Stochasticity in Signaling Pathways and Gene Regulation The














- Slides: 37

Stochasticity in Signaling Pathways and Gene Regulation: The NFκB Example and the Principle of Stochastic Robustness Marek Kimmel Rice University, Houston, TX, USA

Credits • Rice University – Pawel Paszek – Roberto Bertolusso • UTMB – Galveston – Allan Brasier – Bing Tian • Politechnika Slaska – Jaroslaw Smieja – Krzysztof Fujarewicz • Baylor College of Medicine – Michael Mancini – Adam Szafran – Elizabeth Jones • IPPT – Warsaw – Tomasz Lipniacki – Beata Hat

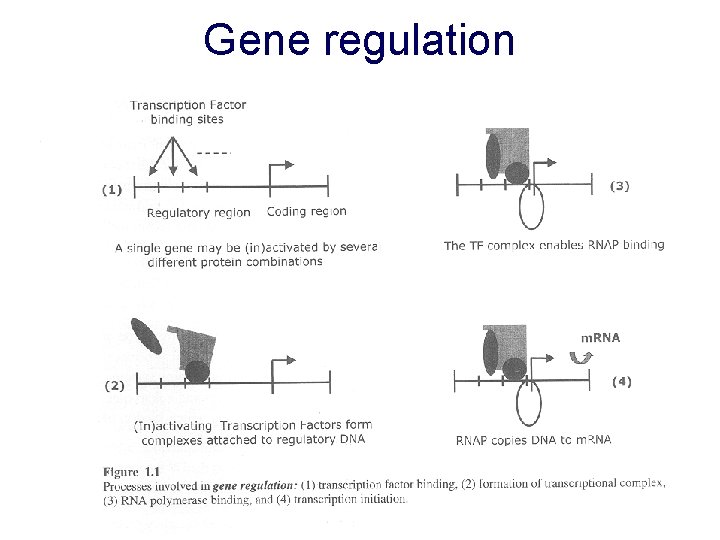
Gene regulation

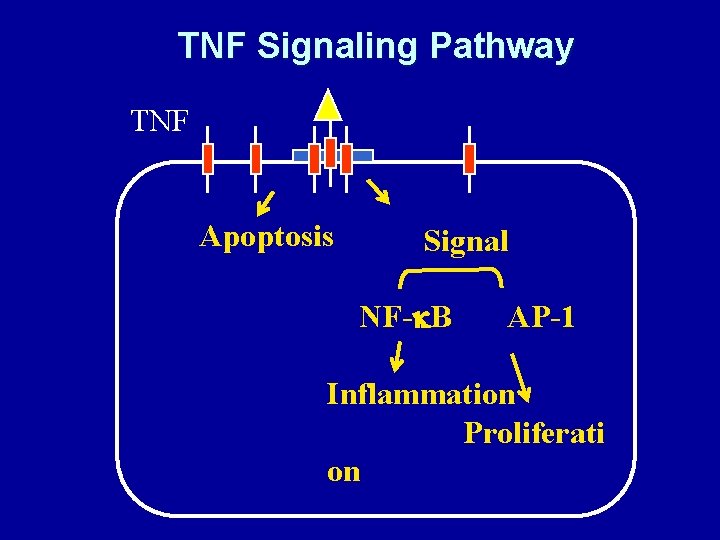
TNF Signaling Pathway TNF Apoptosis Signal NF-k. B AP-1 Inflammation Proliferati on

Nuclear Factor- B (NF- B) • Inducible (cytoplasmic) transcription factor • Mediator of acute phase reactant transcription (angiotensinogen, SAA) • Mediator of cytokine and chemokine expression in pulmonary cytokine cascade • Plays role in anti-apoptosis and confering chemotherapy resistance in drug resistant cancers

