Statistical Thermodynamics Lecture 12 Solvation Models Molecular Mechanics























- Slides: 30

Statistical Thermodynamics Lecture 12: Solvation Models: Molecular Mechanics Modeling of Hydration Effects Dr. Ronald M. Levy ronlevy@temple. edu

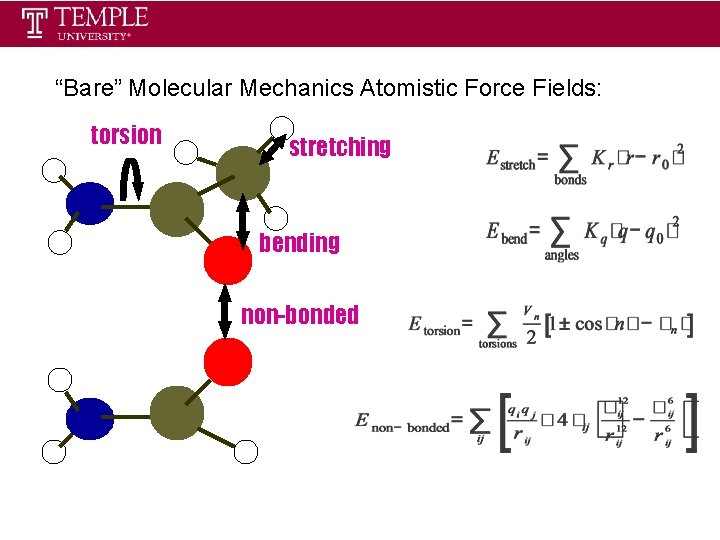
“Bare” Molecular Mechanics Atomistic Force Fields: torsion stretching bending non-bonded

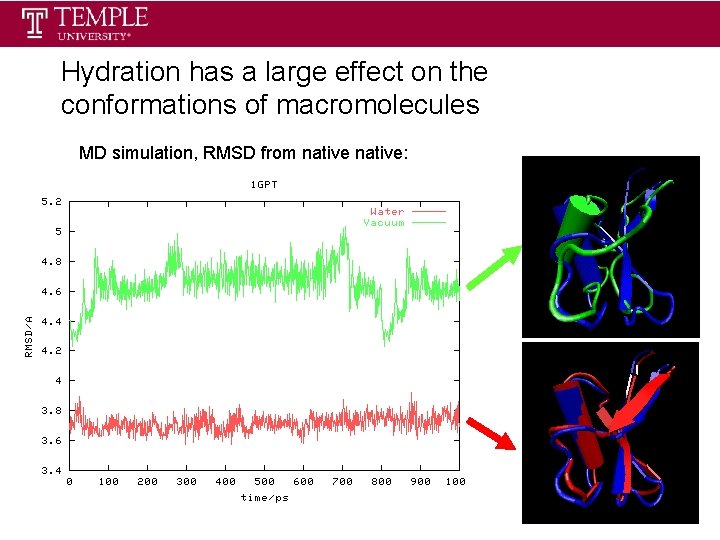
Hydration has a large effect on the conformations of macromolecules MD simulation, RMSD from native:

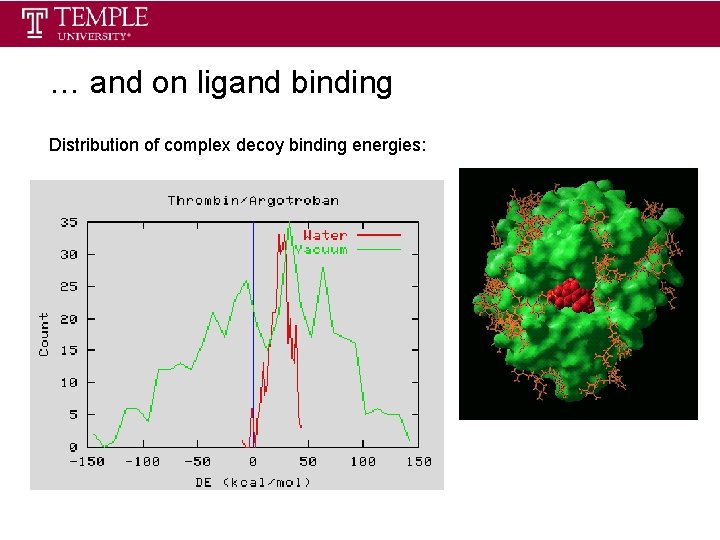
… and on ligand binding Distribution of complex decoy binding energies:

