State of CBIL Current and Future Directions Computational

State of CBIL Current and Future Directions

Computational Biology and Informatics Laboratory October, 2001

CBIL Research • Gene Discovery – EST analysis – Genomic sequence analysis • Gene Regulation – Microarray analysis – Promoter/ regulatory region analysis • Biological data representation – Data integration – Ontology

CBIL: Gene Discovery Ø Gene Annotation (Kolchanov) Ø Gene coding potential Ø Gene function prediction Ø All. Genes Ø EPCon. DB (Kaestner, Permutt, Melton) • Stem. Cell. DB/ Stro. CDB (Lemischka, Moore) Ø Mouse chromosome 5 (Bucan) Ø Plasmo. DB (Roos, Kissinger) • Para. DB (Roos) Ø Posters

CBIL: Gene Regulation Ø Pa. GE Ø PROM_REC • TESS Ø Pancreatic development (Kaestner) Ø TGF-B signaling (Bottinger) • Fetal globin expression in adults (Fortina) • Brain disease and injury (Eberwine, Meaney) • Endothelial cell function (Davies)

CBIL: Biological Data Representation Ø Genomic Unified Schema • RNA Abundance Database • Connecting to a brain atlas (Nissanov, Davidson) Ø Microarray ontology (MGED)

CBIL Project Architecture Sequence & annotation Gene index (ESTs and m. RNAs) GUS Microarray expression data experimental annotation RAD Relational DB (Oracle) with Perl object layer
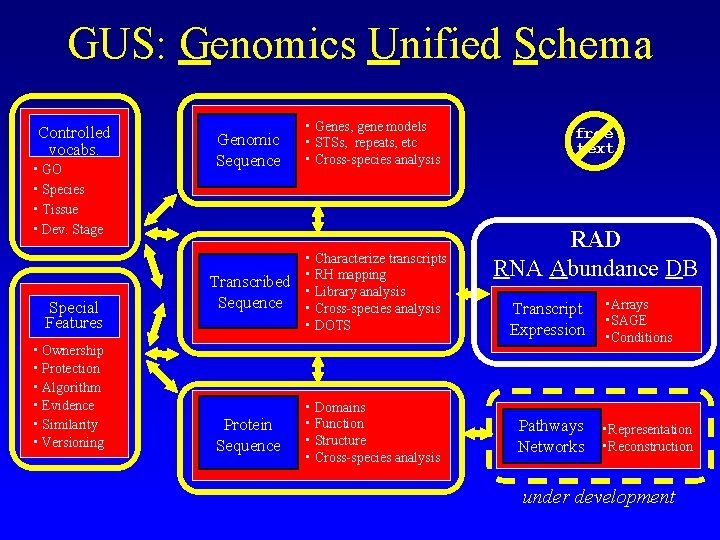
GUS: Genomics Unified Schema Controlled vocabs. • GO • Species • Tissue • Dev. Stage Special Features • Ownership • Protection • Algorithm • Evidence • Similarity • Versioning Genomic Sequence • Genes, gene models • STSs, repeats, etc • Cross-species analysis Transcribed Sequence • Characterize transcripts • RH mapping • Library analysis • Cross-species analysis • DOTS Protein Sequence • Domains • Function • Structure • Cross-species analysis free text RAD RNA Abundance DB Transcript Expression Pathways Networks • Arrays • SAGE • Conditions • Representation • Reconstruction under development
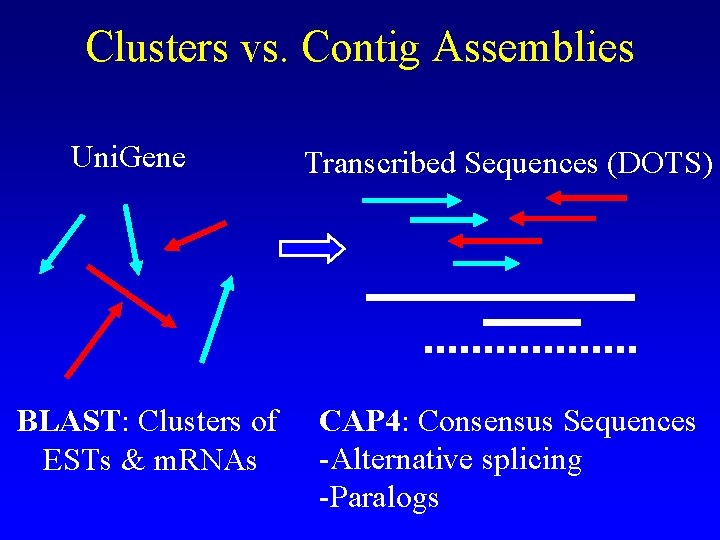
Clusters vs. Contig Assemblies Uni. Gene BLAST: Clusters of ESTs & m. RNAs Transcribed Sequences (DOTS) CAP 4: Consensus Sequences -Alternative splicing -Paralogs

