Structural dynamics of HER 2 and Erb B

Structural dynamics of HER 2 and Erb. B 4: Yin and Yang in Mammary Carcinoma Shannon Telesco Advisor: Ravi Radhakrishnan, Ph. D. Department of Bioengineering
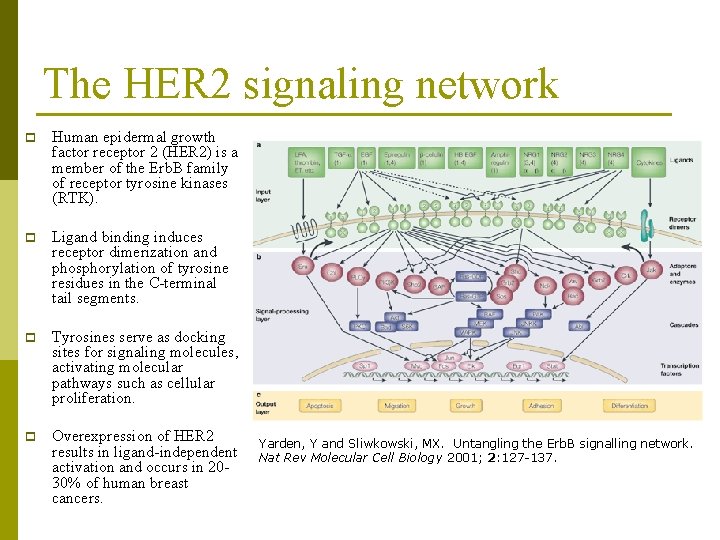
The HER 2 signaling network p Human epidermal growth factor receptor 2 (HER 2) is a member of the Erb. B family of receptor tyrosine kinases (RTK). p Ligand binding induces receptor dimerization and phosphorylation of tyrosine residues in the C-terminal tail segments. p Tyrosines serve as docking sites for signaling molecules, activating molecular pathways such as cellular proliferation. p Overexpression of HER 2 results in ligand-independent activation and occurs in 2030% of human breast cancers. Yarden, Y and Sliwkowski, MX. Untangling the Erb. B signalling network. Nat Rev Molecular Cell Biology 2001; 2: 127 -137.
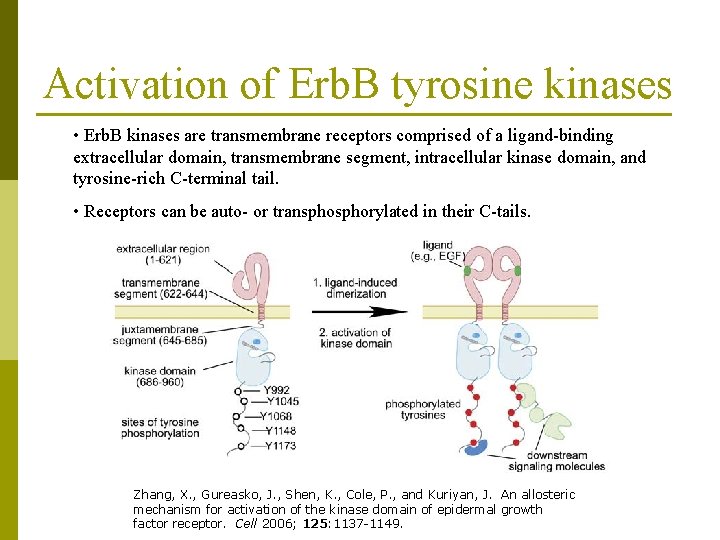
Activation of Erb. B tyrosine kinases • Erb. B kinases are transmembrane receptors comprised of a ligand-binding extracellular domain, transmembrane segment, intracellular kinase domain, and tyrosine-rich C-terminal tail. • Receptors can be auto- or transphorylated in their C-tails. Zhang, X. , Gureasko, J. , Shen, K. , Cole, P. , and Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 2006; 125: 1137 -1149.
