Statistical Significance for Peptide Identification by Tandem Mass

Statistical Significance for Peptide Identification by Tandem Mass Spectrometry Nathan Edwards Center for Bioinformatics and Computational Biology University of Maryland, College Park

Mass Spectrometry for Proteomics • Measure mass of many (bio)molecules simultaneously • High bandwidth • Mass is an intrinsic property of all (bio)molecules • No prior knowledge required 2

Mass Spectrometry for Proteomics • Measure mass of many molecules simultaneously • . . . but not too many, abundance bias • Mass is an intrinsic property of all (bio)molecules • . . . but need a reference to compare to 3
High Bandwidth % Intensity 100 0 250 500 750 4 1000 m/z

Mass is fundamental! 5

Mass Spectrometry for Proteomics • Mass spectrometry has been around since the turn of the century. . . • . . . why is MS based Proteomics so new? • Ionization methods • MALDI, Electrospray • Protein chemistry & automation • Chromatography, Gels, Computers • Protein sequence databases • A reference for comparison 6
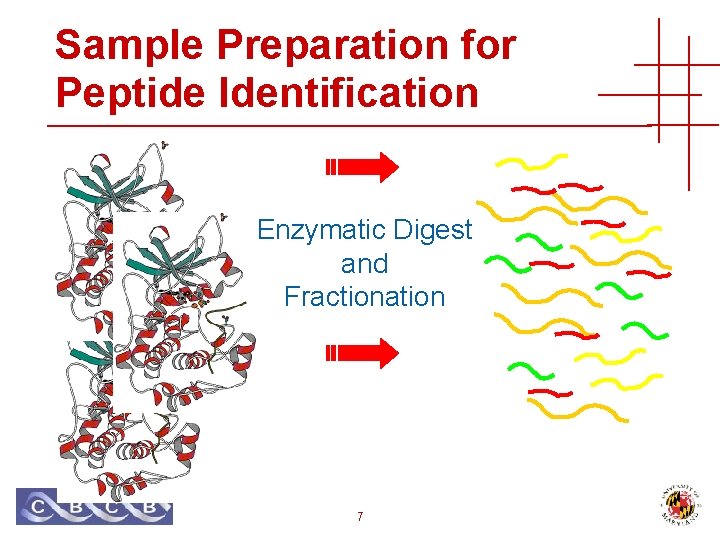
Sample Preparation for Peptide Identification Enzymatic Digest and Fractionation 7
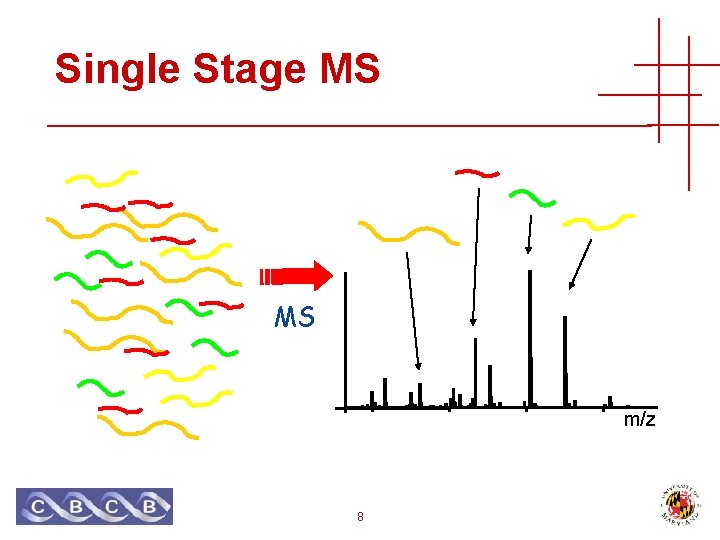
Single Stage MS MS m/z 8
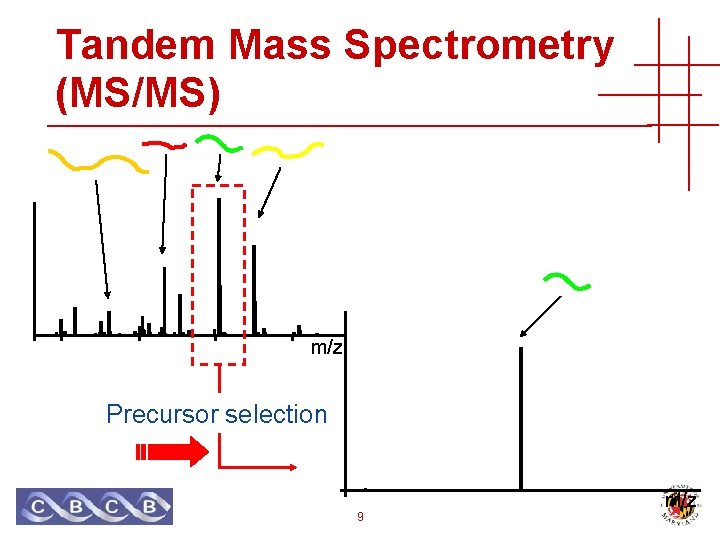
Tandem Mass Spectrometry (MS/MS) m/z Precursor selection 9 m/z
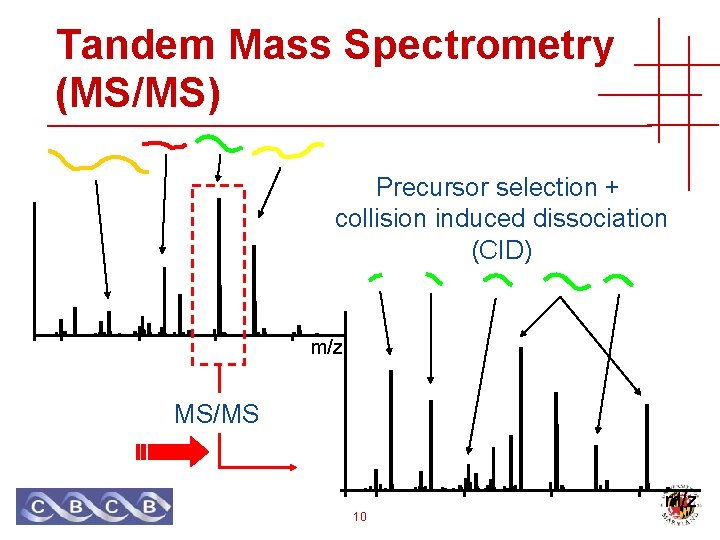
Tandem Mass Spectrometry (MS/MS) Precursor selection + collision induced dissociation (CID) m/z MS/MS 10 m/z
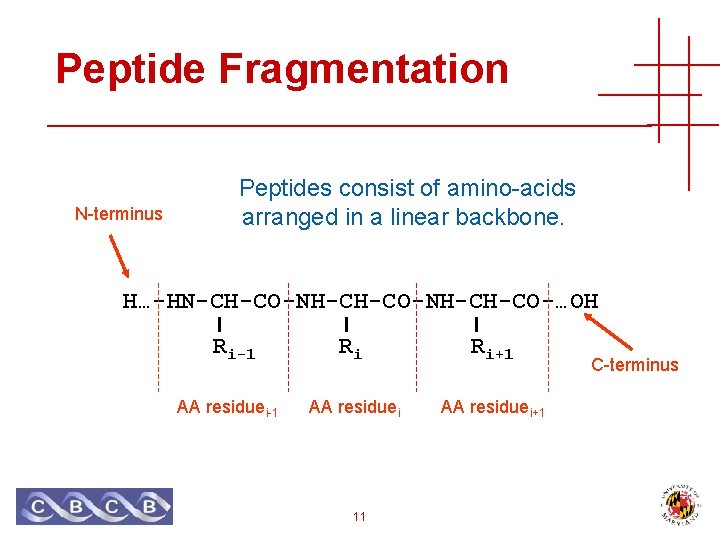
Peptide Fragmentation N-terminus Peptides consist of amino-acids arranged in a linear backbone. H…-HN-CH-CO-NH-CH-CO-…OH Ri-1 AA residuei-1 Ri Ri+1 AA residuei+1 11 C-terminus
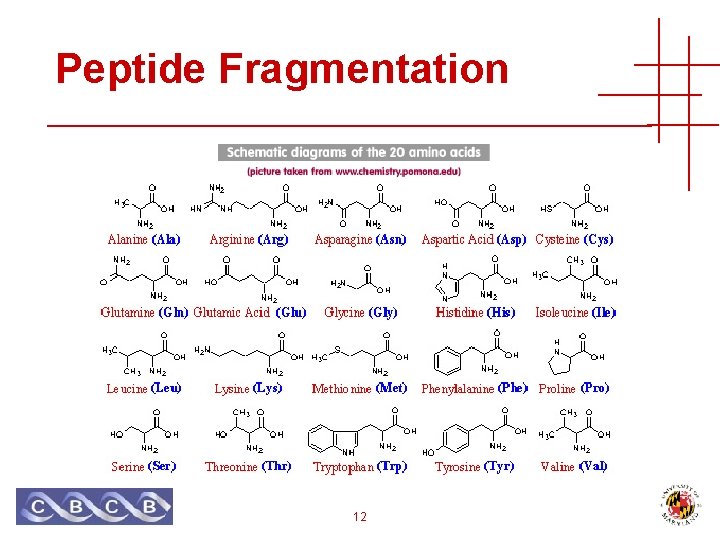
Peptide Fragmentation 12
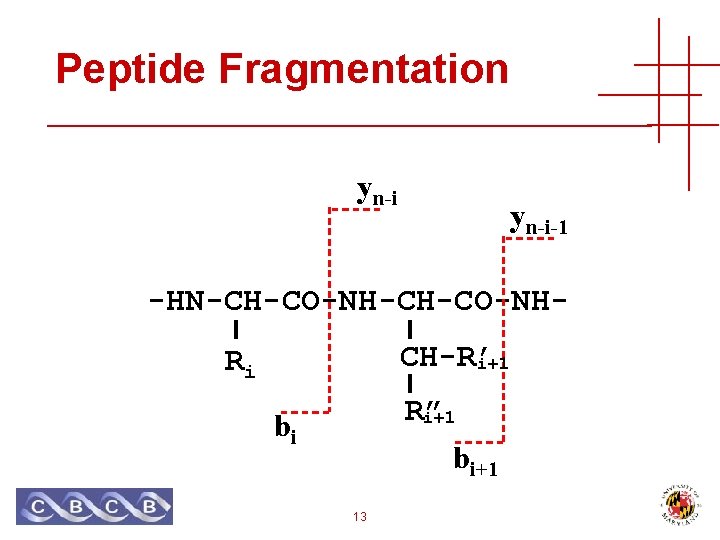