Assembled Transcripts About 3 million human EST and m. RNA sequences used Combined into 797, 028 assemblies Cluster into 150, 006 “genes” Can identify a protein for 76, 771 genes And predict a function for 24, 127 genes About 2 million mouse EST and m. RNA sequences used Combined into 355, 770 assemblies Cluster into 74, 024 “genes” Can identify a protein for 34, 008 genes And predict a function for 15, 403 genes
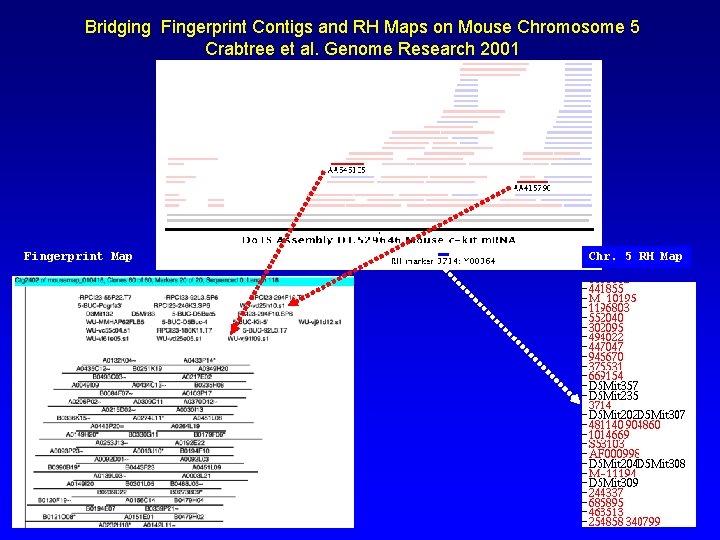
Bridging Fingerprint Contigs and RH Maps on Mouse Chromosome 5 Crabtree et al. Genome Research 2001 Fingerprint Map Chr. 5 RH Map
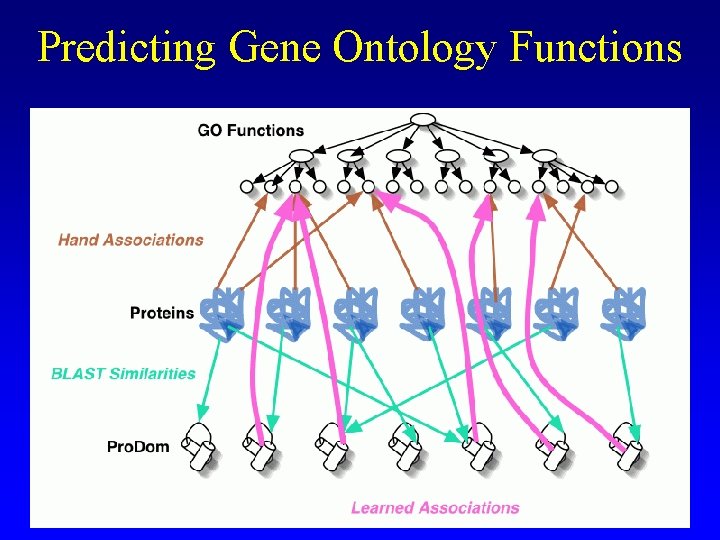
Predicting Gene Ontology Functions
All. Genes