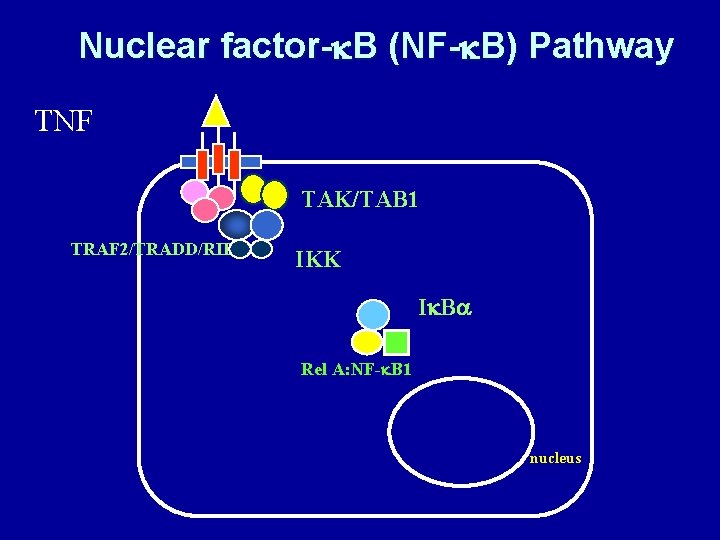
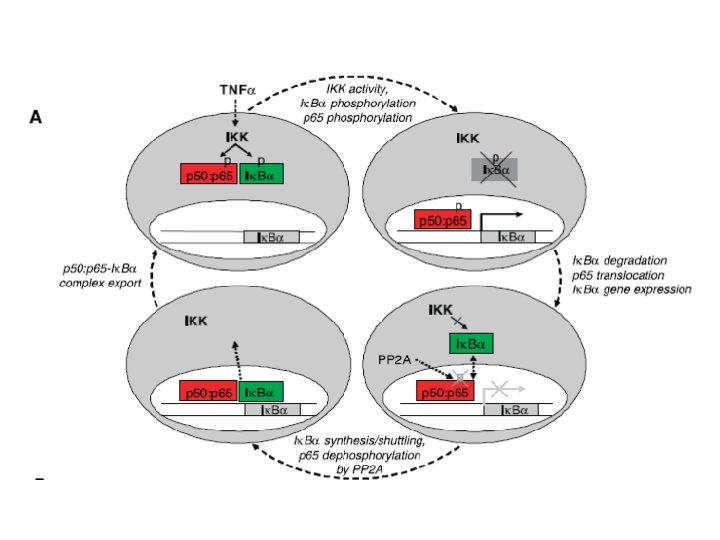
Nuclear factor-k. B (NF-k. B) Pathway TNF TAK/TAB 1 TRAF 2/TRADD/RIP IKK Ik. Ba Rel A: NF-k. B 1 nucleus

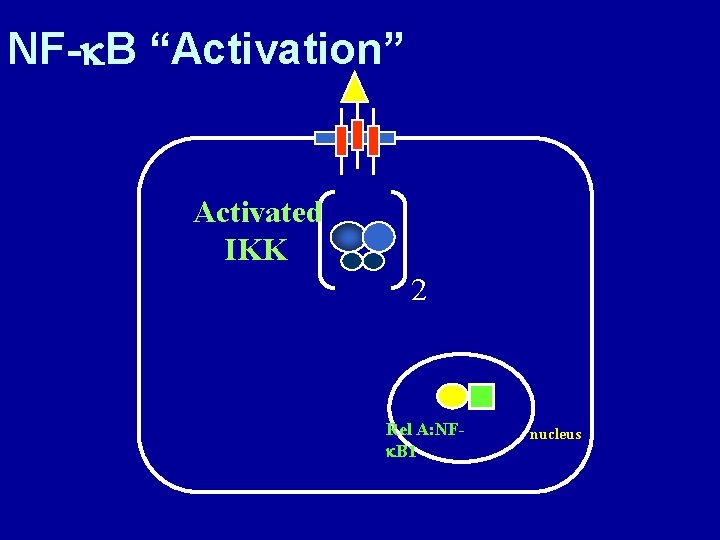
NF-k. B “Activation” Activated IKK 2 Rel A: NFk. B 1 nucleus