Peptide Fragmentation yn-i-1 -HN-CH-CO-NHCH-R’ i+1 Ri R” i+1 bi 13
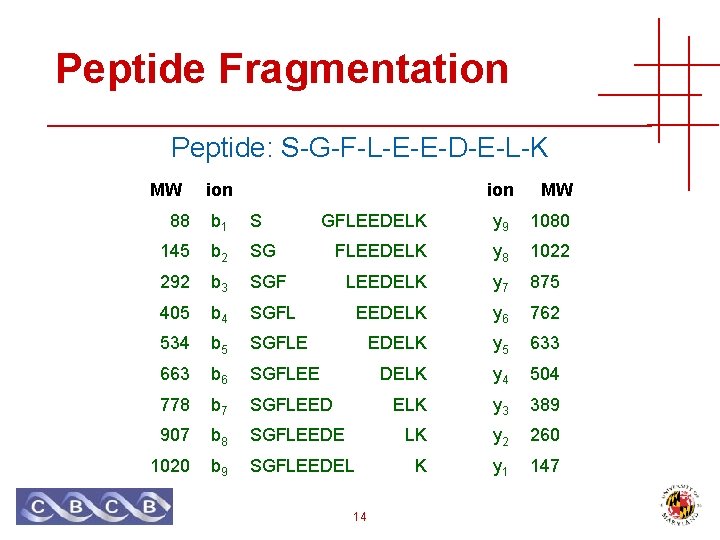
Peptide Fragmentation Peptide: S-G-F-L-E-E-D-E-L-K MW ion 88 b 1 S 145 b 2 SG 292 b 3 405 MW GFLEEDELK y 9 1080 FLEEDELK y 8 1022 SGF LEEDELK y 7 875 b 4 SGFL EEDELK y 6 762 534 b 5 SGFLE EDELK y 5 633 663 b 6 SGFLEE DELK y 4 504 778 b 7 SGFLEED ELK y 3 389 907 b 8 SGFLEEDE LK y 2 260 1020 b 9 SGFLEEDEL K y 1 147 14
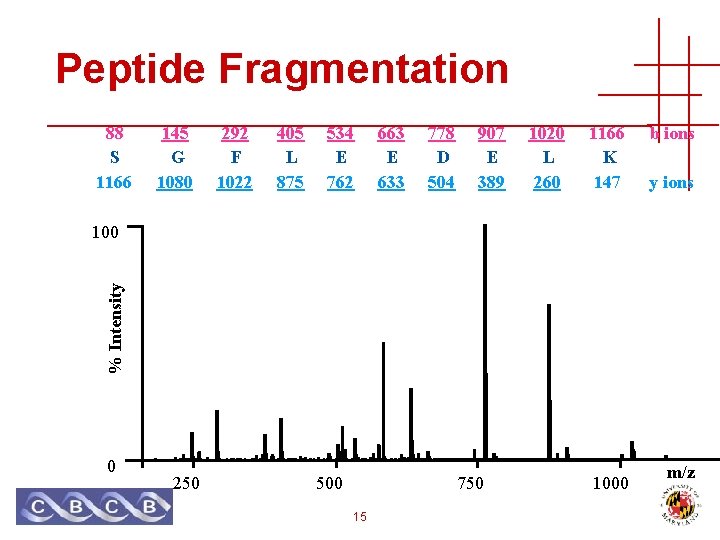
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 907 E 389 1020 L 260 1166 K 147 b ions y ions % Intensity 100 0 250 500 750 15 1000 m/z
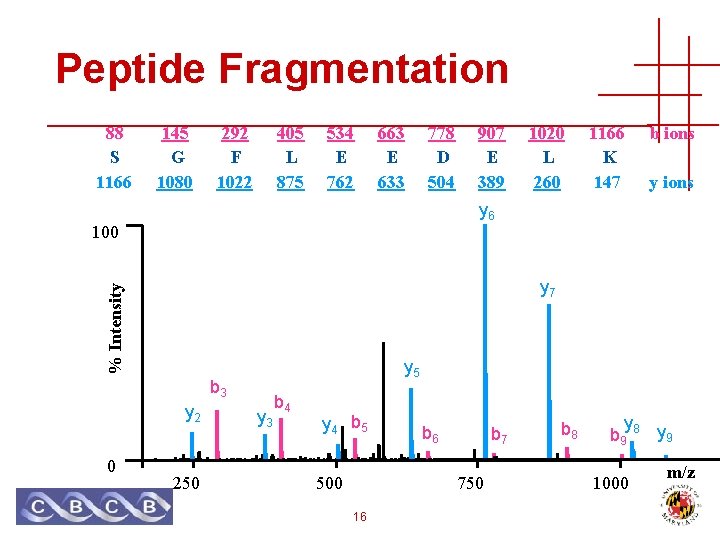
Peptide Fragmentation 88 S 1166 145 G 1080 292 F 1022 405 L 875 534 E 762 663 E 633 778 D 504 1020 L 260 1166 K 147 b ions y 6 100 % Intensity y 7 y 5 b 3 y 2 0 907 E 389 250 y 3 b 4 y 4 b 5 500 b 6 b 7 750 16 b 8 y b 9 8 y 9 1000 m/z