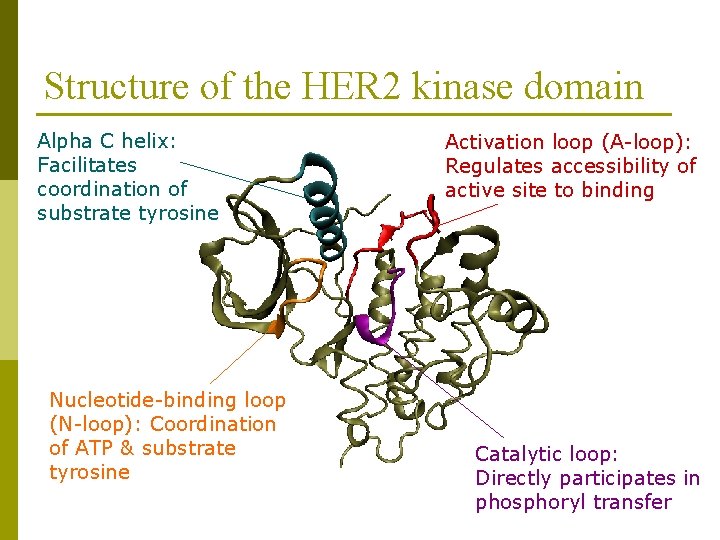
Structure of the HER 2 kinase domain Alpha C helix: Facilitates coordination of substrate tyrosine Nucleotide-binding loop (N-loop): Coordination of ATP & substrate tyrosine Activation loop (A-loop): Regulates accessibility of active site to binding Catalytic loop: Directly participates in phosphoryl transfer
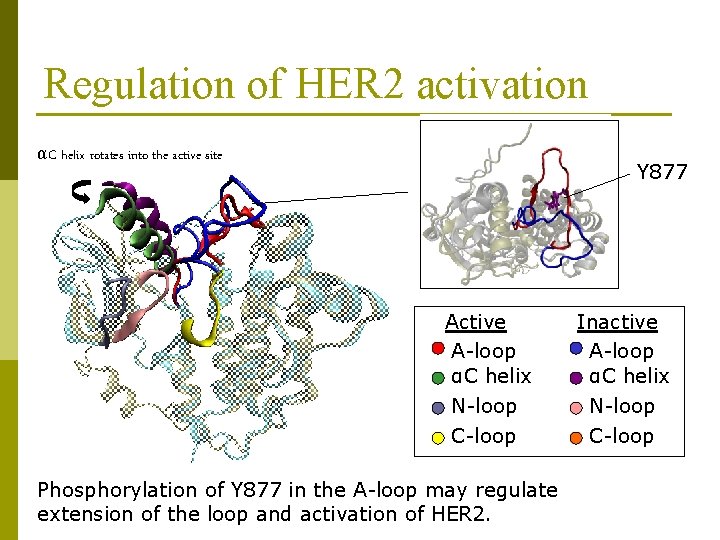
Regulation of HER 2 activation αC helix rotates into the active site Y 877 Active A-loop αC helix N-loop C-loop Phosphorylation of Y 877 in the A-loop may regulate extension of the loop and activation of HER 2. Inactive A-loop αC helix N-loop C-loop

Elucidating HER 2 activation mechanism p Investigate the structural differences between inactive and active HER 2. What are the key bonds that must be formed or broken upon activation? p Define the role of Y 877 phosphorylation in HER 2 activation. Is HER 2 unique from other Erb. B members in that P-Y 877 is necessary for activity? p Predict the behavior of an EGFR/HER 2 heterodimer. How might the dimerization interface trigger conformational changes in HER 2?
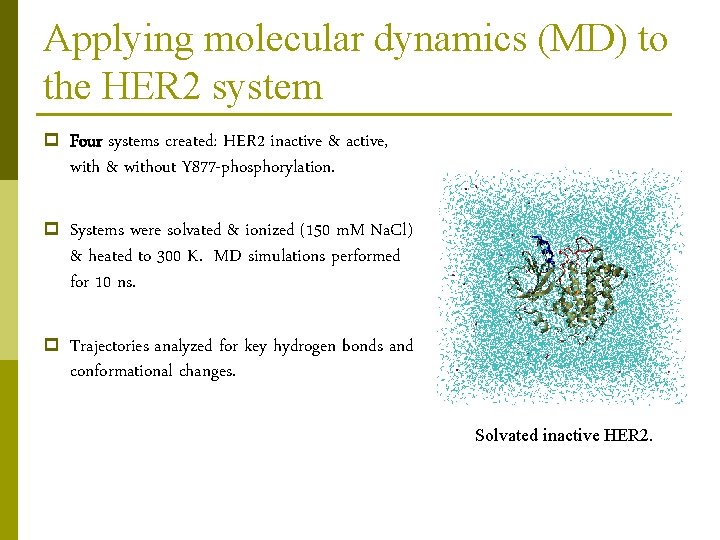
Applying molecular dynamics (MD) to the HER 2 system p Four systems created: HER 2 inactive & active, with & without Y 877 -phosphorylation. p Systems were solvated & ionized (150 m. M Na. Cl) & heated to 300 K. MD simulations performed for 10 ns. p Trajectories analyzed for key hydrogen bonds and conformational changes. Solvated inactive HER 2.

Hydrogen bonds in the A-loop Salt Bridges Conserved bond Hydrogen Bonds A-loop

Hydrogen bonds in the αC helix Salt Bridges Key salt bridge Hydrogen Bonds αC helix
