Statistical Thermodynamics Lecture 20 Binding Free Energy Calculations

Statistical Thermodynamics Lecture 20: Binding Free Energy Calculations Dr. Ronald M. Levy ronlevy@temple. edu

Outline focus on the binding problem of protein-ligand systems. • Statistical theory of non-covalent binding equilibria • Computer models for computing free energy of binding • Binding energy distribution analysis method • Examples from our research group Center for Biophysics and Computational Biology, https: //cb 2. cst. temple. edu/

Statistical Theory of Non-Covalent Binding Equilibria General chemical reaction: Very important type of “reaction”: bimolecular non-covalent binding R(sol) + L(sol) RL(sol) Small molecule dimerization/association Supramolecular complexes Protein-ligand binding Protein-protein binding/dimerization Protein-nucleic acids interactions. . . *Note*: We are implicitly assuming above that we can describe the system as being composed of 3 distinct chemical “species”, R, L, and RL (quasichemical description). If interactions between R and L are weak/non-specific then it would be more appropriate to treat the system as a non-ideal solution of R and L.

From general theory of chemical reactions, for receptor-ligand system: Effective potential energy of solute i in solution.

If the solution is isotropic ( invariant upon rotation of solute), integrate analytically over rotational degrees of freedom (ignoring rotovibarational couplings, OK at physiological temperatures). Internal coordinates When inserting into the expression for Kb(T), Λ's cancel because We get: [Gilson et al. Biophysical Journal 72, 1047 -1069 (1997)]
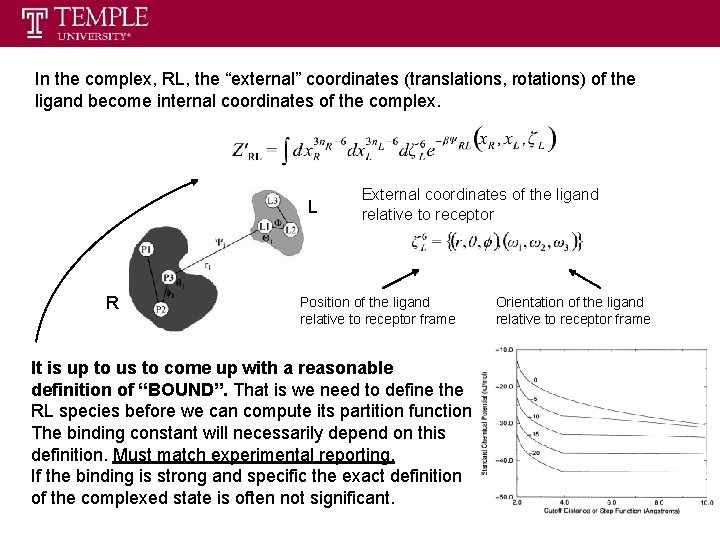
In the complex, RL, the “external” coordinates (translations, rotations) of the ligand become internal coordinates of the complex. L R External coordinates of the ligand relative to receptor Position of the ligand relative to receptor frame It is up to us to come up with a reasonable definition of “BOUND”. That is we need to define the RL species before we can compute its partition function. The binding constant will necessarily depend on this definition. Must match experimental reporting. If the binding is strong and specific the exact definition of the complexed state is often not significant. Orientation of the ligand relative to receptor frame

It is convenient to introduce an “indicator” function for the complex: then: Next, define “binding energy” of a conformation of the complex: basically, change in effective energy for bringing ligand receptor together at fixed internal conformation: +

In terms of binding energy: then: Now: we are not very good at computing partition functions. We are much better at computing ensemble averages:

To transform the expression for Kb so that it looks like an average: multiply and divide by: then: or:

Summary of Binding Free Energy Theory Binding energy: Binding Constant: We can see that binding constant can be expressed in terms of an average of the exponential of the binding energy over the ensemble of conformations of the complex in which the ligand the receptor are not interacting while the ligand is placed in the binding site. Standard free energy of binding: (analytic formulas) (numerical computation)
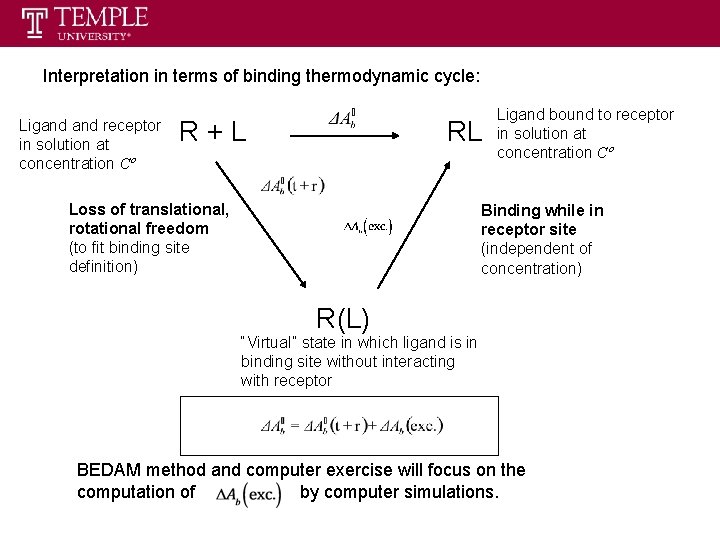
Interpretation in terms of binding thermodynamic cycle: Ligand receptor in solution at concentration Cº R+L RL Loss of translational, rotational freedom (to fit binding site definition) Ligand bound to receptor in solution at concentration Cº Binding while in receptor site (independent of concentration) R(L) “Virtual” state in which ligand is in binding site without interacting with receptor BEDAM method and computer exercise will focus on the computation of by computer simulations.
![Binding Free Energy Models [Gallicchio and Levy, Adv. Prot. Chem (2012)] Double Decoupling Method Binding Free Energy Models [Gallicchio and Levy, Adv. Prot. Chem (2012)] Double Decoupling Method](http://slidetodoc.com/presentation_image_h/20489e42676c1d89cb98748f611ee376/image-12.jpg)
Binding Free Energy Models [Gallicchio and Levy, Adv. Prot. Chem (2012)] Double Decoupling Method (DDM) Relative Binding Free Energies (FEP) λ-dynamics Potential of Mean Force/ Pathway Methods Docking & Scoring Statistical mechanics theory BEDAM (Implicit solvation) Exhaustive docking MM/PBSA Binding Energy Distribution Analysis Method Mining Minima (M 2)