Peptide Identification • For each (likely) peptide sequence 1. Compute fragment masses 2. Compare with spectrum 3. Retain those that match well • Peptide sequences from protein sequence databases • Swiss-Prot, IPI, NCBI’s nr, . . . • Automated, high-throughput peptide identification in complex mixtures 17
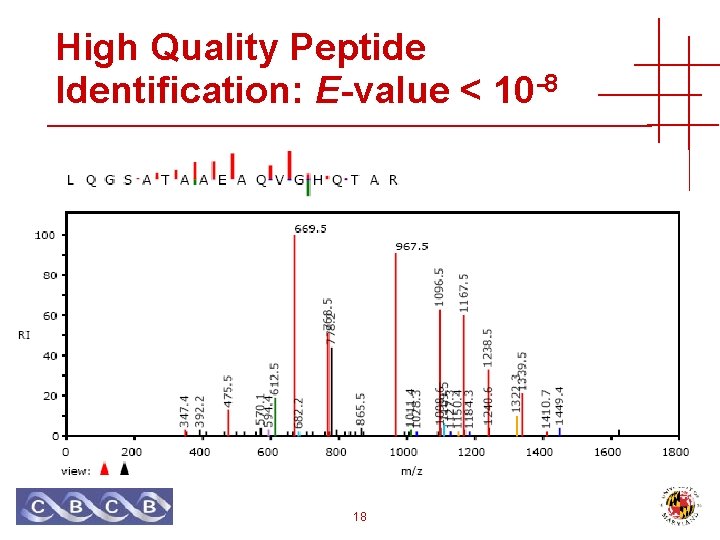
High Quality Peptide Identification: E-value < 10 -8 18
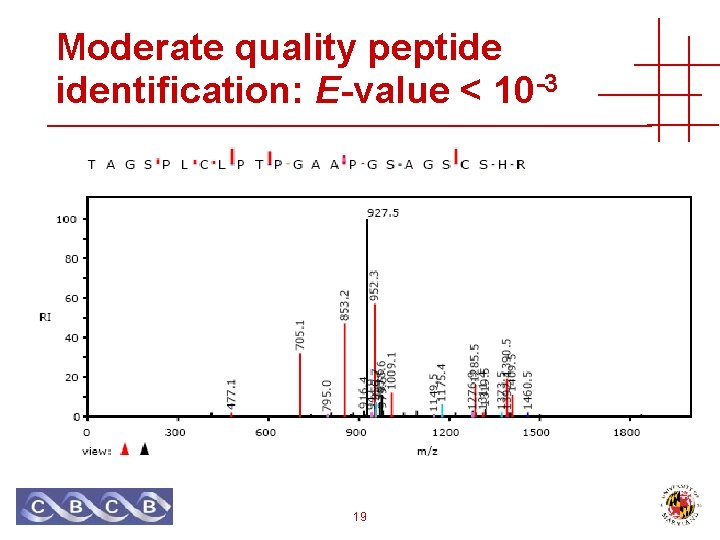
Moderate quality peptide identification: E-value < 10 -3 19
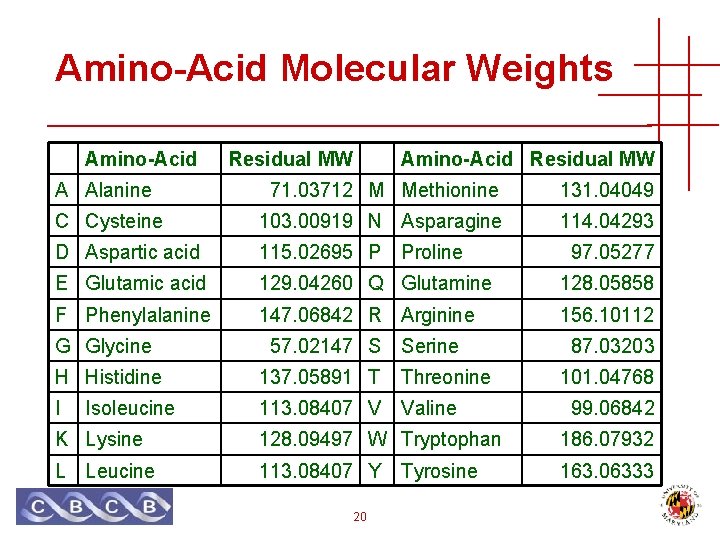
Amino-Acid Molecular Weights Amino-Acid Residual MW A Alanine 71. 03712 M Methionine 131. 04049 C Cysteine 103. 00919 N Asparagine 114. 04293 D Aspartic acid 115. 02695 P E Glutamic acid 129. 04260 Q Glutamine 128. 05858 F Phenylalanine 147. 06842 R Arginine 156. 10112 G Glycine 57. 02147 S Proline Serine H Histidine 137. 05891 T Threonine I 113. 08407 V Valine Isoleucine 97. 05277 87. 03203 101. 04768 99. 06842 K Lysine 128. 09497 W Tryptophan 186. 07932 L Leucine 113. 08407 Y 163. 06333 20 Tyrosine

