Structural Bioinformatics in Drug Discovery Melissa Passino Structural

Structural Bioinformatics in Drug Discovery Melissa Passino

Structural Bioinformatics • What is SBI? “Structural bioinformatics is a subset of bioinformatics concerned with the use of biological structures – proteins, DNA, RNA, ligands etc. and complexes thereof to further our understanding of biological systems. ” http: //biology. sdsc. edu/strucb. html
SBI in Drug Design and Discovery • SBI can be used to examine: • drug targets (usually proteins) • binding of ligands ↓ “rational” drug design (benefits = saved time and $$$)

Traditional Methods of Drug Discovery natural (plant-derived) treatment for illness/ailments ↓ isolation of active compound (small, organic)
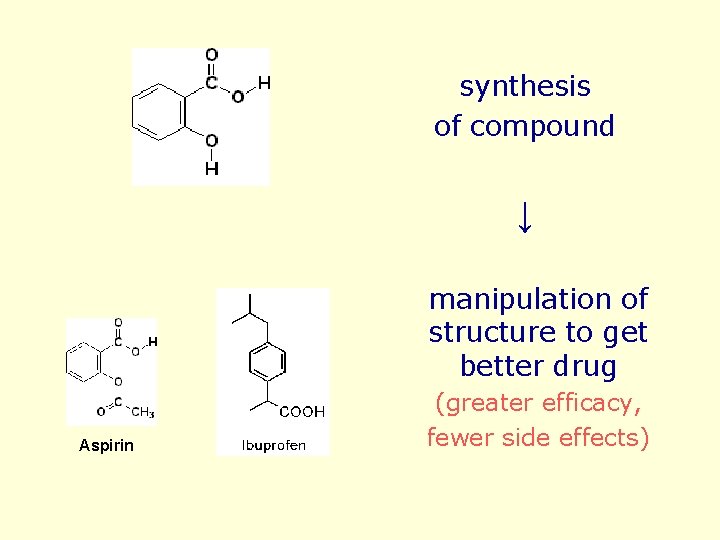
synthesis of compound ↓ manipulation of structure to get better drug Aspirin (greater efficacy, fewer side effects)

Modern Methods of Drug Discovery What’s different? • Drug discovery process begins with a disease (rather than a treatment) • Use disease model to pinpoint relevant genetic/biological components (i. e. possible drug targets)
Modern Drug Discovery disease → genetic/biological target ↓ discovery of a “lead” molecule - design assay to measure function of target - use assay to look for modulators of target’s function ↓ high throughput screen (HTS) - to identify “hits” (compounds with binding in low n. M to low μM range)

Modern Drug Discovery small molecule hits ↓ manipulate structure to increase potency i. e. decrease Ki to low n. M affinity ↓ *optimization of lead molecule into candidate drug* fulfillment of required pharmacological properties: potency, absorption, bioavailability, metabolism, safety ↓ clinical trials

Interesting facts. . . • Over 90% of drugs entering clinical trials fail to make it to market • The average cost to bring a new drug to market is estimated at $770 million
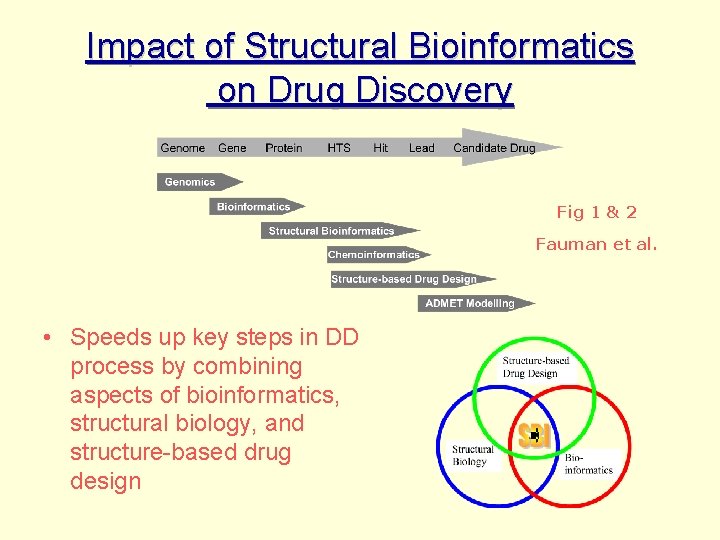
Impact of Structural Bioinformatics on Drug Discovery Fig 1 & 2 Fauman et al. • Speeds up key steps in DD process by combining aspects of bioinformatics, structural biology, and structure-based drug design