Free Energy Perturbation and Double Decoupling Methods Statistical mechanics based, in principle account for: • Total binding free energy • Entropic costs • Ligand/receptor reorganization Free Energy Perturbation (FEP/TI) Jorgensen, Kollman, Mc. Cammon (1980’s – present) Double Decoupling (DDM) Jorgensen, Gilson, Roux, . . . (2000’s – to present) : Challenges: • Dissimilar ligand sets • Numerical instability • Dependence on starting conformations • Multiple bound poses • Slow convergence
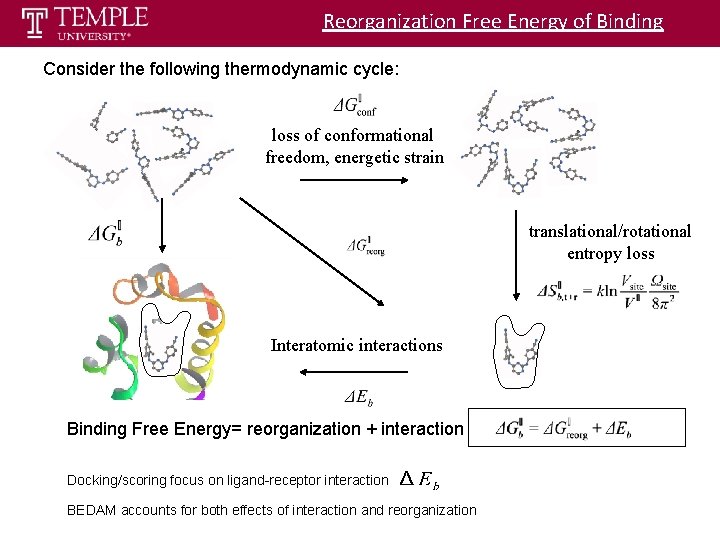
Reorganization Free Energy of Binding Consider the following thermodynamic cycle: loss of conformational freedom, energetic strain translational/rotational entropy loss Interatomic interactions Binding Free Energy= reorganization + interaction Docking/scoring focus on ligand-receptor interaction BEDAM accounts for both effects of interaction and reorganization
![Binding Energy Distribution Analysis Method (Statistical Theory) [Gilson, Mc. Cammon et al. , (1997)] Binding Energy Distribution Analysis Method (Statistical Theory) [Gilson, Mc. Cammon et al. , (1997)]](http://slidetodoc.com/presentation_image_h/20489e42676c1d89cb98748f611ee376/image-15.jpg)