Peptide Identification • Peptide fragmentation by CID is poorly understood • MS/MS spectra represent incomplete information about amino-acid sequence • I/L, K/Q, GG/N, … • Correct identifications don’t come with a certificate! 21

Peptide Identification • High-throughput workflows demand we analyze all spectra, all the time. • Spectra may not contain enough information to be interpreted correctly • …bad static on a cell phone • Peptides may not match our assumptions • …its all Greek to me • “Don’t know” is an acceptable answer! 22

Peptide Identification • Rank the best peptide identifications • Is the top ranked peptide correct? 23

Peptide Identification • Rank the best peptide identifications • Is the top ranked peptide correct? 24

Peptide Identification • Rank the best peptide identifications • Is the top ranked peptide correct? 25

Peptide Identification • Incorrect peptide has best score • Correct peptide is missing? • Potential for incorrect conclusion • What score ensures no incorrect peptides? • Correct peptide has weak score • Insufficient fragmentation, poor score • Potential for weakened conclusion • What score ensures we find all correct peptides? 26

Statistical Significance • Can’t prove particular identifications are right or wrong. . . • . . . need to know fragmentation in advance! • A minimal standard for identification scores. . . • . . . better than guessing. • p-value, E-value, statistical significance 27

Pin the tail on the donkey… 28

Probability Concepts Throwing darts • One at a time • Blindfolded Uniform distribution? Independent? Identically distributed? Pr [ Dart hits 20 ] = 0. 05 29

Probability Concepts Throwing darts • One at a time • Blindfolded • Three darts Pr [Hitting 20 3 times] = 0. 05 * 0. 05 Pr [Hit 20 at least twice] = 0. 007125 + 0. 000125 30 0 times 1 times 2 times 3 times 0. 857375 0. 135375 0. 007125 0. 000125
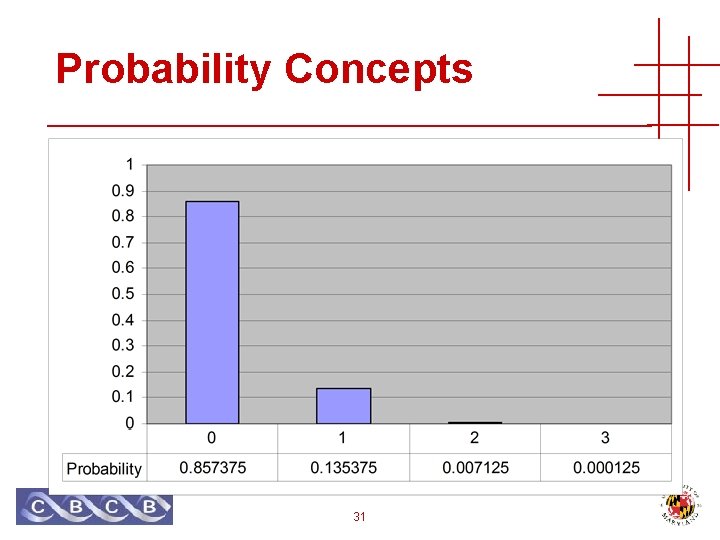
Probability Concepts 31

Probability Concepts Throwing darts • One at a time • Blindfolded • 100 darts Pr [Hitting 20 3 times] = 0. 139575 0 times 1 times 2 times 3 times Pr [Hit 20 at least twice] = 0. 9629188 32 0. 005920 0. 031160 0. 081181 0. 139575
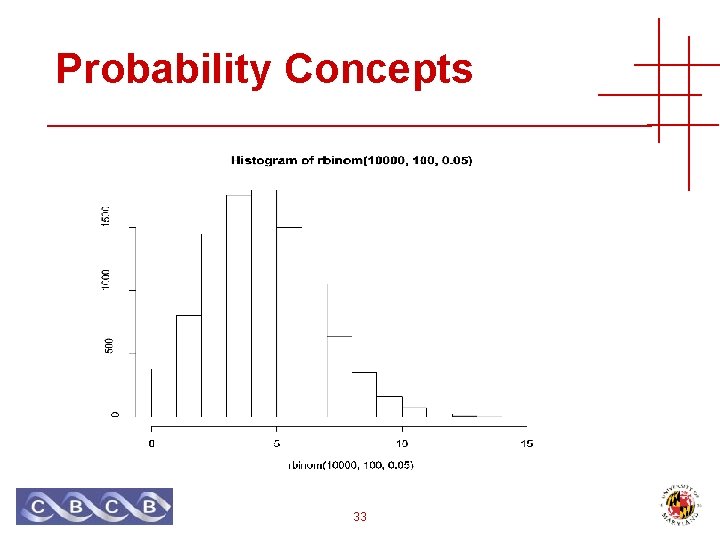
Probability Concepts 33

Match Score • Dartboard represents the mass range of the spectrum • Peaks of a spectrum are “slices” • Width of slice corresponds to mass tolerance • Darts represent • random masses • masses of fragments of a random peptide • masses of peptides of a random protein • masses of biomarkers from a random class • How many darts do we get to throw? 34
Match Score 100 % Intensity • What is the probability that we match at least 5 peaks? 270 0 250 500 750 1000 m/z 330 870 550 35 755 580
![Match Score • Pr [ Match ≥ s peaks ] = Binomial( p , Match Score • Pr [ Match ≥ s peaks ] = Binomial( p ,](http://slidetodoc.com/presentation_image_h2/e833e4185f660c7cd5d128ac732b19ac/image-36.jpg)
Match Score • Pr [ Match ≥ s peaks ] = Binomial( p , n ) ≈ Poisson( p n ), for small p and large n p is prob. of random mass / peak match, n is number of darts (fragments in our answer) 36