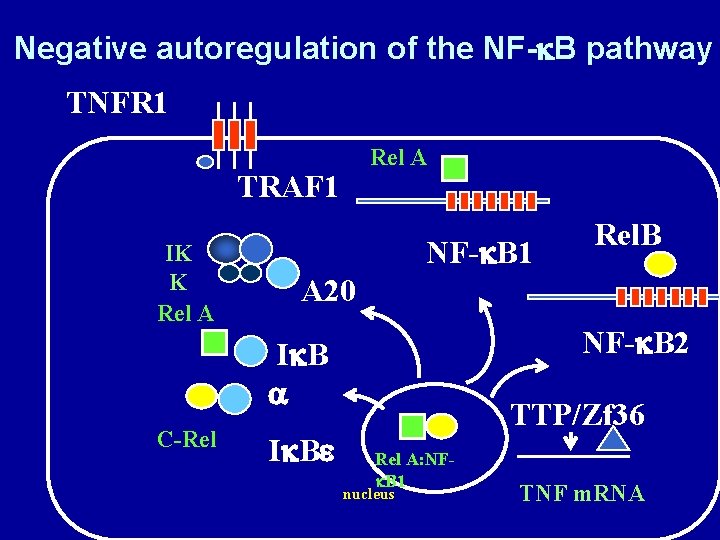
Negative autoregulation of the NF-k. B pathway TNFR 1 Rel A TRAF 1 IK K Rel A NF-k. B 1 A 20 NF-k. B 2 Ik. B a C-Rel Rel. B Ik. Be TTP/Zf 36 Rel A: NFk. B 1 nucleus TNF m. RNA

Intrinsic sources of stochasticity • In bacteria, single-cell level stochasticity is quite well-recognized, since the number of m. RNA or even protein of given type, per cell, might be small (1 gene, several m. RNA, protein ~10) • Eukaryotic cells are much larger (1 -2 genes, m. RNA ~100, protein ~100, 000), so the source of stochasticity is mainly the regulation of gene activity.

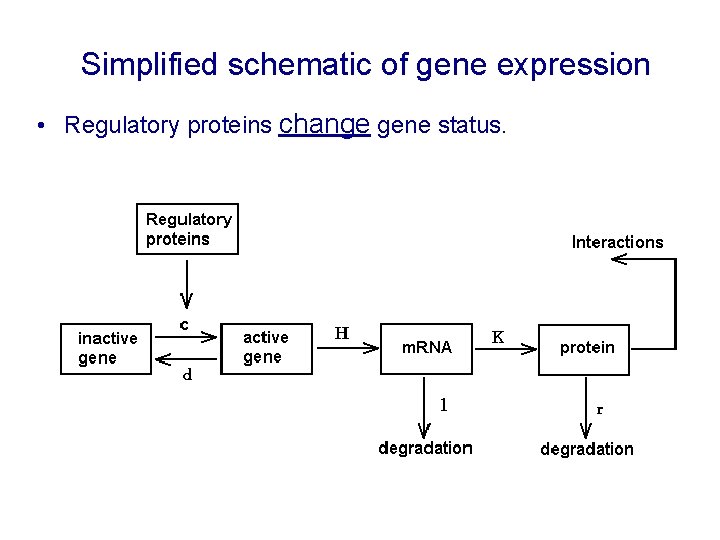
Simplified schematic of gene expression • Regulatory proteins change gene status.

Discrete Stochastic Model Time-continuous Markov chain with state space and transition intensities

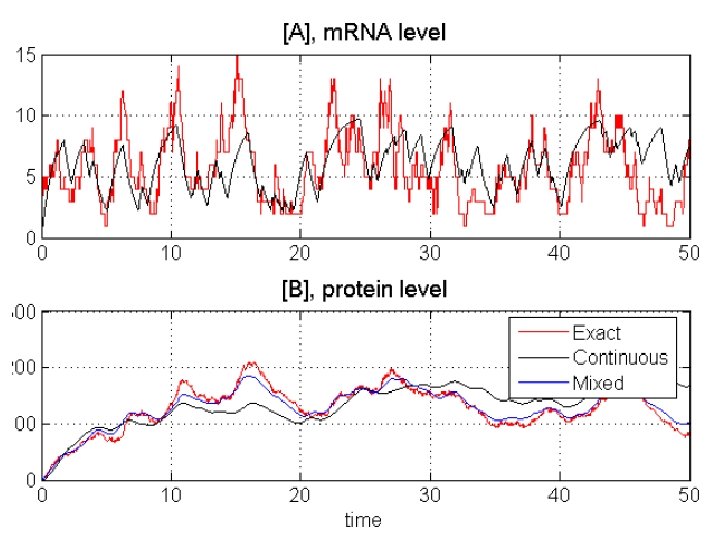
Continuous Approximation only gene on/off discrete stochastic