Binding Energy Distribution Analysis Method (Statistical Theory) [Gilson, Mc. Cammon et al. , (1997)] Binding “energy” of a fixed conformation of the complex. W(): solvent PMF (implicit solvation model) Ligand in binding site in absence of ligand-receptor interactions Entropically favored
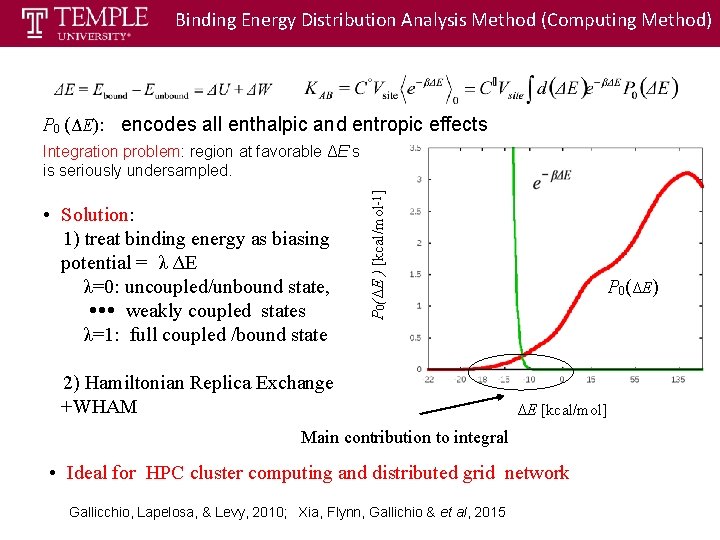
Binding Energy Distribution Analysis Method (Computing Method) P 0 (ΔE): encodes all enthalpic and entropic effects • Solution: 1) treat binding energy as biasing potential = λ ΔE λ=0: uncoupled/unbound state, weakly coupled states λ=1: full coupled /bound state P 0(ΔE ) [kcal/mol-1] Integration problem: region at favorable ΔE’s is seriously undersampled. 2) Hamiltonian Replica Exchange +WHAM P 0(ΔE) ΔE [kcal/mol] Main contribution to integral • Ideal for HPC cluster computing and distributed grid network ] Gallicchio, Lapelosa, & Levy, 2010; Xia, Flynn, Gallichio & et al, 2015
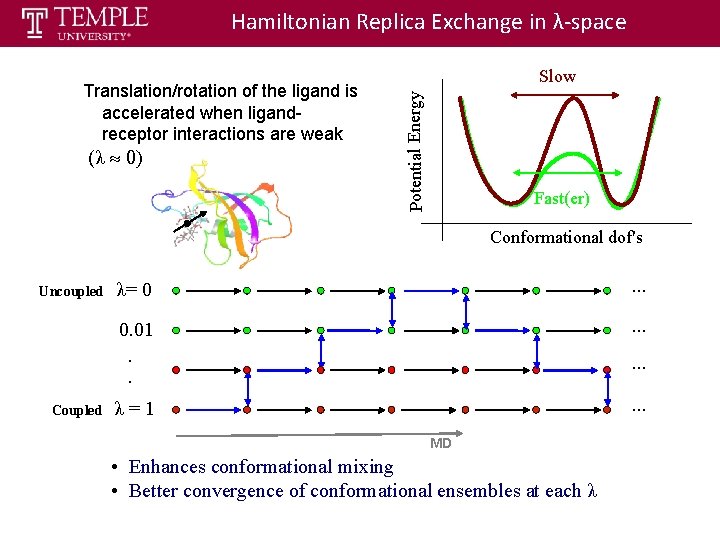
Hamiltonian Replica Exchange in λ-space (λ » 0) Potential Energy Translation/rotation of the ligand is accelerated when ligandreceptor interactions are weak Slow Fast(er) Conformational dof's Uncoupled Coupled λ= 0 . . . 0. 01 . . . . λ=1 . . . MD • Enhances conformational mixing • Better convergence of conformational ensembles at each λ