Match Score Theoretical distribution • Used by OMSSA • Proposed, in various forms, by many. • Probability of random mass / peak match • IID (independent, identically distributed) • Based on match tolerance 37

Match Score Theoretical distribution assumptions • Each dart is independent • Peaks are not “related” • Each dart is identically distributed • Chance of random mass / peak match is the same for all peaks 38
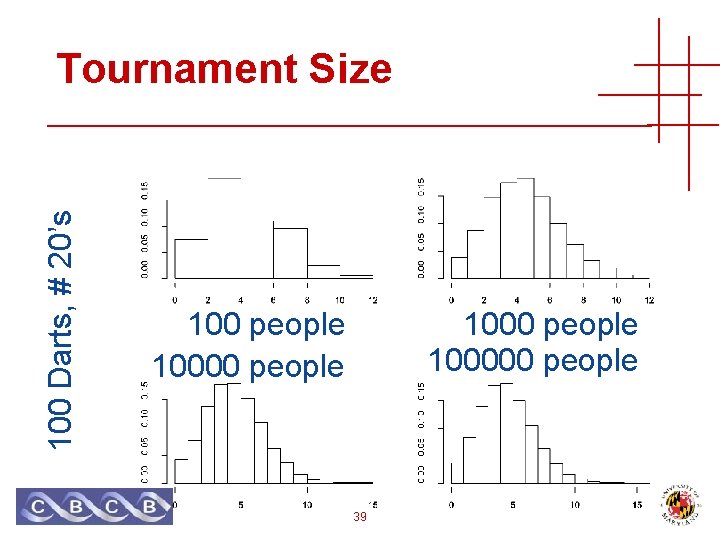
100 Darts, # 20’s Tournament Size 100 people 100000 people 39
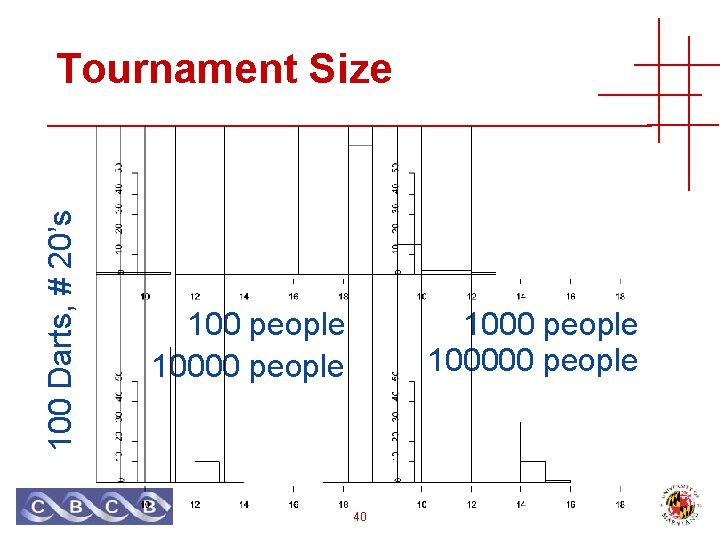
100 Darts, # 20’s Tournament Size 100 people 100000 people 40

Number of Trials • Tournament size == number of trials • Number of peptides tried • Related to sequence database size • Probability that a random match score is ≥ s • 1 – Pr [ all match scores < s ] • 1 – Pr [ match score < s ] Trials • Assumes IID! • Expect value (*) • E = Trials * Pr [ match ≥ s ] • Corresponds to Bonferroni bound on (*) 41

Better Dart Throwers 42

Better Random Models • Comparison with completely random model isn’t really fair • Match scores for real spectra with real peptides obey rules • Even incorrect peptides match with non-random structure! 43

Better Random Models • Want to generate random fragment masses (darts) that behave more like the real thing: • Some fragments are more likely than others • Some fragments depend on others • Theoretical models can only incorporate this structure to a limited extent. 44