Challenge • Model hydration conveniently, as accurately as needed by the application and with the least computational cost. Two main approaches • Explicit solvation models. • Implicit solvation models.

Explicit Solvation • Each solvent molecule is represented with a set of atomic interaction centers (just as for the solute). • • Most accurate/detailed. Computationally expensive. Requires averaging over solvent coordinates. Difficult to obtain relative free energies of solute conformations.

Implicit Solvation • The solvent is represented by a continuum described by macroscopic parameters such as the dielectric constant, density, surface tension, etc. • Theoretical framework based on solvent PMF. • Not as accurate, especially for short-range solute-solvent interactions. • Reduced dimensionality. • Relative solvation free energies from single point effective potential energy calculations.

• • Many recent developments in molecular modeling have focused on solvation models. – Explicit: long-range electrostatic models – Implicit: improve coverage, accuracy and efficiency. Sometimes it is hard to keep track of all the choices. From a 2007 publication: The peptide is simulated in TIP 3 P and several variations of the GB implicit solvent model: GBHCT, GBOBC, and GBNeck (igb = 1, 5, and 7, respectively, in Amber 9). […] For consistency, MBondi radii were used in both the GB REMD simulations and subsequent GB and PE energy calculations described below. For TIP 3 P simulations, Ala 10 was solvated in a truncated octahedral box with 983 solvent molecules […] long-range electrostatic interactions were calculated using periodic boundary conditions via the particle mesh Ewald (PME). Roe, Wickstrom, et al. JPC B 111: 1846 (2007)

Explicit Hydration: Water Models • • • A water model specifies the interaction sites of a water molecule and their partial charges and LJ parameters. There is a large variety of models: rigid/flexible, 3 -points/4 -points or more, polarizable, dissociable, … The 3 rigid models most frequently used are SPC, TIP 3 P and TIP 4 P: SPC TIP 3 P TIP 4 P r(OH), Å 1. 0 0. 96 HOH, deg 109. 5 104. 5 σ(O), Å 3. 16 3. 15 ε(O), kcal/mol 0. 1554 0. 155 q(O) -0. 82 -0. 834 0. 0 q(H) 0. 417 0. 52 q(M) -1. 04

Periodic Boundary Conditions (PBC’s) • PBC’s are often used to simulate a bulk solution with a relatively small number of molecules. • The “central” simulation box is replicated in 3 D space so that the system has no edges. • The “minimum image convention” says that the actual distance between two particles is the smallest distance between all of the distances between their images: where L is the box size and x is the distance (along x) between any two images of a pair of particles. nint()=nearest integer. • A variety of space filling box shapes can be used. The “truncated octahedron” has particularly nice properties.

Treatment of long-range electrostatic interactions Electrostatic energy (per cell) of a periodic system: • The sum for the electrostatic potential φ includes all of the particles in the unit cell plus all of their images. • Straight sum is “conditionally convergent”; for example it would diverge if the positive and negative charges are summed separately. • Unlike LJ interactions, in this case truncation of the sum (with a distance cutoff) can be inaccurate. • In explicit solvent molecular simulation programs the infinite sum problem is circumvented using the “Ewald summation” class of models.

Ewald summation methods The Ewald sum is based on a decomposition of the 1/r interaction into a “fast decaying” component (handled with distance cutoffs) and a “smooth” component (summed using a Fourier transformation in reciprocal space). Think of surrounding each charge by a canceling Gaussian charge distribution: ρk is the Fourier transform of the charge distribution: measures the “softness” of the canceling distribution - the larger the more convergent is the direct sum (but less convergent is the reciprocal sum).