Identifying Targets: The “Druggable Genome”

human genome polysaccharides lipids nucleic acids proteins Problems with toxicity, specificity, and difficulty in creating potent inhibitors eliminate the first 3 categories. . .

human genome polysaccharides lipids nucleic acids proteins with binding site “druggable genome” = subset of genes which express proteins capable of binding small drug-like molecules
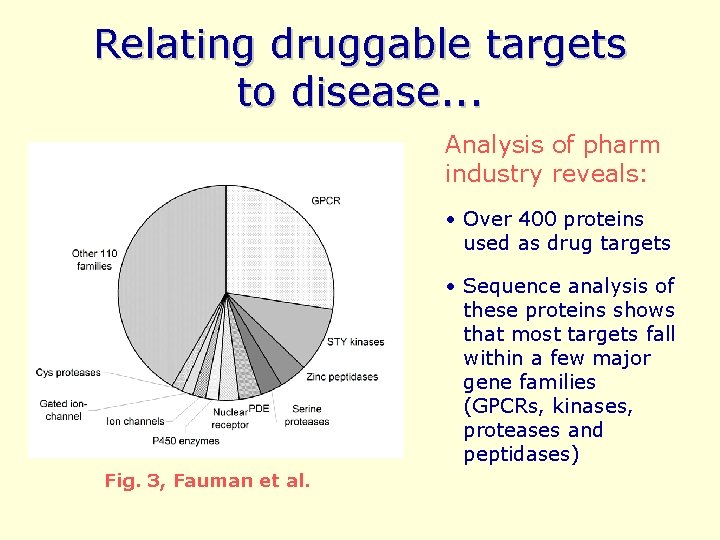
Relating druggable targets to disease. . . Analysis of pharm industry reveals: • Over 400 proteins used as drug targets • Sequence analysis of these proteins shows that most targets fall within a few major gene families (GPCRs, kinases, proteases and peptidases) Fig. 3, Fauman et al.

Assessing Target Druggability • Once a target is defined for your disease of interest, SBI can help answer the question: Is this a “druggable” target? • Does it have sequence/domains similar to known targets? • Does the target have a site where a drug can bind, and with appropriate affinity?
Other roles for SBI in drug discovery • Binding pocket modeling • Lead identification • Similarity with known proteins or ligands • Chemical library design / combinatorial chemistry • Virtual screening • *Lead optimization* • Binding • ADMET

SBI in cancer therapy: MMPIs

• Inability to control metastasis is the leading cause of death in patients with cancer (Zucker et al. Oncogene. 2000, 19, 6642 -6650. ) • Matrix metalloproteinase inhibitors (MMPIs) are a newer class of cancer therapeutics • can prevent metastasis (but not cytotoxic); may also play role in blocking tumor angiogenesis (growth inhibition) • Used to treat “major” cancers: lung, GI, prostate

