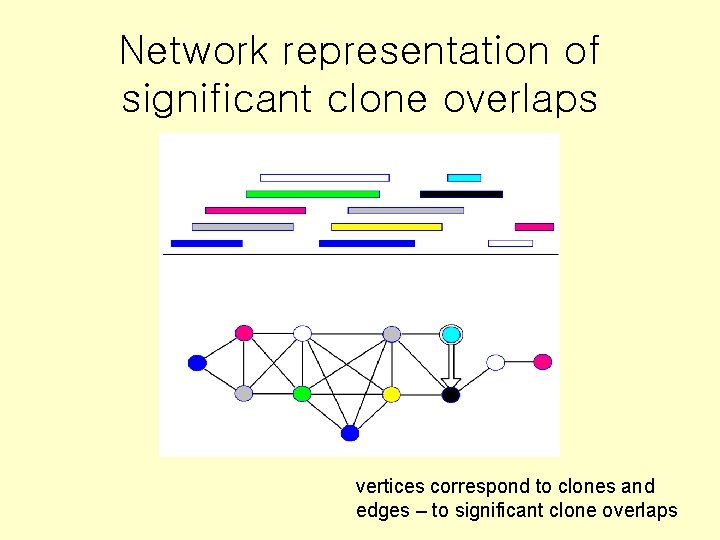
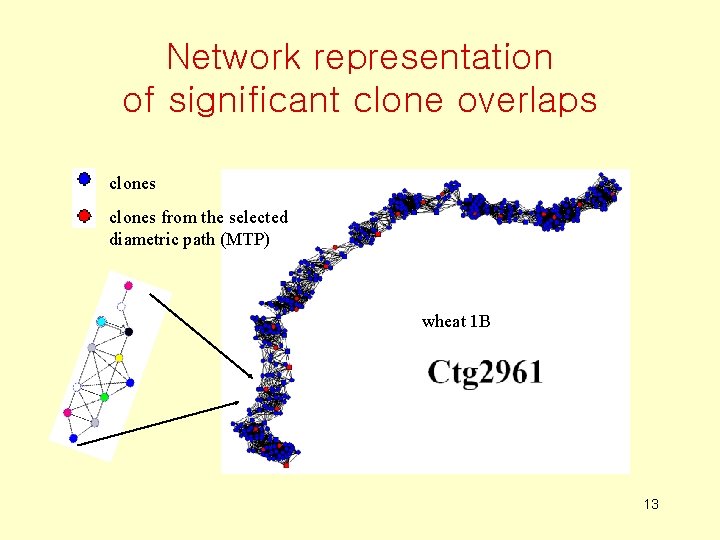
Network representation of significant clone overlaps vertices correspond to clones and edges – to significant clone overlaps

Network representation of significant clone overlaps clones from the selected diametric path (MTP) wheat 1 B 13

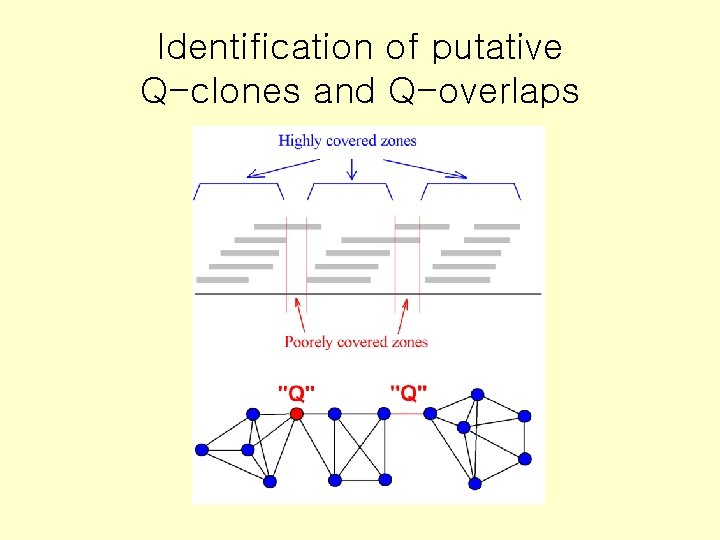
Identification of putative Q-clones and Q-overlaps