Reweighting Techniques in Free Energy Calculations Reweighting techniques are necessary to recover the unbiased true observables (results) from more advanced sampling methods: • Weighted Histogram Analysis Method (WHAM) Ferrenberg & Swendsen (1989) Kumar, Kollman et al. (1992) Bartels & Karplus (1997) Gallicchio, Levy et al. (2005) • Unbinned Weighted Histogram Analysis Method (UWHAM) equivalent to Multistate Bennett Acceptance Ratio (MBAR) Shirts & Chodera J. Chem. Phys. (2008). Tan, Gallicchio, Lapelosa, Levy JCTC (2012).
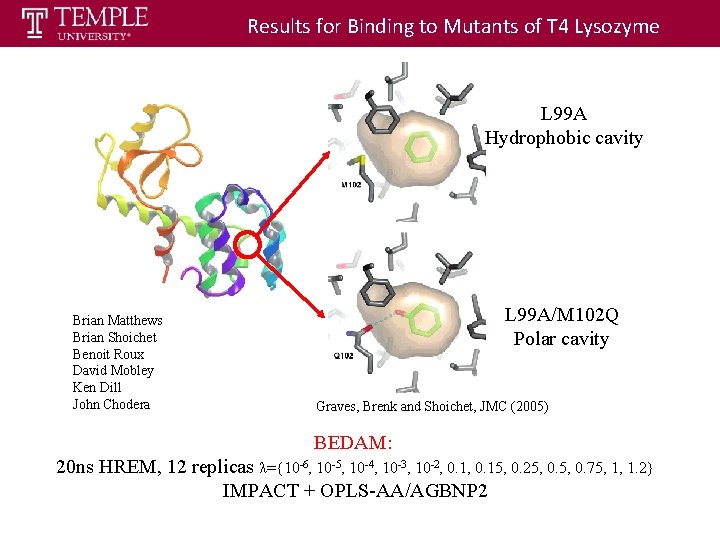
Results for Binding to Mutants of T 4 Lysozyme L 99 A Hydrophobic cavity Brian Matthews Brian Shoichet Benoit Roux David Mobley Ken Dill John Chodera L 99 A/M 102 Q Polar cavity Graves, Brenk and Shoichet, JMC (2005) BEDAM: 20 ns HREM, 12 replicas λ={10 -6, 10 -5, 10 -4, 10 -3, 10 -2, 0. 15, 0. 25, 0. 75, 1, 1. 2} IMPACT + OPLS-AA/AGBNP 2
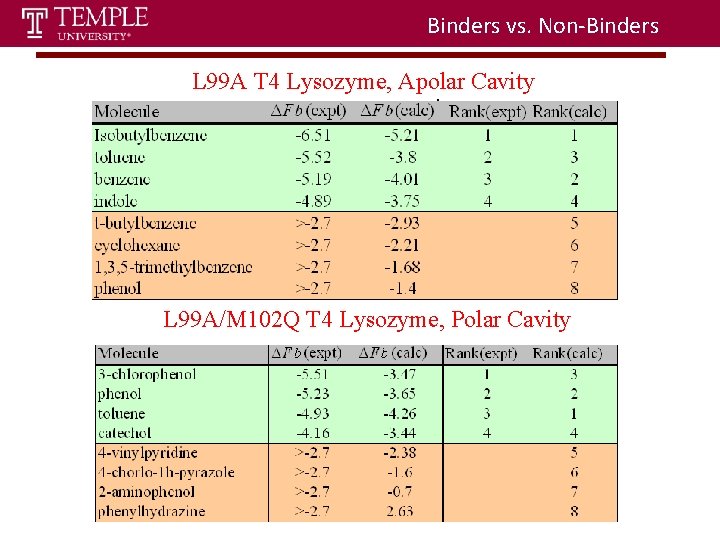
Binders vs. Non-Binders L 99 A T 4 Lysozyme, Apolar Cavity L 99 A/M 102 Q T 4 Lysozyme, Polar Cavity