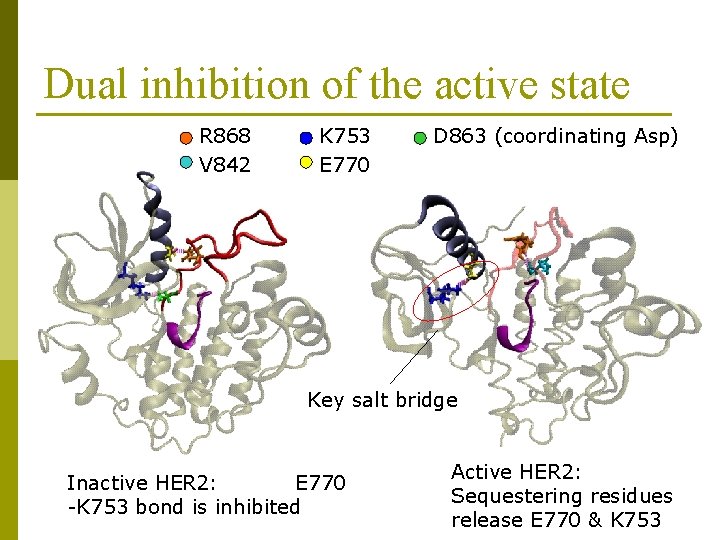
Dual inhibition of the active state R 868 V 842 K 753 E 770 D 863 (coordinating Asp) Key salt bridge Inactive HER 2: E 770 -K 753 bond is inhibited Active HER 2: Sequestering residues release E 770 & K 753

Stabilizing H-bonds in the active state Active A-loop Active αC helix K 883 Inactive A-loop Inactive αC helix E 766 • Key salt bridge in active HER 2 is K 883 -E 766 • Connects the αC helix with the A-loop, stabilizing the helix in the active site • Bond is conserved among Erb. B family members: K 851 -E 734 (EGFR), K 856 E 739 (Erb. B 4) Inactive/active HER 2 (superimposed)
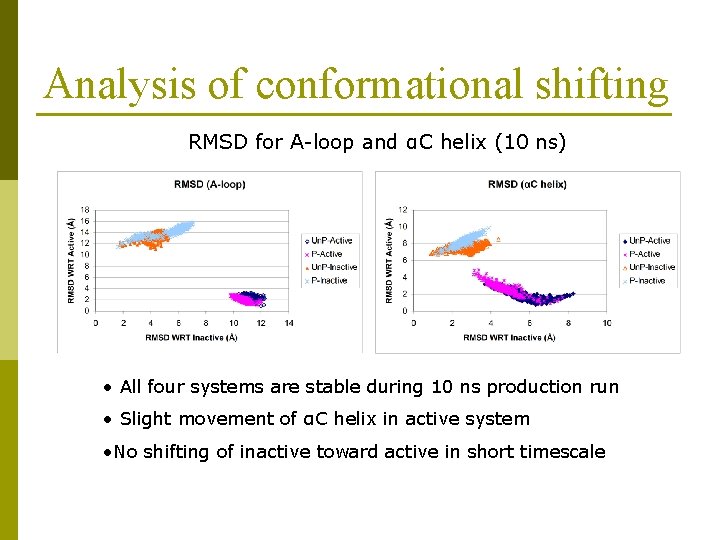
Analysis of conformational shifting RMSD for A-loop and αC helix (10 ns) • All four systems are stable during 10 ns production run • Slight movement of αC helix in active system • No shifting of inactive toward active in short timescale
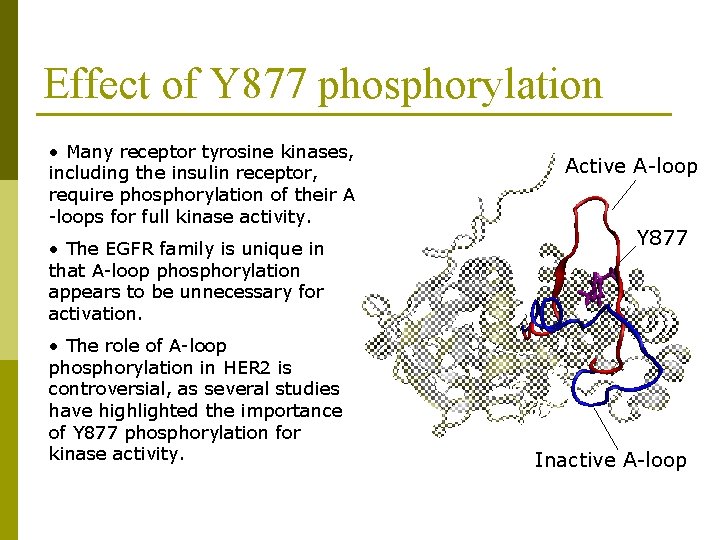
Effect of Y 877 phosphorylation • Many receptor tyrosine kinases, including the insulin receptor, require phosphorylation of their A -loops for full kinase activity. • The EGFR family is unique in that A-loop phosphorylation appears to be unnecessary for activation. • The role of A-loop phosphorylation in HER 2 is controversial, as several studies have highlighted the importance of Y 877 phosphorylation for kinase activity. Active A-loop Y 877 Inactive A-loop
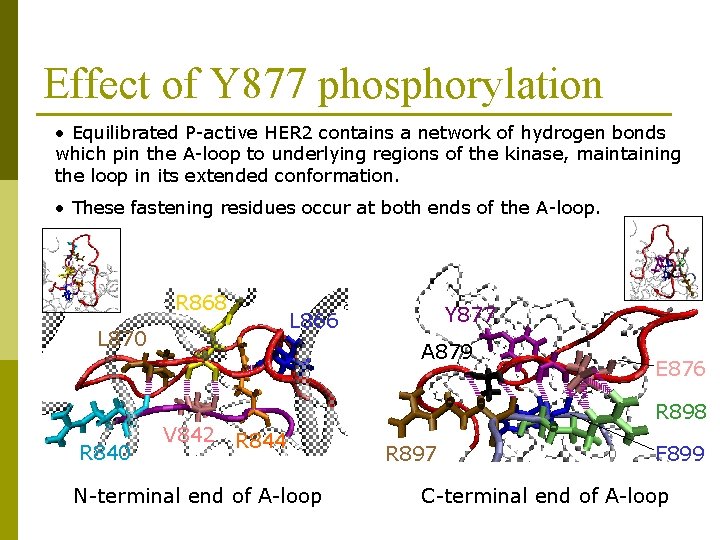