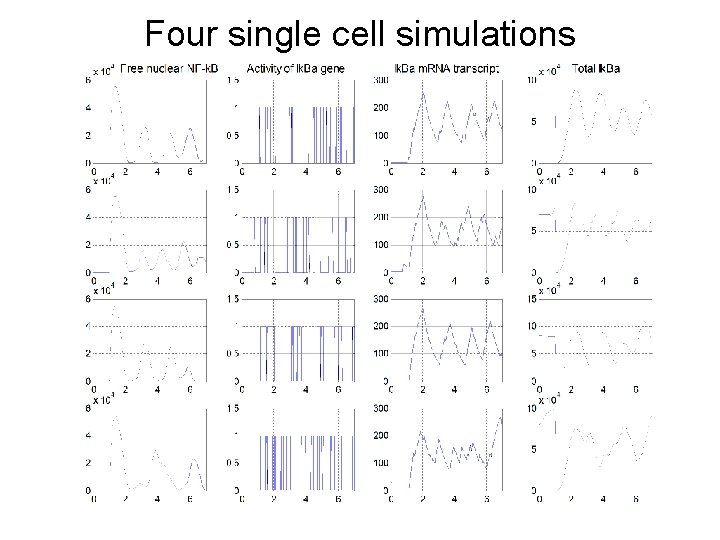
Four single cell simulations

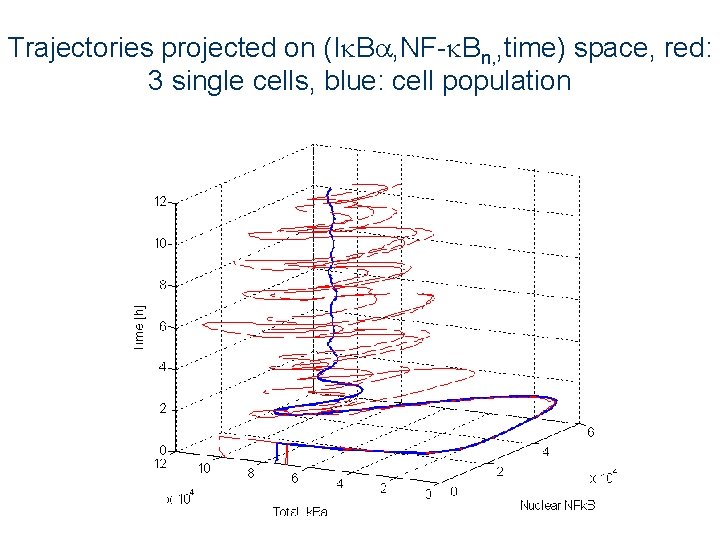
Trajectories projected on (I B , NF- Bn, , time) space, red: 3 single cells, blue: cell population Any single cell trajectory differs from the “averaged” trajectory