Particle mesh Ewald methods • Straightforward implementations of the Ewald sum scale at best as N 3/2 • More efficient implementations of complexity Nlog. N exploit FFT techniques by expressing the charge density on a regular grid. • The original idea of Hockeny and Eastwood (1981) consisted of assigning fractional charges to grid points: PPPM or P 3 M • Darden, York and Pedersen (1993) presented an (equivalent) approach based on Lagrange interpolation of the structure factors exp(ikr) on a grid: PME • PME was later refined using B-spline interpolation by Essmann et al. (1995) to obtain gradients for MD: SPME − In most recent papers, “PME” is SPME

Fast Multipole Methods • Based on representing distant charge distributions by multipole expansions. • Very favorable asymptotic scaling of order N – but with high initial overhead. • For values of N typical for biomolecular simulations FMM tends not to be as efficient as PME-based methods. • Still the only efficient method to treat longrange interactions without periodic boundary conditions. • Used routinely in simulations of million-body simulations of galaxies, etc. • A future for FMM in the context of implicit solvent modeling? Figuerido, Levy, Zhou, Berne. JCP 106: 9835 (1997)

Implicit Solvation: the solvent potential of mean force (SPMF) • Write the canonical partition function Qxy of the solution in terms of solute coordinates x, and solvent coordinates y. • The average of any observable that depends only on x can be written in terms of an “effective solvation potential” W(x). where defines the solvent potential of mean force • So called because the gradients of W(x) with respect to the solute coordinates are equal to the average forces of the solvent on the solute atoms at fixed solute conformation. • W(x) implicitly contains all of the effects of solvation.

The solvent potential of mean force is a free energy Conformational equilibria: x 2 x 1 Solvation (fixed solute conformation): y y x x

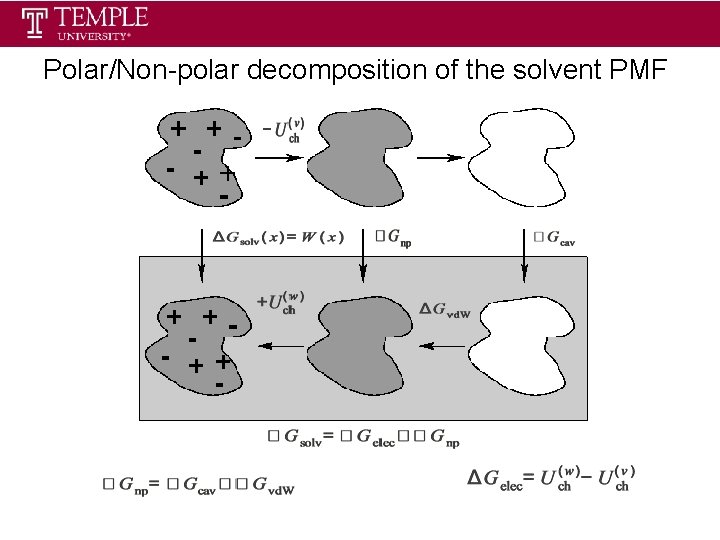
Polar/Non-polar decomposition of the solvent PMF

Typical Modern Implicit Solvent Model Electrostatic Component: Continuum Dielectric • Poisson-Boltzmann solvers (accurate but numerical and slow). • Generalized Born models (faster, can be expressed as analytic function). Non-Polar Component: • Solute surface area models • Cavity + van der Waals NP models.

The Poisson Equation (PE) and the Poisson-Boltzmann (PB) equations PE for a non-homogeneous dielectric: ε=1 ε= εslv Electrostatic free energy of solute: + - Electrostatic solvation free energy: direct Coulomb The reaction field potential φrf is due to the polarization of the medium.