Effect of Y 877 phosphorylation • Equilibrated P-active HER 2 contains a network of hydrogen bonds which pin the A-loop to underlying regions of the kinase, maintaining the loop in its extended conformation. • These fastening residues occur at both ends of the A-loop. R 868 L 870 R 840 Y 877 L 866 A 879 V 842 E 876 R 898 R 844 N-terminal end of A-loop R 897 F 899 C-terminal end of A-loop
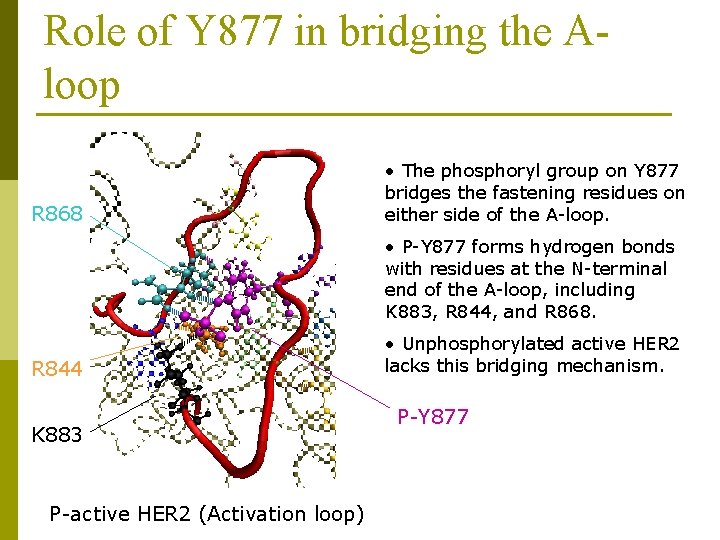
Role of Y 877 in bridging the Aloop R 868 • The phosphoryl group on Y 877 bridges the fastening residues on either side of the A-loop. • P-Y 877 forms hydrogen bonds with residues at the N-terminal end of the A-loop, including K 883, R 844, and R 868. R 844 K 883 P-active HER 2 (Activation loop) • Unphosphorylated active HER 2 lacks this bridging mechanism. P-Y 877
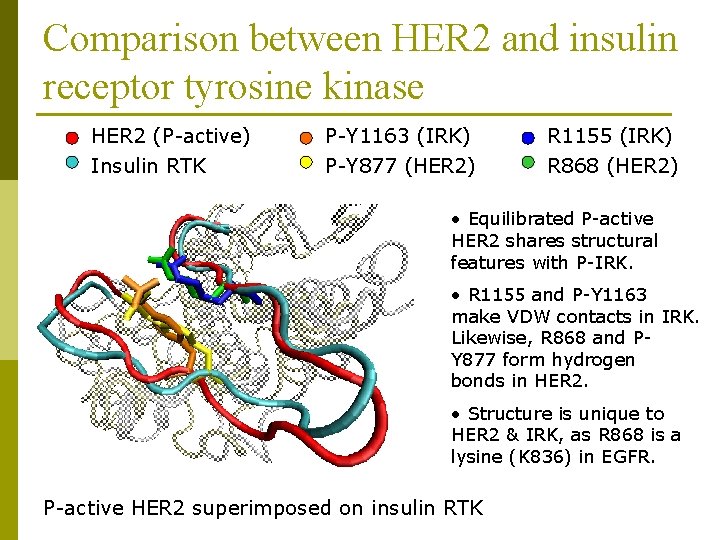
Comparison between HER 2 and insulin receptor tyrosine kinase HER 2 (P-active) P-Y 1163 (IRK) R 1155 (IRK) Insulin RTK P-Y 877 (HER 2) R 868 (HER 2) • Equilibrated P-active HER 2 shares structural features with P-IRK. • R 1155 and P-Y 1163 make VDW contacts in IRK. Likewise, R 868 and PY 877 form hydrogen bonds in HER 2. • Structure is unique to HER 2 & IRK, as R 868 is a lysine (K 836) in EGFR. P-active HER 2 superimposed on insulin RTK
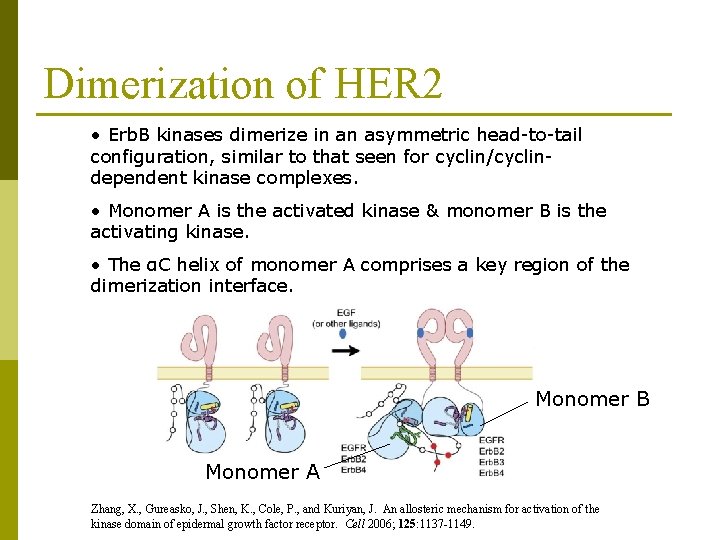
Dimerization of HER 2 • Erb. B kinases dimerize in an asymmetric head-to-tail configuration, similar to that seen for cyclin/cyclindependent kinase complexes. • Monomer A is the activated kinase & monomer B is the activating kinase. • The αC helix of monomer A comprises a key region of the dimerization interface. Monomer B Monomer A Zhang, X. , Gureasko, J. , Shen, K. , Cole, P. , and Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 2006; 125: 1137 -1149.