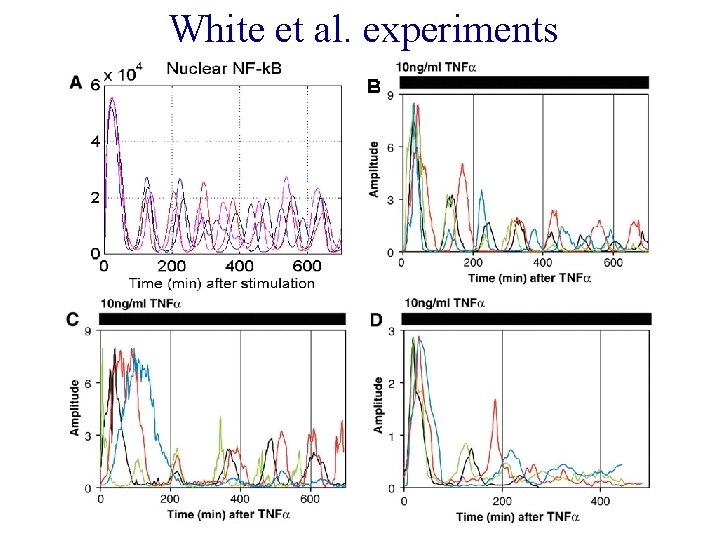
White et al. experiments

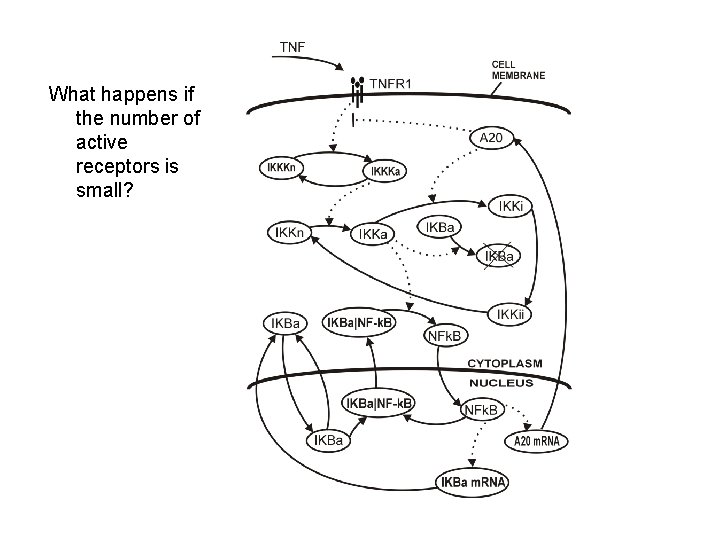
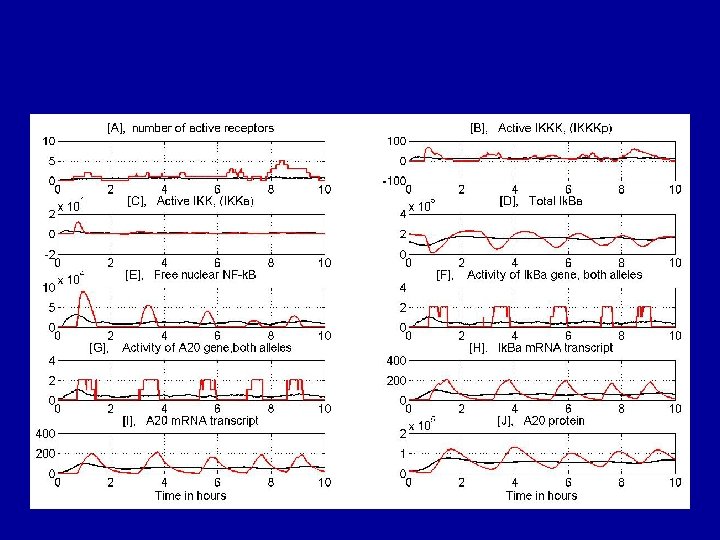
What happens if the number of active receptors is small?

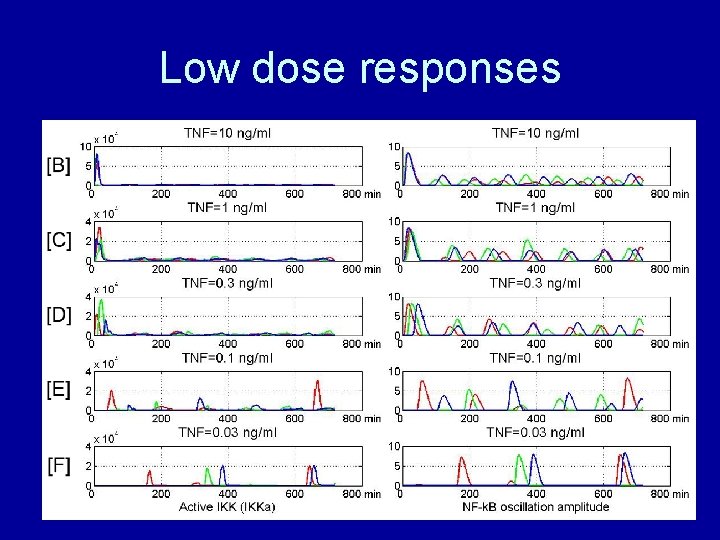
Low dose responses


How to find out if on/off transcrition stochasticity plays a role? • If on/off rapid enough, its influence on the system is damped • Recent photobleaching experiments → TF turnover ~10 sec • However, does this quick turnover reflect duration of transcription “bursts”?