Better Random Models • Generate random peptides • Real looking fragment masses • No theoretical model! • Must use empirical distribution • Usually require they have the correct precursor mass • Score function can model anything we like! 45
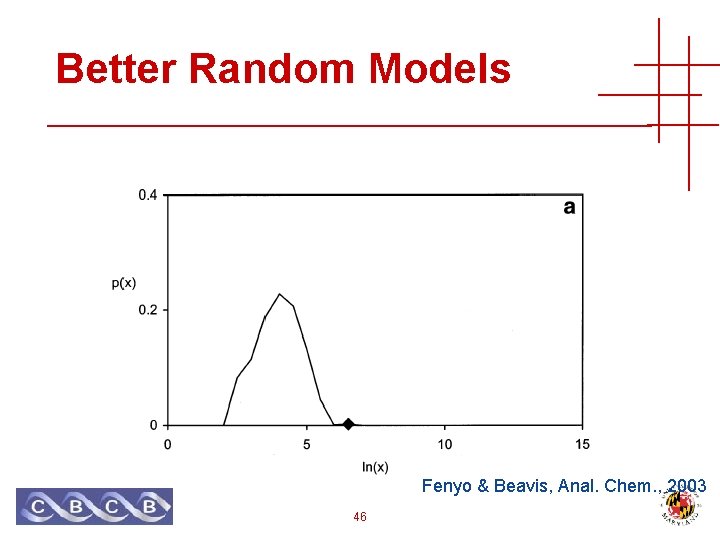
Better Random Models Fenyo & Beavis, Anal. Chem. , 2003 46
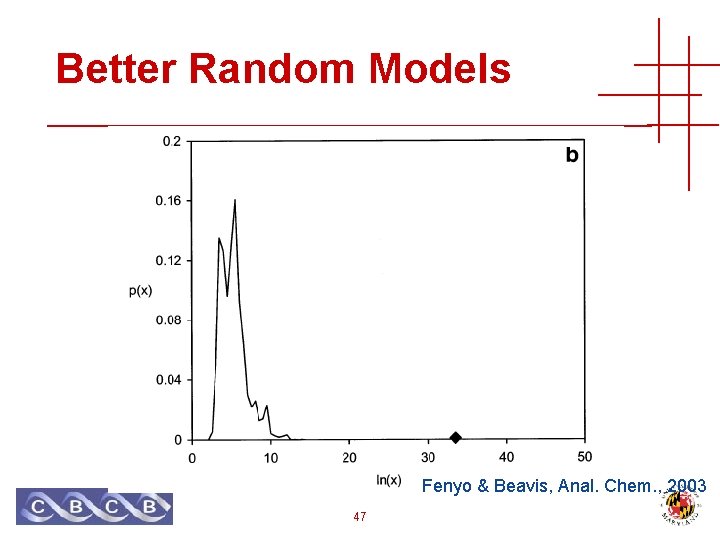
Better Random Models Fenyo & Beavis, Anal. Chem. , 2003 47

Better Random Models • Truly random peptides don’t look much like real peptides • Just use peptides from the sequence database! • Caveats: • Correct peptide (non-random) may be included • Peptides are not independent • Reverse sequence avoids only the first problem 48
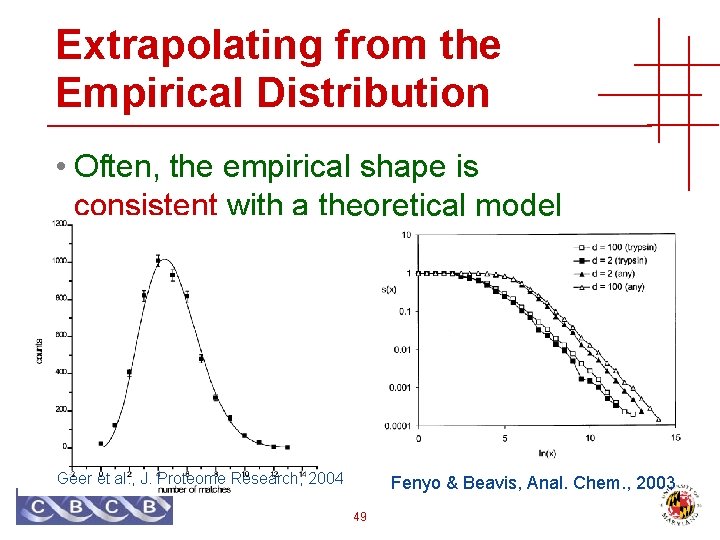
Extrapolating from the Empirical Distribution • Often, the empirical shape is consistent with a theoretical model Geer et al. , J. Proteome Research, 2004 Fenyo & Beavis, Anal. Chem. , 2003 49

False Positive Rate Estimation • Each spectrum is a chance to be right, wrong, or inconclusive. • How many decisions are wrong? • Given identification criteria: • SEQUEST Xcorr, E-value, Score, etc. , plus. . . • . . . threshold • Use “decoy” sequences • random, reverse, cross-species • Identifications must be incorrect! 50

False Positive Rate Estimation • # FP in real search = # hits in decoy search • Need same size database, or rate conversion • FP Rate: # decoy hits # real hits • FP Rate: 2 x # decoy hits. (# real hits + # decoy hits) 51

False Positive Rate Estimation • A form of statistical significance • In “theory”, E-value and a FP rate are the same. • Search engine independent • Easy to implement • Assumes a single threshold for all spectra • Spectrum/Peptide Identification scores are not iid!. . . • . . . but E-values, in principle, are. 52