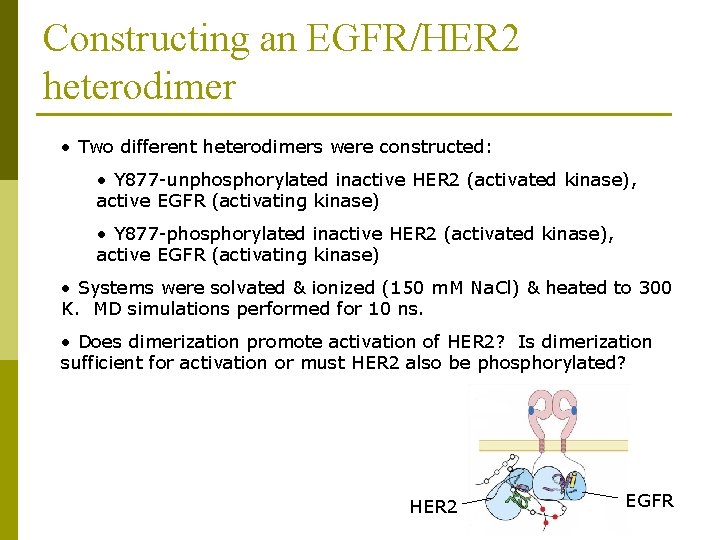
Constructing an EGFR/HER 2 heterodimer • Two different heterodimers were constructed: • Y 877 -unphosphorylated inactive HER 2 (activated kinase), active EGFR (activating kinase) • Y 877 -phosphorylated inactive HER 2 (activated kinase), active EGFR (activating kinase) • Systems were solvated & ionized (150 m. M Na. Cl) & heated to 300 K. MD simulations performed for 10 ns. • Does dimerization promote activation of HER 2? Is dimerization sufficient for activation or must HER 2 also be phosphorylated? HER 2 EGFR
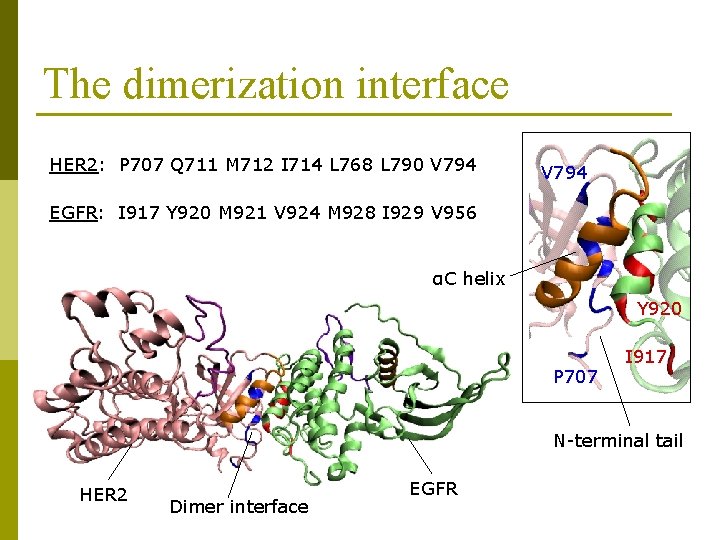
The dimerization interface HER 2: P 707 Q 711 M 712 I 714 L 768 L 790 V 794 EGFR: I 917 Y 920 M 921 V 924 M 928 I 929 V 956 αC helix Y 920 P 707 I 917 N-terminal tail HER 2 Dimer interface EGFR
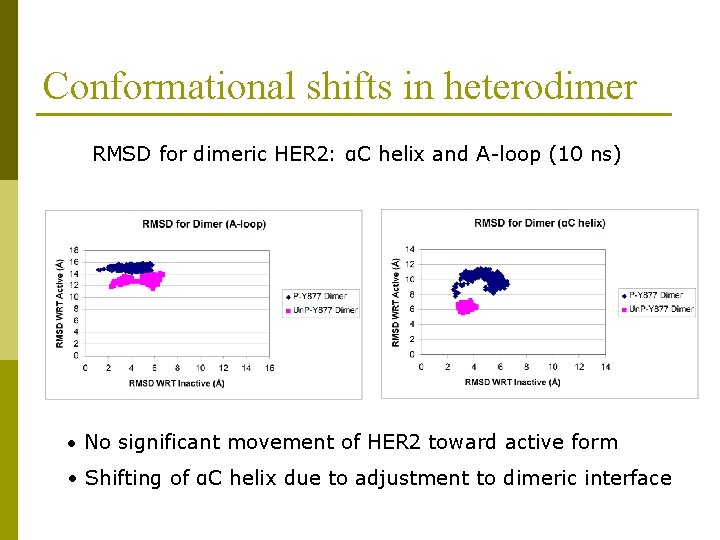
Conformational shifts in heterodimer RMSD for dimeric HER 2: αC helix and A-loop (10 ns) • No significant movement of HER 2 toward active form • Shifting of αC helix due to adjustment to dimeric interface
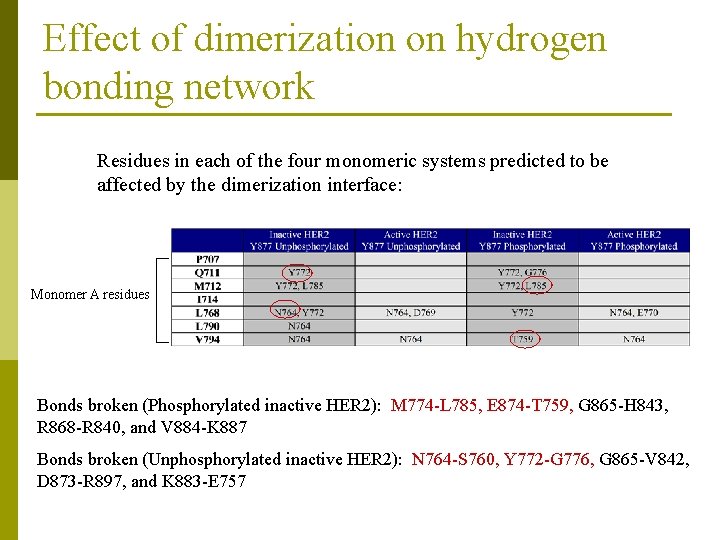
Effect of dimerization on hydrogen bonding network Residues in each of the four monomeric systems predicted to be affected by the dimerization interface: Monomer A residues Bonds broken (Phosphorylated inactive HER 2): M 774 -L 785, E 874 -T 759, G 865 -H 843, R 868 -R 840, and V 884 -K 887 Bonds broken (Unphosphorylated inactive HER 2): N 764 -S 760, Y 772 -G 776, G 865 -V 842, D 873 -R 897, and K 883 -E 757

Conclusions • Inactive and active HER 2 structures reveal distinctive hydrogen bonding patterns that stabilize each conformation. A dual inhibitory mechanism maintains the inactive state through sequestration of key residues required for activation. • Phosphorylation of Y 877 may serve to bridge the stabilizing hydrogen bonds on either side of the A-loop in the active conformation. Unphosphorylated active HER 2 lacks this bridging mechanism. • Formation of EGFR/HER 2 heterodimer results in repositioning of the αC helix and breakage of several key bonds that are present in the inactive state.