SAMPL 4 Rutgers/Temple – E. Gallicchio, N. Deng, P. He, R. Levy Scripps - A. Perryman, S. Forli, D. Santiago, A. Olson,
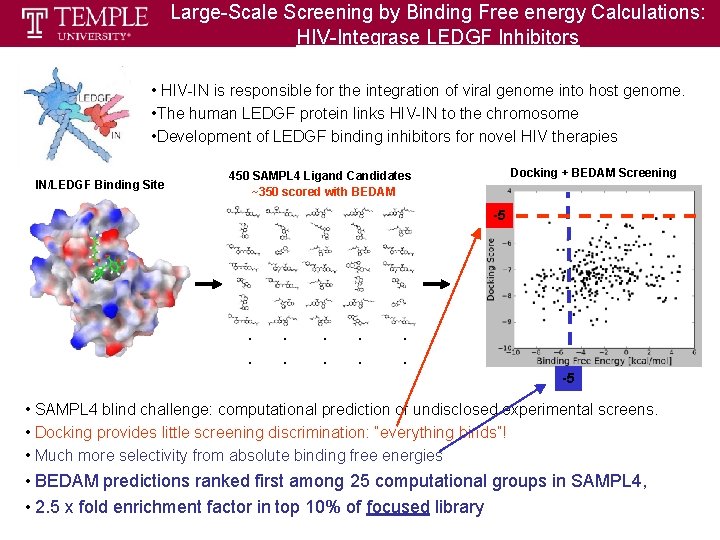
Large-Scale Screening by Binding Free energy Calculations: HIV-Integrase LEDGF Inhibitors • HIV-IN is responsible for the integration of viral genome into host genome. • The human LEDGF protein links HIV-IN to the chromosome • Development of LEDGF binding inhibitors for novel HIV therapies IN/LEDGF Binding Site Docking + BEDAM Screening 450 SAMPL 4 Ligand Candidates ~350 scored with BEDAM -5 . . -5 • SAMPL 4 blind challenge: computational prediction of undisclosed experimental screens. • Docking provides little screening discrimination: “everything binds”! • Much more selectivity from absolute binding free energies • BEDAM predictions ranked first among 25 computational groups in SAMPL 4, • 2. 5 x fold enrichment factor in top 10% of focused library
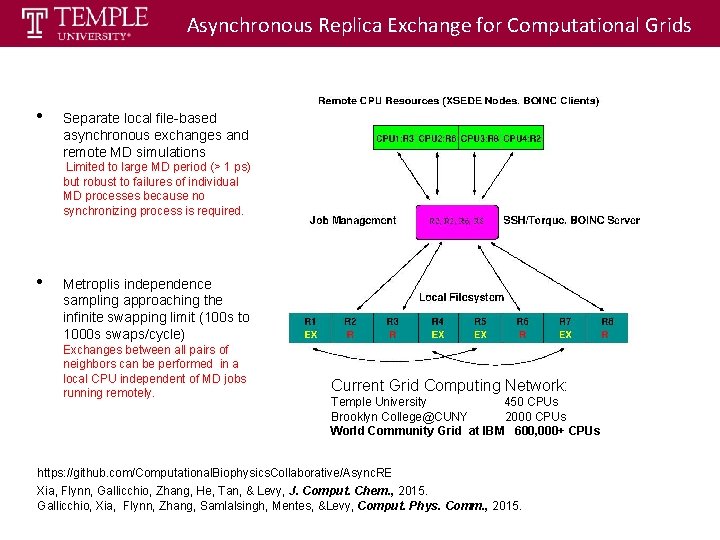
Asynchronous Replica Exchange for Computational Grids • Separate local file-based asynchronous exchanges and remote MD simulations MD running remotely Limited to large MD period (> 1 ps) but robust to failures of individual MD processes because no synchronizing process is required. • Metroplis independence sampling approaching the infinite swapping limit (100 s to 1000 s swaps/cycle) Exchanges between all pairs of neighbors can be performed in a local CPU independent of MD jobs running remotely. Exchange locally Current Grid Computing Network: Temple University 450 CPUs Brooklyn College@CUNY 2000 CPUs World Community Grid at IBM 600, 000+ CPUs https: //github. com/Computational. Biophysics. Collaborative/Async. RE Xia, Flynn, Gallicchio, Zhang, He, Tan, & Levy, J. Comput. Chem. , 2015. Gallicchio, Xia, Flynn, Zhang, Samlalsingh, Mentes, &Levy, Comput. Phys. Comm. , 2015.
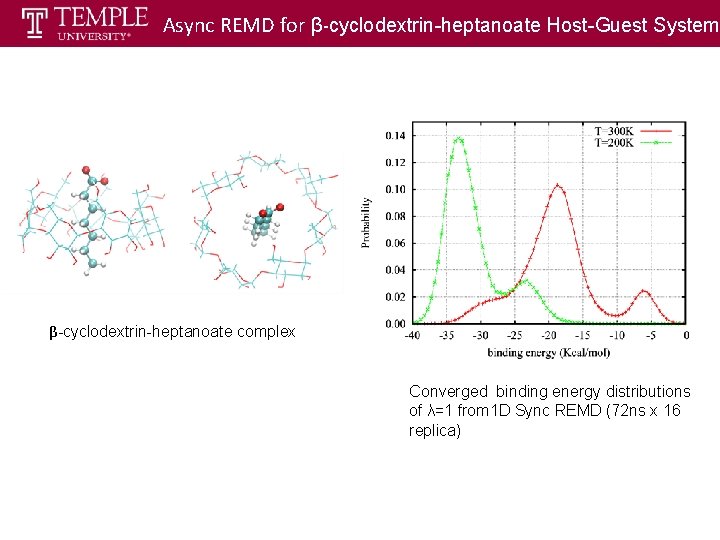
Async REMD for β-cyclodextrin-heptanoate Host-Guest System MD running remotely Exchange locally ) β-cyclodextrin-heptanoate complex Converged binding energy distributions of λ=1 from 1 D Sync REMD (72 ns x 16 replica)

2 D Async RE Results for β-cyclodextrin-heptanoate Complex MD running remotely Exchange locally ) Binding energy distributions of λ=1 from different REMD simulations at T=300 K Binding free energy as a function of λ from different REMD simulations at T=300 K
- Slides: 25