All. Genes Enhancements: Annotated Entries
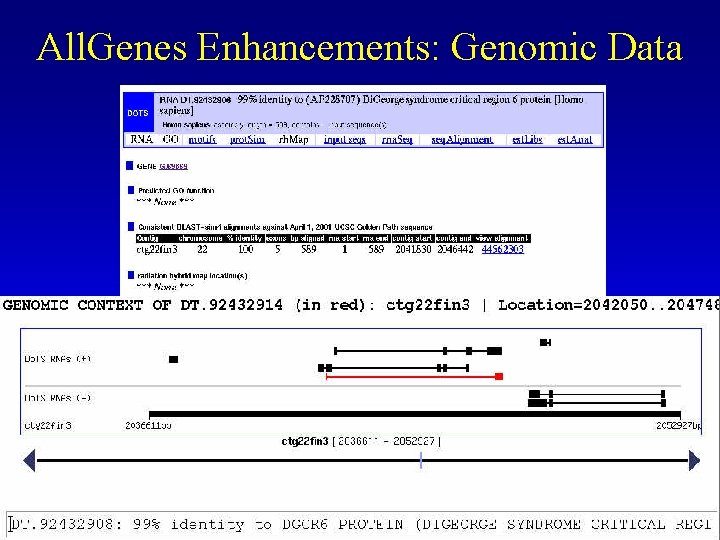
All. Genes Enhancements: Genomic Data
http: //plasmodb. org
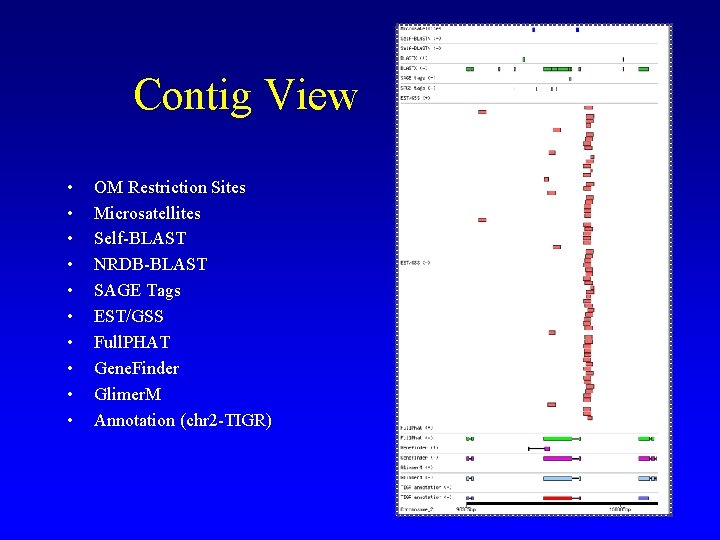
Contig View • • • OM Restriction Sites Microsatellites Self-BLAST NRDB-BLAST SAGE Tags EST/GSS Full. PHAT Gene. Finder Glimer. M Annotation (chr 2 -TIGR)
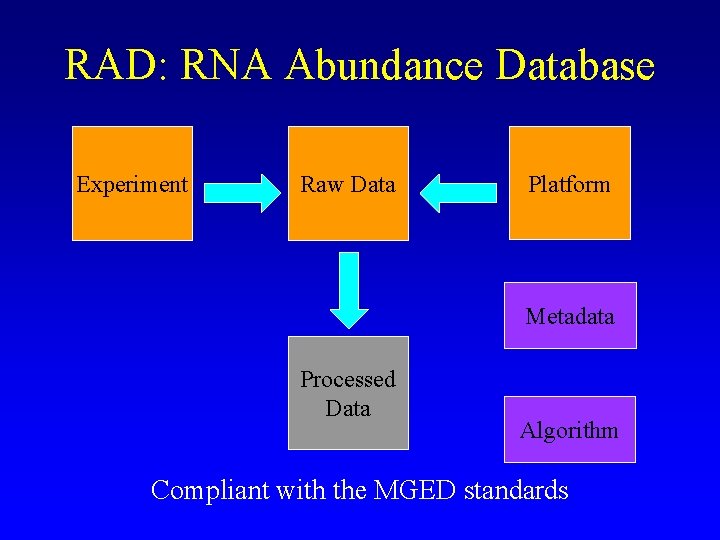
RAD: RNA Abundance Database Experiment Raw Data Platform Metadata Processed Data Algorithm Compliant with the MGED standards