Part II: Role of Erb. B 4 signaling in the mammary gland
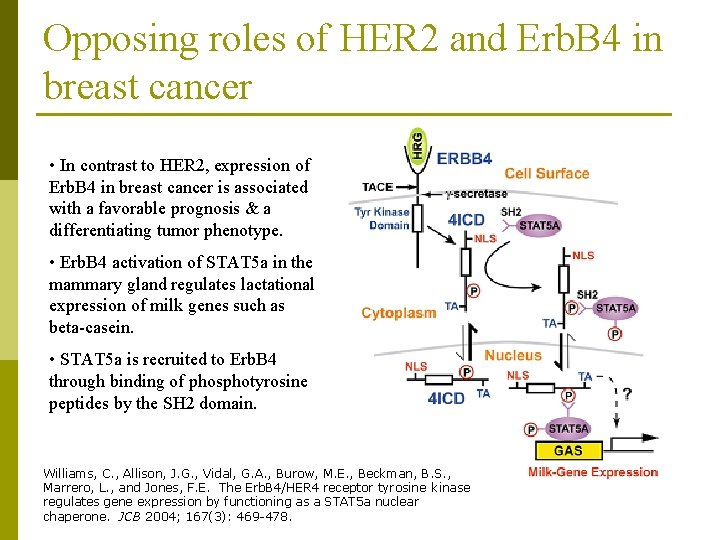
Opposing roles of HER 2 and Erb. B 4 in breast cancer • In contrast to HER 2, expression of Erb. B 4 in breast cancer is associated with a favorable prognosis & a differentiating tumor phenotype. • Erb. B 4 activation of STAT 5 a in the mammary gland regulates lactational expression of milk genes such as beta-casein. • STAT 5 a is recruited to Erb. B 4 through binding of phosphotyrosine peptides by the SH 2 domain. Williams, C. , Allison, J. G. , Vidal, G. A. , Burow, M. E. , Beckman, B. S. , Marrero, L. , and Jones, F. E. The Erb. B 4/HER 4 receptor tyrosine kinase regulates gene expression by functioning as a STAT 5 a nuclear chaperone. JCB 2004; 167(3): 469 -478.

PAINTing a picture of the interaction between Erb. B 4 and STAT 5 a • Goal is to connect Erb. B 4 to its regulated transcription factors, such as STAT 5 a, by applying a bioinformatics program called PAINT. • PAINT (Promoter Analysis & Interaction Network Toolset) is a computational tool which analyzes microarray data & generates networks connecting upregulated genes to their respective transcription factors. • Given a list of genes (microarray data), PAINT can: Fetch potential promoter sequences for the genes in the list. Find Transcription Factor (TF) binding sites on the sequences. Analyze the TF-binding site occurrences for over/under-representation compared to a reference. Vadigepalli, R, Chakravarthula, P, Zak DE, Schwaber JS, and Gonye, GE. PAINT: a promoter analysis and interaction network generation tool for gene regulatory network identification. OMICS 2003; 7(3): 235 -53.