FRAP (Mancini Lab) Fluorescence recovery after photobleaching

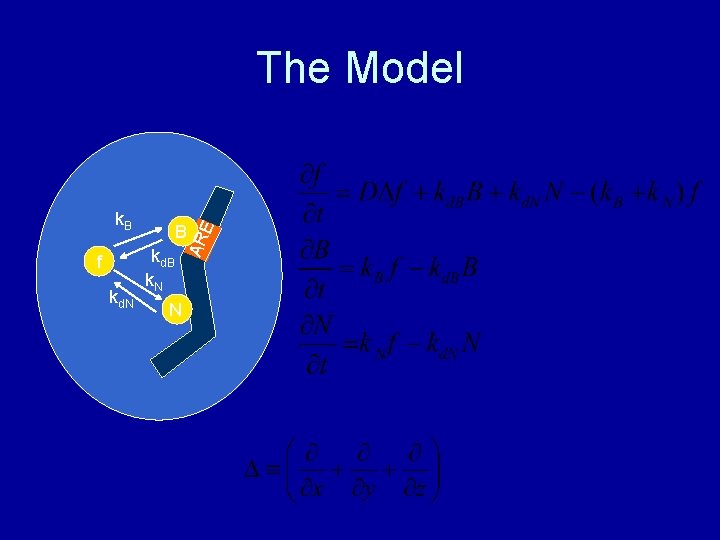
k. B kd. N kd. B k. N N AR f B E The Model

The Model • Fit the model to photobleaching data • Obtain estimates of binding constants of the factor • Invert binding constants to obtain mean residence times • Effect: ~10 seconds

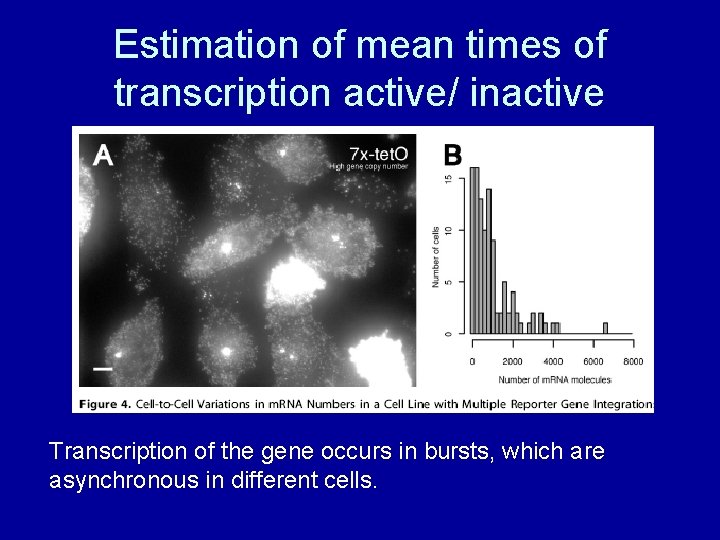
Estimation of mean times of transcription active/ inactive

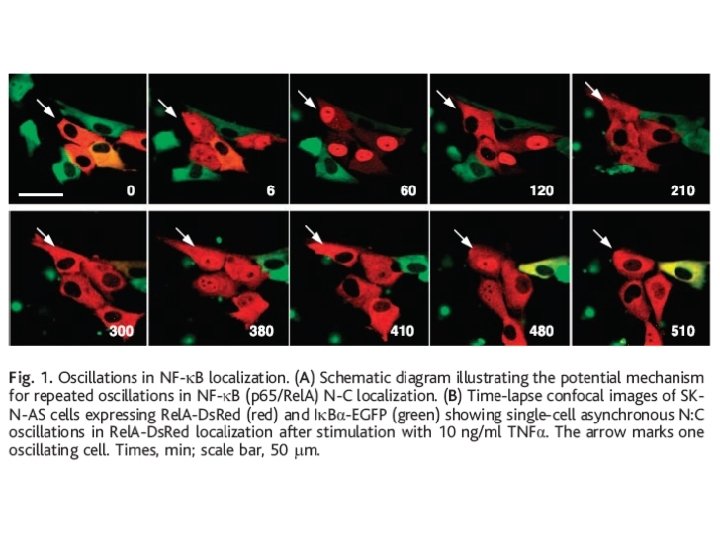
Estimation of mean times of transcription active/ inactive Transcription of the gene occurs in bursts, which are asynchronous in different cells.