What is an MMP? • Family of over 20 structurally related proteinases • Principal substrates: • protein components of extracellular matrix (collagen, fibronectin, laminin, proteoglycan core protein) • Functions: • Breakdown of connective tissue; tissue remodeling • Role in cancer: • Increased levels/activity of MMPs in area surrounding tumor
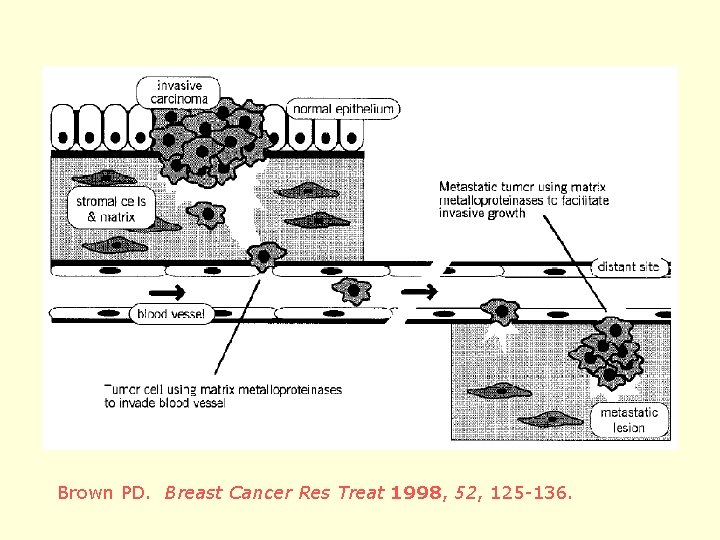
Brown PD. Breast Cancer Res Treat 1998, 52, 125 -136.

Whittaker et al. Chem. Rev. 1999, 2735 -2776
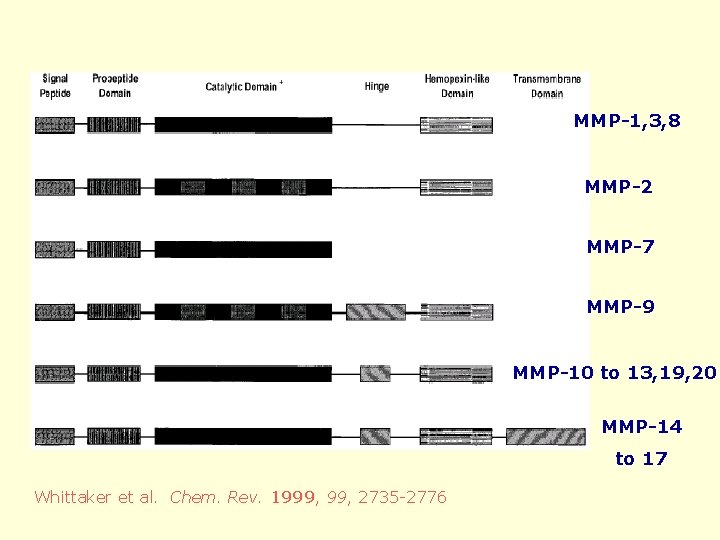
MMP-1, 3, 8 MMP-2 MMP-7 MMP-9 MMP-10 to 13, 19, 20 MMP-14 to 17 Whittaker et al. Chem. Rev. 1999, 2735 -2776
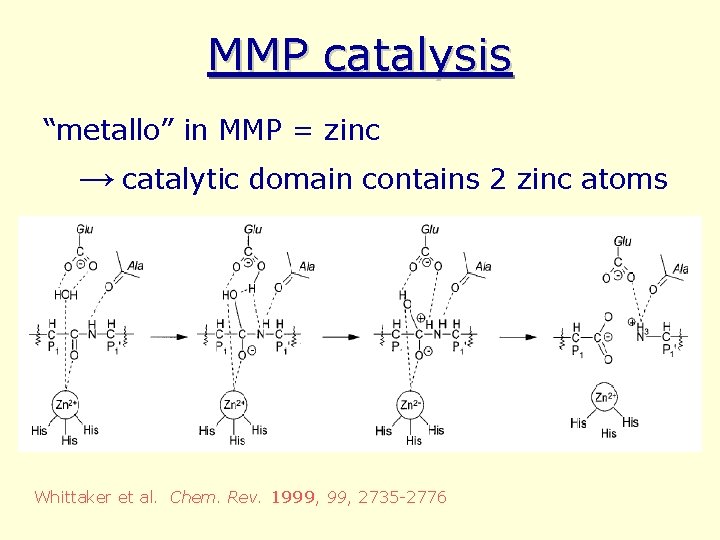
MMP catalysis “metallo” in MMP = zinc → catalytic domain contains 2 zinc atoms Whittaker et al. Chem. Rev. 1999, 2735 -2776
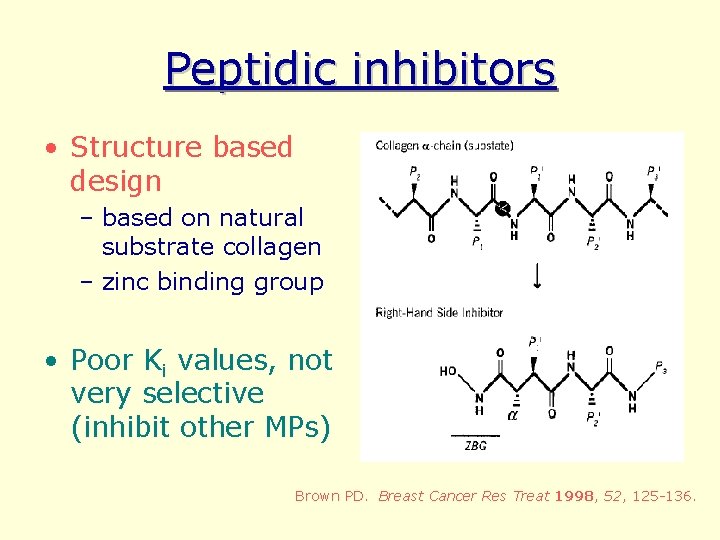
Peptidic inhibitors • Structure based design – based on natural substrate collagen – zinc binding group • Poor Ki values, not very selective (inhibit other MPs) Brown PD. Breast Cancer Res Treat 1998, 52, 125 -136.
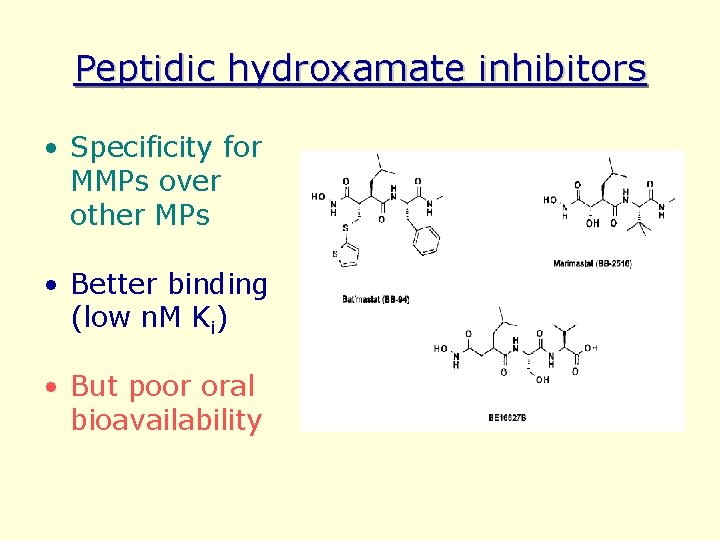
Peptidic hydroxamate inhibitors • Specificity for MMPs over other MPs • Better binding (low n. M Ki) • But poor oral bioavailability

A (not very) long time ago, in a town (not too) far away… …lived a company named Agouron… …and this company had a dream, a dream to design a nonpeptidic hydroxamate inhibitor of MMPs…

. . . so they made some special crystals… used x-ray crystallography/3 D structure of recombinant human MMPs bound to various inhibitors ↓ to determine key a. a. residues, ligand substituents needed for binding Gelatinase A http: //www. rcsb. org/pdb/
…and used the magic of structural bioinformatics to design many, many nonpeptidic hydroxylates. Ki oral bioavailabity antigrowth repeat… antimetastasis

Results… AG 3340 “Prinomastat” • Good oral bioavailability • Selective for specific MMPs – may implicate their roles in certain cancers

Prinomastat • Evidence showing prevention of lung cancer metastasis in rat and mice models • Clinical trials → non small cell lung cancer → hormone refractory prostate cancer …stopped at Phase 3 (Aug 2000) because did not show effects against late stage metastasis

Morals of the story… • SBI can be used as basis for lead discovery and optimization • MMPs are good targets for chemotherapy to help control metastasis… …but MMPIs must be combined with other cytotoxic drugs to get maximum benefits, and used at earliest stage possible
- Slides: 31