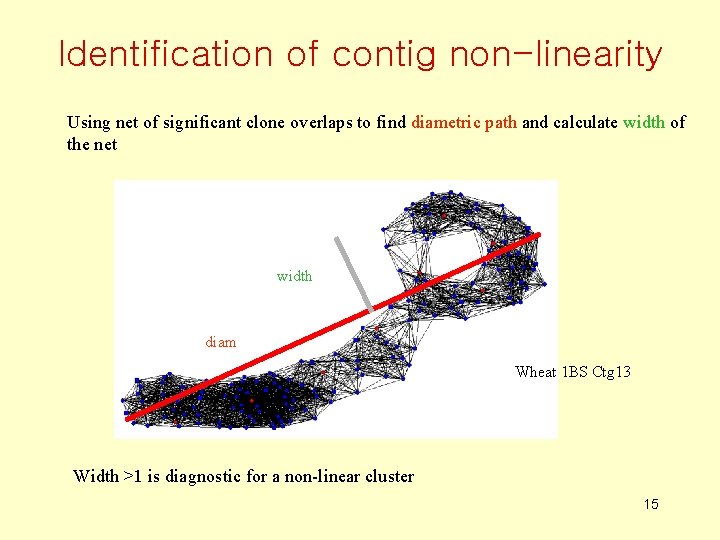
Identification of contig non-linearity Using net of significant clone overlaps to find diametric path and calculate width of the net width diam Wheat 1 BS Ctg 13 Width >1 is diagnostic for a non-linear cluster 15

Identification of contig non-linearity 0 Diametric path: 1 • Calculate ranks rj=rj(ci) for all clones cj relative to clone ci (through significant clone overlaps). 2 • Diametric path ( MTP) is the shortest path through significant clone overlaps connecting clones ci and cj with maximal rj(ci). 6 3 4 5 7 8 9 • Width of net: maximal rank relative to diametric path • Width >1 non-linear cluster 0 1 2 16

Identification of contig non-linearity Example with Q-clone: 17

Identification of contig non-linearity • Using net of significant clone overlaps, for each clone ci calculate ranks rij for all clones cj. • Diametric path: for pair of clones with maximal rij identify the shortest path through significant clone overlaps MTP • Width of net: maximal rank relative to diametric path • Width >1 is diagnostic for a non-linear cluster PAG-19 2011

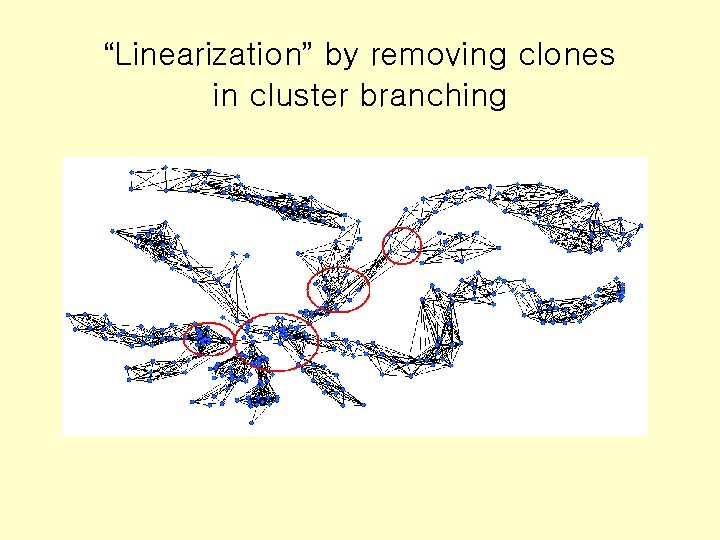
“Linearization” by removing clones in cluster branching

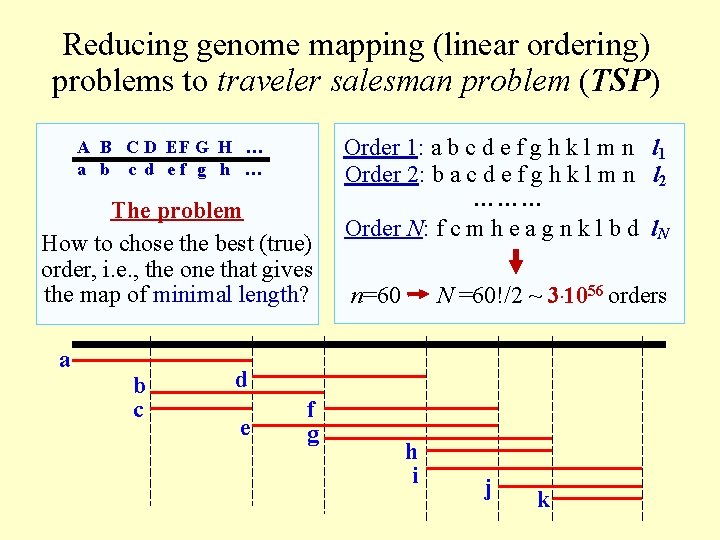
Reducing genome mapping (linear ordering) problems to traveler salesman problem (TSP) A B C D EF G H … a b c d ef g h … The problem How to chose the best (true) order, i. e. , the one that gives the map of minimal length? a b c Order 1: a b c d e f g h k l m n l 1 Order 2: b a c d e f g h k l m n l 2 ……… Order N: f c m h e a g n k l b d l. N n=60 N =60!/2 ~ 3. 1056 orders d e f g h i j k