The PB equation takes into account the effect of free ions in solution Debye-Hückel ion distribution - When (otherwise ion adsorption): ε=1 ε= εslv linearized form of the PB equation. where with (ionic strength). + +

Numerical solutions of the PB equation • The PB equation is solved on a grid in both surface and volume formulations. • Finite difference: solves the PB equation on a volume grid (APBS, Delphi, UHBD) • Finite element: solves integral form of the equation on a volume grid (PBF) • Boundary element: surface grid. • PB solvers often available in molecular simulation packages: Amber, CHARMM, IMPACT, etc. • Main drawback: continuum dielectric models are not suitable for specific short-range solute-solvent interactions, finite size effects, non-linear effects, high ionization states. • Other limitations are dependence on atomic radii parameters, speed, lack of analytical derivatives, dependence on frame of reference.

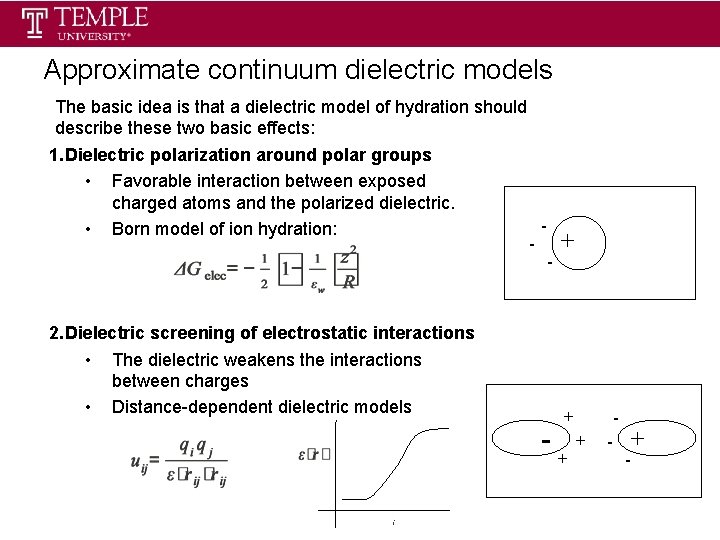
Approximate continuum dielectric models The basic idea is that a dielectric model of hydration should describe these two basic effects: 1. Dielectric polarization around polar groups • Favorable interaction between exposed charged atoms and the polarized dielectric. • Born model of ion hydration: - - 2. Dielectric screening of electrostatic interactions • The dielectric weakens the interactions between charges • Distance-dependent dielectric models - + + - - +

Generalized Born Model Bi is the Born radius of atom i defined by: • Satisfies the Born model in the two limits of infinite separation and complete overlap of solute atoms. • Basically it’s an interpolation formula.

Overall Features of Generalized Born Models The GB model “works” because it describes both dielectric polarization and dielectric screening effects. Polarization i=j (“self” energy): Favors the solvent exposure (small Bi) of polar groups (large q). Dielectric screening i≠j (pair energies):

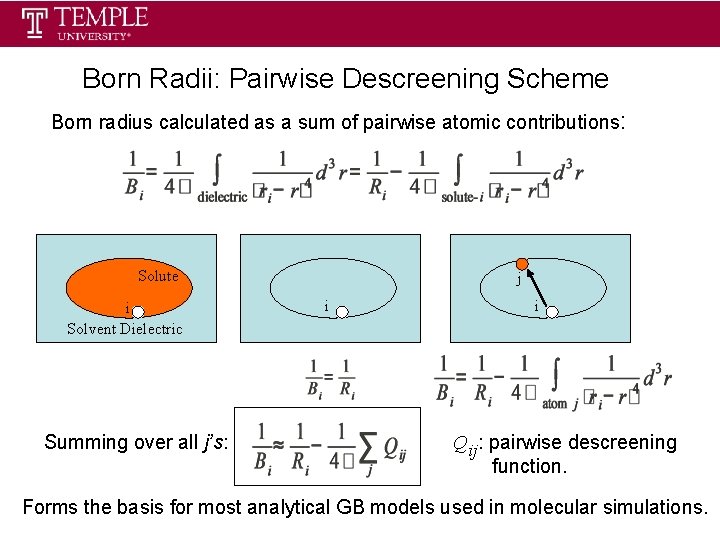
Born Radii: Pairwise Descreening Scheme Born radius calculated as a sum of pairwise atomic contributions: Solute i Solvent Dielectric Summing over all j’s: j i i Qij: pairwise descreening function. Forms the basis for most analytical GB models used in molecular simulations.