Peptide Prophet • From the Institute for Systems Biology • Keller et al. , Anal. Chem. 2002 • Re-analysis of SEQUEST results • Spectra are trials • Assumes that many of the spectra are not correctly identified 53
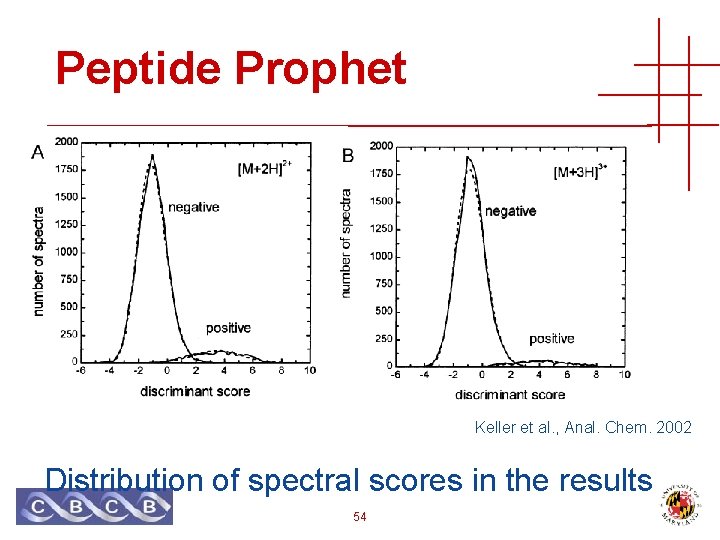
Peptide Prophet Keller et al. , Anal. Chem. 2002 Distribution of spectral scores in the results 54

Peptide Prophet • Assumes a bimodal distribution of scores, with a particular shape • Ignores database size • …but it is included implicitly • Like empirical distribution for peptide sampling, can be applied to any score function • Can be applied to any search engines’ results 55

Peptide Prophet • Caveats • Are spectra scores sampled from the same distribution? • Is there enough correct identifications for second peak? • Are spectra independent observations? • Are distributions appropriately shaped? • Huge improvement over raw SEQUEST results 56
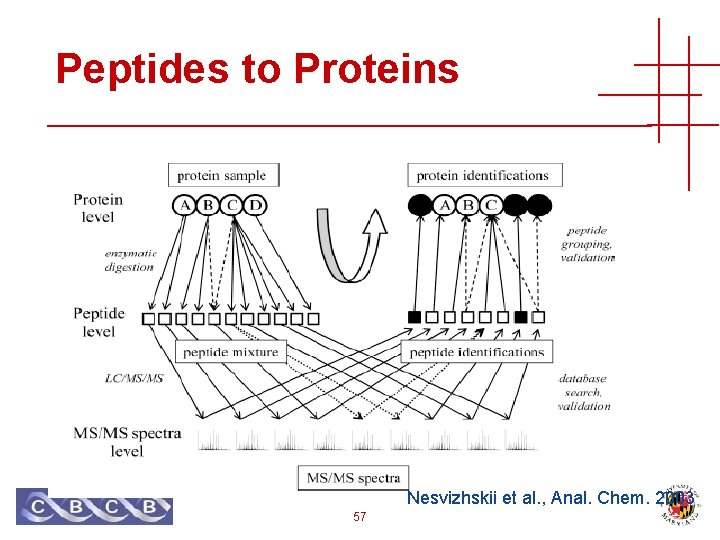
Peptides to Proteins Nesvizhskii et al. , Anal. Chem. 2003 57
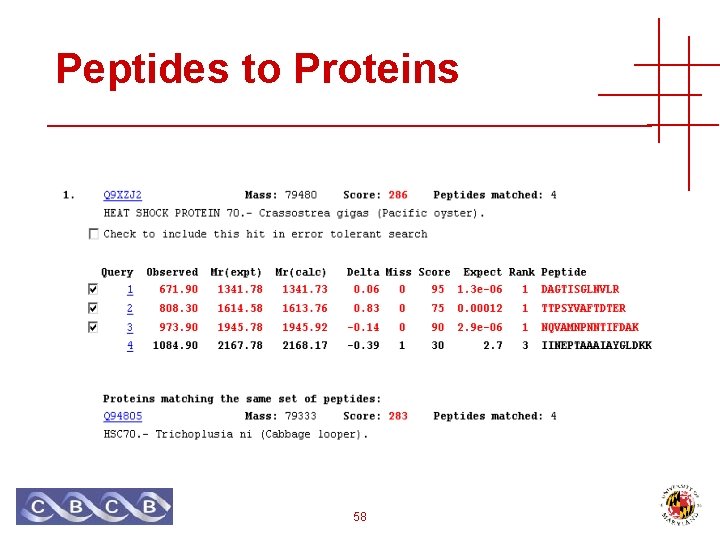
Peptides to Proteins 58

Peptides to Proteins • A peptide sequence may occur in many different protein sequences • Variants, paralogues, protein families • Separation, digestion and ionization is not well understood • Proteins in sequence database are extremely non-random, and very dependent 59

Publication Guidelines 60

Publication Guidelines 1. Computational parameters • • Spectral processing Sequence database Search program Statistical analysis 2. Number of peptides per protein • • Each peptide sequence counts once! Multiple forms of the same peptide count once! 61

Publication Guidelines 3. Single-peptide proteins must be explicitly justified by • • • Peptide sequence N and C terminal amino-acids Precursor mass and charge Peptide Scores Multiple forms of the peptide counted once! 4. Biological conclusions based on singlepeptide proteins must show the spectrum 62

Publication Guidelines 5. More stringent requirements for PMF data analysis • Similar to that for tandem mass spectra 6. Management of protein redundancy • Peptides identified from a different species? 7. Spectra submission encouraged 63

Summary • Could guessing be as effective as a search? • More guesses improves the best guess • Better guessers help us be more discriminating • Peptide to proteins is not as simple as it seems • Publication guidelines reflect sound statistical principles. 64
- Slides: 64