Example: A Contig

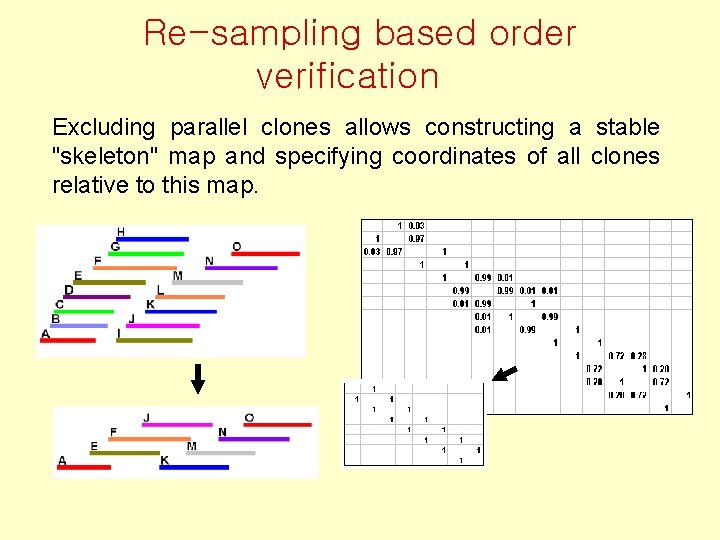
Re-sampling based order verification Excluding parallel clones allows constructing a stable "skeleton" map and specifying coordinates of all clones relative to this map.

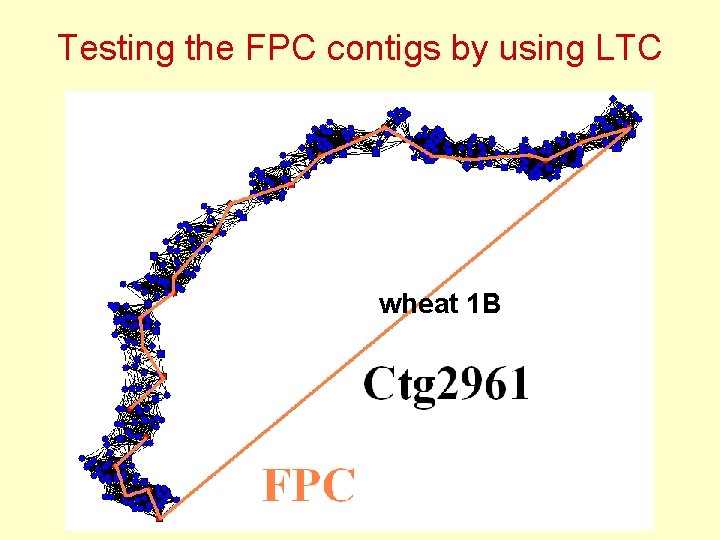
Testing the FPC contigs by using LTC wheat 1 B

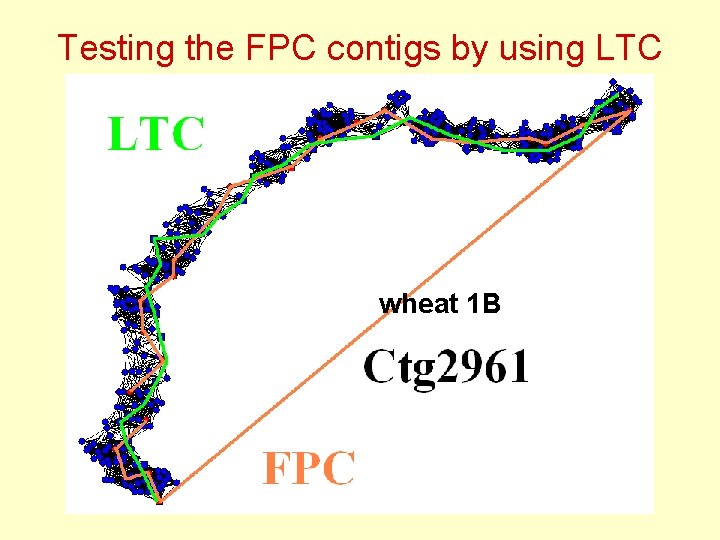
Testing the FPC contigs by using LTC wheat 1 B