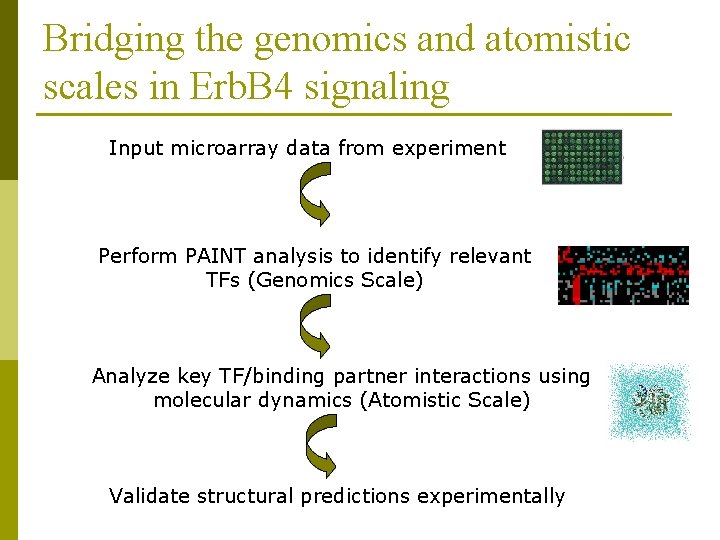
Bridging the genomics and atomistic scales in Erb. B 4 signaling Input microarray data from experiment Perform PAINT analysis to identify relevant TFs (Genomics Scale) Analyze key TF/binding partner interactions using molecular dynamics (Atomistic Scale) Validate structural predictions experimentally

Preliminary results: PAINT analysis • The PAINT method was applied to the following study, which focused on stimulation of Erb. B 4 in mammary epithelial cells: Amin, DN, Perkins AS, and Stern DF. Gene expression profiling of Erb. B receptor and ligand-dependent transcription. Oncogene 2004 Feb 19; 23(7): 1428 -38. • In the study, agonistic antibodies as well as natural ligands (neuregulin) were used to activate the Erb. B 4 pathway, and Erb. B 4 -stimulated gene expression was assessed by microarray analysis. • Several novel Erb. B 4 gene targets were identified and their associated transcription factors were predicted by PAINT.
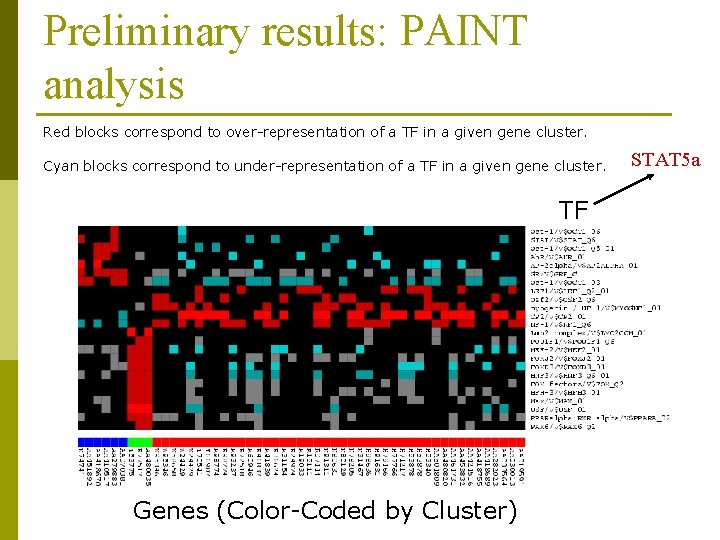
Preliminary results: PAINT analysis Red blocks correspond to over-representation of a TF in a given gene cluster. Cyan blocks correspond to under-representation of a TF in a given gene cluster. TF Genes (Color-Coded by Cluster) STAT 5 a

STAT 5 a interaction with Erb. B 4 IFN-γ phosphopeptide • The SH 2 domain of STAT 5 a binds to P-Y 959 at the C-terminal end of Erb. B 4’s kinase domain. • Structural details of the SH 2 domainphosphotyrosine peptide interaction are known (STAT 1 -IFN-γ crystal structure). • Can we predict features of the interaction between Erb. B 4 and STAT 5 a? STAT 1 SH 2 domain Mao, X. , et al. Structural bases of unphosphorylated STAT 1 association and receptor binding. Mol Cell 2005; 17(6): 761 -71. STAT 1 bound to phosphotyrosine peptide on IFN-γ receptor.
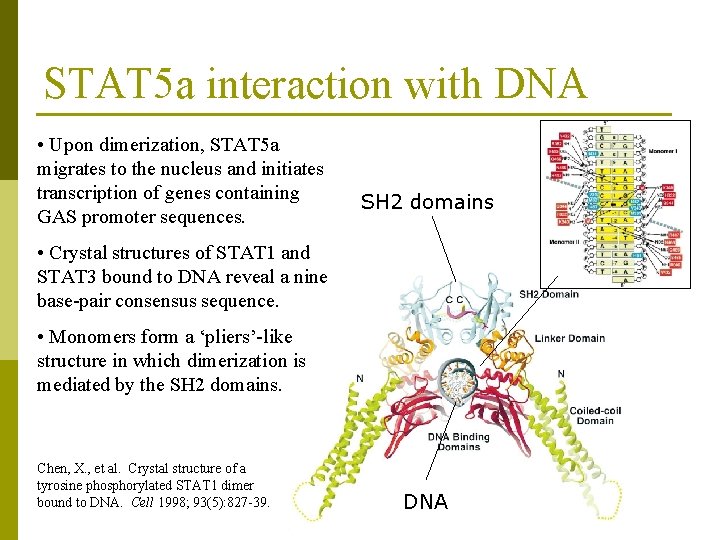
STAT 5 a interaction with DNA • Upon dimerization, STAT 5 a migrates to the nucleus and initiates transcription of genes containing GAS promoter sequences. SH 2 domains • Crystal structures of STAT 1 and STAT 3 bound to DNA reveal a nine base-pair consensus sequence. • Monomers form a ‘pliers’-like structure in which dimerization is mediated by the SH 2 domains. Chen, X. , et al. Crystal structure of a tyrosine phosphorylated STAT 1 dimer bound to DNA. Cell 1998; 93(5): 827 -39. DNA

Predicting STAT 5 a interaction with Erb. B 4 and DNA • To summarize, goals of Erb. B 4 study are two-fold: • Predict binding of Erb. B 4 to STAT 5 a SH 2 domain. Is it structurally possible for Erb. B 4 to phosphorylate the key tyrosine on STAT 5 a? Does Erb. B 4 bind to STAT 5 a as a monomer or as a dimer? • Assess STAT 5 a binding to DNA consensus sequence. Which interactions regulate specificity of nucleotide-binding? What mutations in the DNA sequence or STAT 5 a DNA-binding domain abolish the interaction? • Experimentally validate through mutagenesis studies and EMSA assays. • Elucidate structural features of key interactions involved in the Erb. B 4 signaling pathway.