Pairwise Descreening Gotchas Qij is not simply the integral of 1/|ri - r|4 over atom j because: 1. Overlaps with atom j and other atoms would be over-counted j i Reduce descreening contribution from atom j 2. Atoms i and j may overlap j i Integrate only over portion of j not overlapping with i 3. Both situations can and do occur simultaneously.

GB implementations • Most major biomolecular simulation packages (CHARMM, Amber, IMPACT, Gromacs, etc. ) include pairwise descreening GB implementations suitable for MD calculations. • Key ingredients are the atomic radii and the description of the solute volume. • The atoms overlap problem is generally addressed by empirical scaling coefficients parameterized with respect to higher level calculations – that is the geometric model is parameterized in addition to the energetic model (ACE, GB/SA, GBHCT, GBSW) • Work on the AGBNP series of models shows that “geometric” parameterization is unnecessary. • Some implementations (GBMV, SGB) perform numerical integration on a grid (volume or surface) – non-analytic, higher computational cost, difficulties with derivatives, dependence on coordinate frame. • Some implementations differ in the choice of the GB distance function f(r) • Many of the models include continuum dielectric “correction” terms. • Recent developments have focused on the “interstitial” volumes problem (GBneck, GBMV, AGBNP 2).

Non-Polar Hydration Free Energy • Defined as the work of introducing the “uncharged” solute into solution. • In some ways a harder problem for modeling than electrostatics. • Often (and, nearly as often, incorrectly) ignored because appears “small” in magnitude. • Traditionally modeled in terms of the solvent-exposed surface of the solute by means of a surface tension parameter (SA models) • Motivated by macroscopic interfacial models and theories of hydration of cavities (probability of spontaneous occurrence of voids) in water. • Recent developments have moved beyond surface-only models adopting distinct geometric models for the cavity and van der Waals components of non-polar hydration (NP models):

Example of an analytical NP model (the “NP” in AGBNP) : Surface area of atom i : Geometrical predictor based on Born radius : Surface tension and van der Waals adjustable parameters

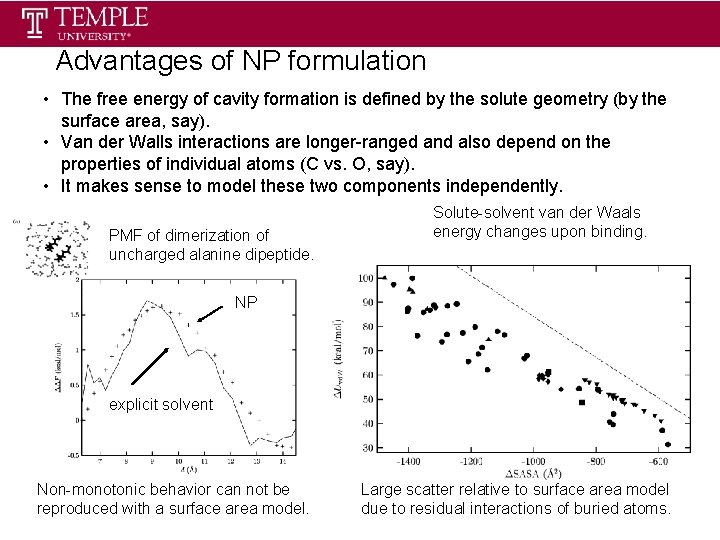
Advantages of NP formulation • The free energy of cavity formation is defined by the solute geometry (by the surface area, say). • Van der Walls interactions are longer-ranged and also depend on the properties of individual atoms (C vs. O, say). • It makes sense to model these two components independently. PMF of dimerization of uncharged alanine dipeptide. Solute-solvent van der Waals energy changes upon binding. NP explicit solvent Non-monotonic behavior can not be reproduced with a surface area model. Large scatter relative to surface area model due to residual interactions of buried atoms.