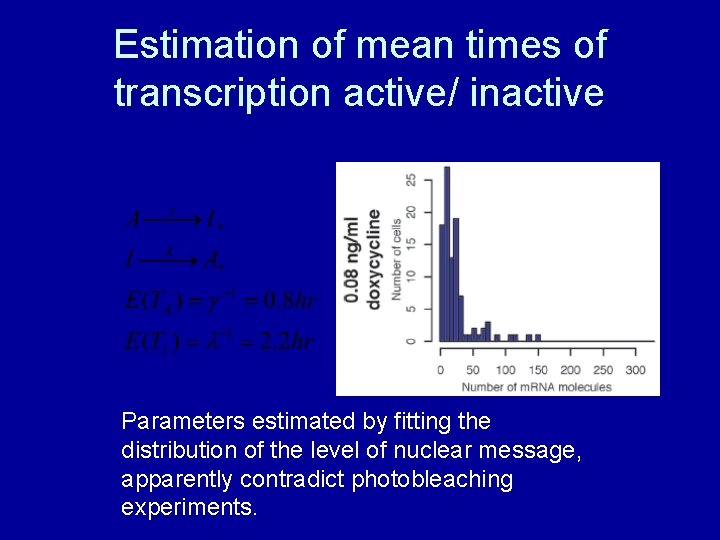
Estimation of mean times of transcription active/ inactive Parameters estimated by fitting the distribution of the level of nuclear message, apparently contradict photobleaching experiments.

A single gene (one copy) using K-E approximation Amount of protein: Where: • and are the constitutive activation and deactivation rates, respectively, • is an inducible activation rate due to the action of protein dimers.

Deterministic description The system has one or two stable equilibrium points depending on the parameters.

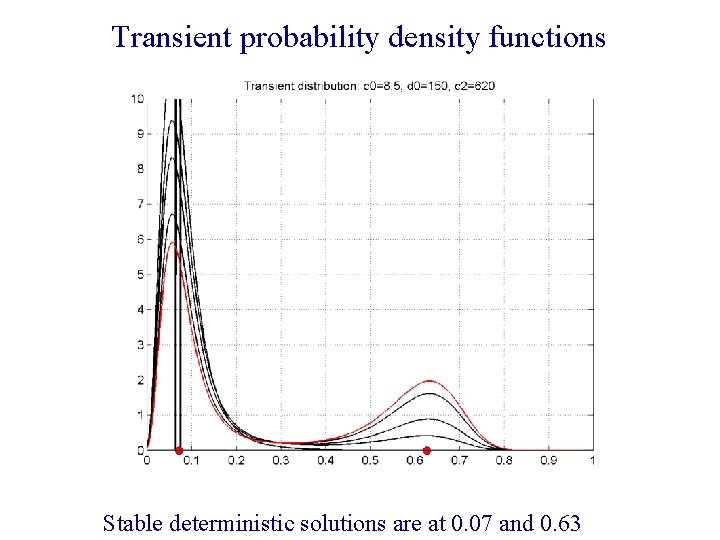
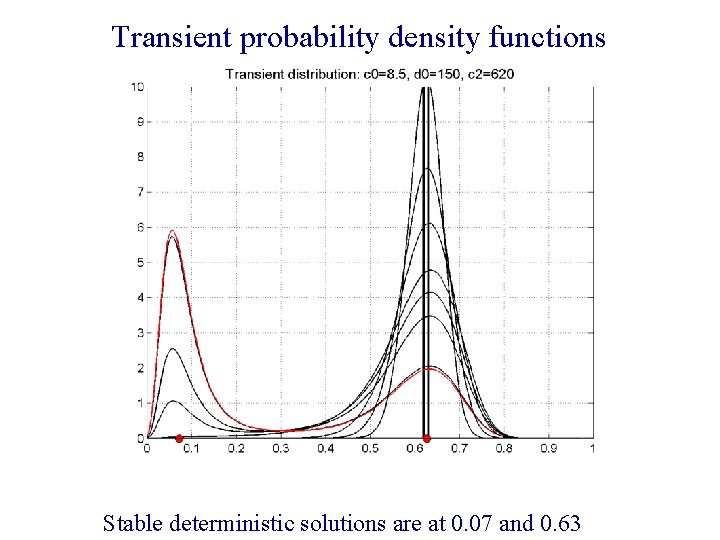
Transient probability density functions Stable deterministic solutions are at 0. 07 and 0. 63