Acknowledgments Ravi Radhakrishnan, Ph. D. Mark Lemmon, Ph. D. Rajanikanth Vadigepalli, Ph. D. Andrew Shih

References Bose, R, Molina, H, Patterson, AS, Bitok, JK, Periaswamy, B, Bader, JS, Pandey, A, and Cole, PA. Phosphoproteomic analysis of Her 2/neu signaling and inhibition. PNAS 2006; 103(26): 9773 -9778. Chen, X, Vinkemeier, U, Zhao, Y, Jeruzalmi, D, Darnell, JE Jr, and Kuriyan, J. Crystal structure of a tyrosine phosphorylated STAT 1 dimer bound to DNA. Cell 1998; 93(5): 827 -39. Fiser, A, Sali, A. Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol 2003; 374: 461 -91. Hubbard, SR, Wei, L, Ellis, L, and Hendrickson, W. A. Crystal structure of the tyrosine kinase domain of the human insulin receptor. Nature 1994; 372: 746 -54. Lemmon, MA. The EGF receptor family as therapeutic targets in breast cancer. Breast Dis 2003; 18: 33 -43. Linggi, B, Cheng, QC, Rao AR, and Carpenter G. The Erb. B 4 s 80 intracellular domain is a constitutively active tyrosine kinase. Oncogene 2006; 25: 160 -63. Liu Y, Purvis J, Shih A, Weinstein J, Agrawal N, Radhakrishnan R. A multiscale computational approach to dissect early events in the Erb family receptor mediated activation, differential signaling, and relevance to oncogenic transformations. Ann Biomed Eng 2007; 35(6): 1012 -25. Mao, X, Ren, Z, Parker, GN, Sondermann, H, Pastorello, MA, Wang, W, Mc. Murray, JS, Demeler, B, Darnell, JE Jr, and Chen, X. Structural bases of unphosphorylated STAT 1 association and receptor binding. Mol Cell 2005; 17(6): 761 -71. Schulze, WX, Deng, L, and Mann, M. Phosphotyrosine interactome of the Erb. B-receptor kinase family. Mol Systems Biology 2005; 1: 2005. 0008. Stamos, J, Sliwkowski, MX, and Eigenbrot, C. Structure of the epidermal growth factor receptor kinase domain alone and in complex with a 4 -anilinoquinazoline inhibitor. JBC 2002; 277(48): 46265 -46272.

References Stein, RA and Staros, JV. Insights into the evolution of the Erb. B receptor family and their ligands from sequence analysis. BMC Evol Biol 2006; Oct 6; 6: 79. Sundvall, M, Peri, L, Maatta, JA, Tvorogov, D, Paatero, I, Savisalo, M, Silvennoinen, O, Yarden, Y and Elenius K. Differential nuclear localization and kinase activity of alternative Erb. B 4 intracellular domains. Oncogene 2007; 26(48): 6905 -14. Vadigepalli, R, Chakravarthula, P, Zak DE, Schwaber JS, and Gonye, GE. PAINT: a promoter analysis and interaction network generation tool for gene regulatory network identification. OMICS 2003; 7(3): 235 -53. Wang, SE, Narasanna, A, Perez-Torres, M, Xiang, B, Wu, FY, Yang, S, Carpenter, G, Gazdar AF, Muthuswamy, SK, and Arteaga, CL. HER 2 kinase domain mutation results in constitutive phosphorylation and activation of HER 2 and EGFR and resistance to EGFR tyrosine kinase inhibitors. Cancer Cell 2006; 10: 25 -38. Williams, C. , Allison, JG, Vidal, GA, Burow, ME, Beckman, BS, Marrero, L. , and Jones, FE. The Erb. B 4/HER 4 receptor tyrosine kinase regulates gene expression by functioning as a STAT 5 a nuclear chaperone. JCB 2004; 167(3): 469 -478. Wood ER, Truesdale AT, Mc. Donald OB, Yuan D, Hassell A, Dickerson SH, Ellis B, Pennisi C, Horne E, Lackey K, Alligood KJ, Rusnak DW, Gilmer TM, and Shewchuk L. A unique structure for epidermal growth factor receptor bound to GW 572016 (Lapatinib): relationships among protein conformation, inhibitor off-rate, and receptor activity in tumor cells. Cancer Res 2004; 64(18): 6652 -9. Yarden, Y and Sliwkowski, MX. Untangling the Erb. B signalling network. Nat Rev Molecular Cell Biology 2001; 2: 127 -137. Zhang, X. , Gureasko, J. , Shen, K. , Cole, P. , and Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 2006; 125: 1137 -1149.
- Slides: 34