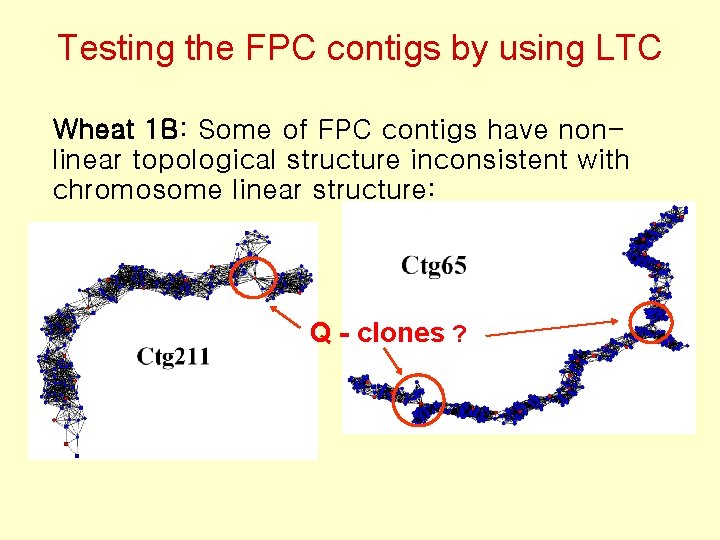
Testing the FPC contigs by using LTC Wheat 1 B: Some of FPC contigs have nonlinear topological structure inconsistent with chromosome linear structure: Q - clones ?

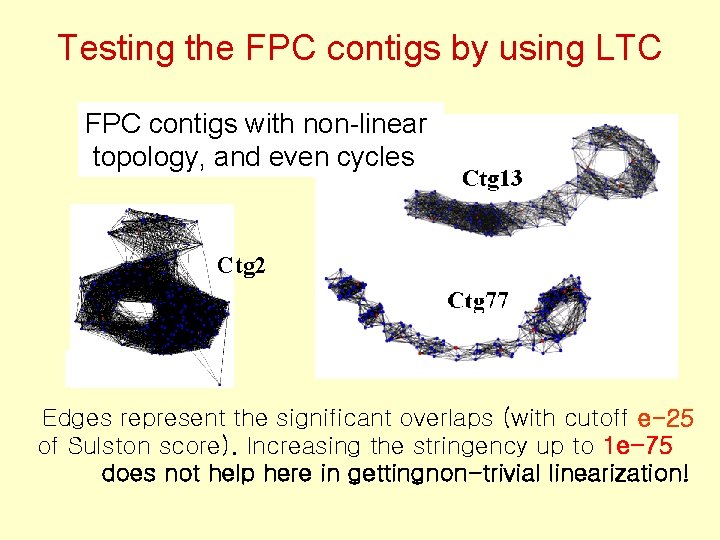
Testing the FPC contigs by using LTC FPC contigs with non-linear topology, and even cycles Ctg 2 Edges represent the significant overlaps (with cutoff e-25 of Sulston score). Increasing the stringency up to 1 e-75 does not help here in gettingnon-trivial linearization!

Problematic contigs (simulated maize)

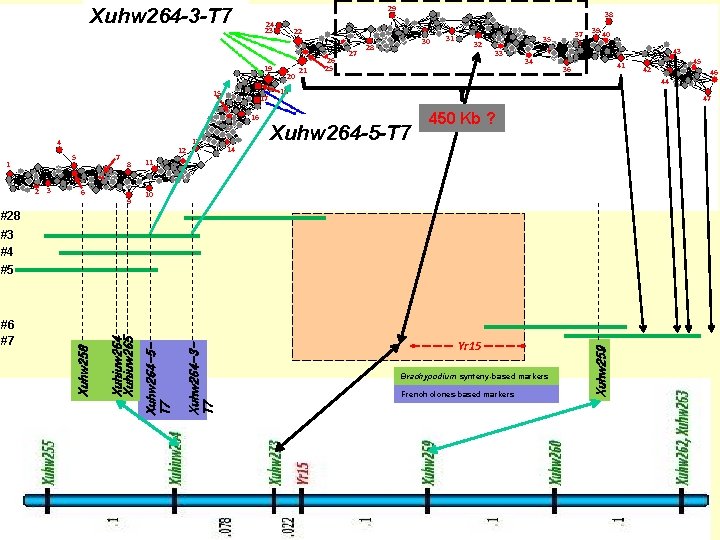
29 Xuhw 264 -3 -T 7 24 23 19 15 17 16 4 5 1 2 3 7 6 12 8 9 13 38 22 20 21 26 25 27 30 28 31 32 37 35 33 39 40 43 34 41 36 44 18 Xuhw 264 -5 -T 7 450 Kb ? 14 11 10 #3 #4 Yr 15 Brachypodium synteny-based markers French clones-based markers Xuhw 259 Xuhw 264 -3 T 7 Xuhw 264 -5 T 7 Xuhiuw 264 Xuhiuw 265 #5 Xuhw 258 46 47 #28 #6 #7 45 42