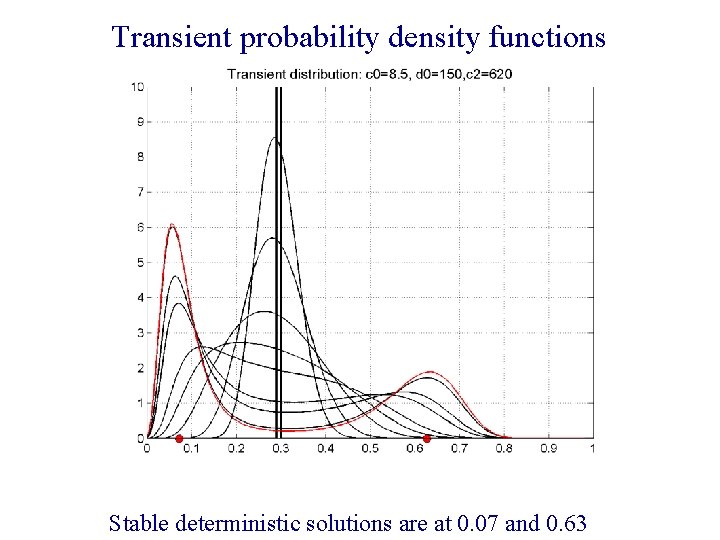
Transient probability density functions Stable deterministic solutions are at 0. 07 and 0. 63

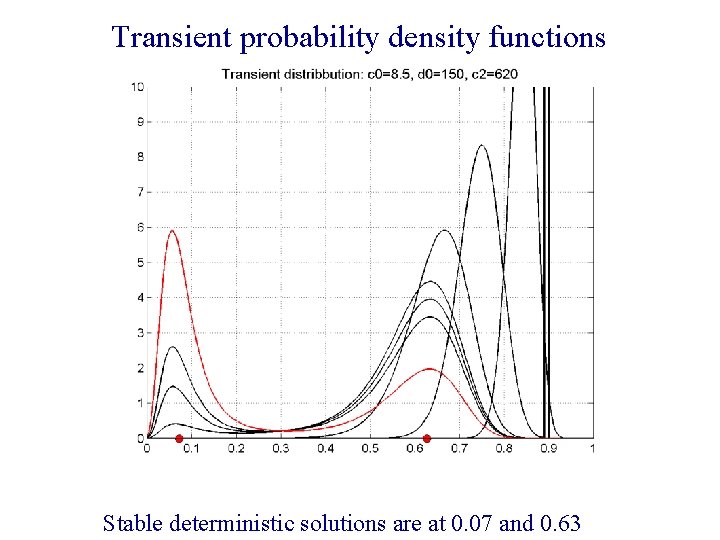
Transient probability density functions Stable deterministic solutions are at 0. 07 and 0. 63

Transient probability density functions Stable deterministic solutions are at 0. 07 and 0. 63

Conclusions from modeling • Stochastic event of gene activation results in a burst of m. RNA molecules, each serving as a template for numerous protein molecules. • No single cell behaves like an average cell. • Decreasing magnitude of the signal below a threshold value lowers the probability of response but not its amplitude. • “Stochastic robustness” allows individual cells to respond differently to the same stimulus, but makes responses well-defined (proliferation vs. apoptopsis).

References • Lipniacki T, Paszek P, Brasier AR, Luxon BA, Kimmel M. Stochastic regulation in early immune response. Biophys J. 2006 Feb 1; 90(3): 725 -42. • Paszek P, Lipniacki T, Brasier AR, Tian B, Nowak DE, Kimmel M. Stochastic effects of multiple regulators on expression profiles in eukaryotes. J Theor Biol. 2005 Apr 7; 233(3): 423 -33. • Lipniacki T, Paszek P, Brasier AR, Luxon B, Kimmel M. Mathematical model of NF-kappa. B regulatory module. J Theor Biol. 2004 May 21; 228(2): 195 -215.