Microarray Gene Expression Database group (MGED) International effort on microarray data standards: – Develop standards for storing and communicating microarray-based gene expression data • defining the minimal information required to ensure reproducibility and verifiability of results and to facilitate data exchange (MIAME, MAGEML-MAGEDOM) • collecting (and where needed creating) controlled vocabularies/ ontologies. • developing standards for data comparison and normalization. http: //www. mged. org

EPCon. DB Pathway query
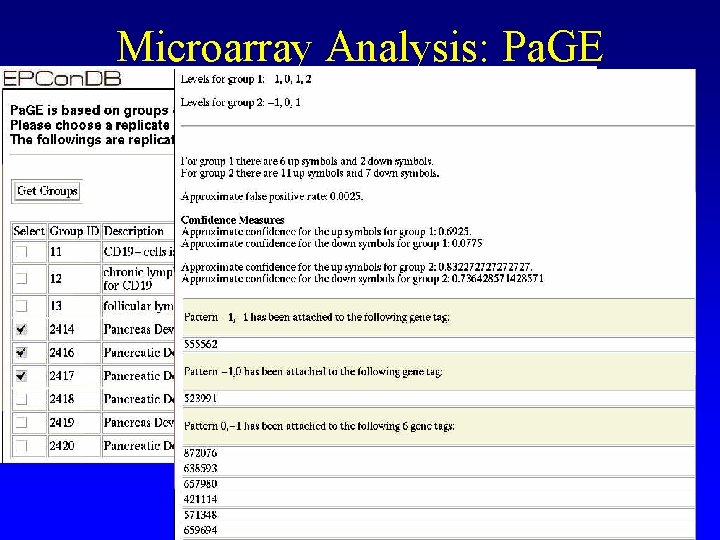
Microarray Analysis: Pa. GE
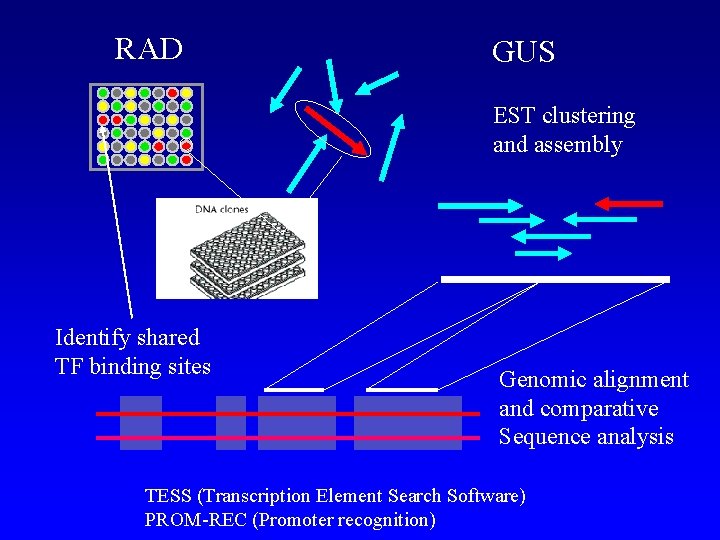
RAD GUS EST clustering and assembly Identify shared TF binding sites Genomic alignment and comparative Sequence analysis TESS (Transcription Element Search Software) PROM-REC (Promoter recognition)

Promoter Analysis: PROM_REC

http: www. cbil. upenn. edu

Acknowledgements CBIL: Chris Stoeckert Vladimir Babenko Brian Brunk Jonathan Crabtree Sharon Diskin Greg Grant Yuri Kondrakhin Georgi Kostov Phil Le Li Li Junmin Liu Elisabetta Manduchi Joan Mazzarelli Shannon Mc. Weeney Debbie Pinney Angel Pizarro Jonathan Schug Plasmo. DB collaborators: David Roos Martin Fraunholz Jesse Kissinger Jules Milgram Ross Koppel, Monash U. Malarial Genome Sequencing Consortium (Sanger Centre, Stanford U. , TIGR/NMRC) EPCon. DB collaborators: Klaus Kaestner Marie Scearce Doug Melton, Harvard Alan Permutt, Wash. U Comparative Sequence Analysis Collaborators: Maja Bucan Shaying Zhao Whitehead/MIT Center for Genome Research CAP 4 provided by Paracel

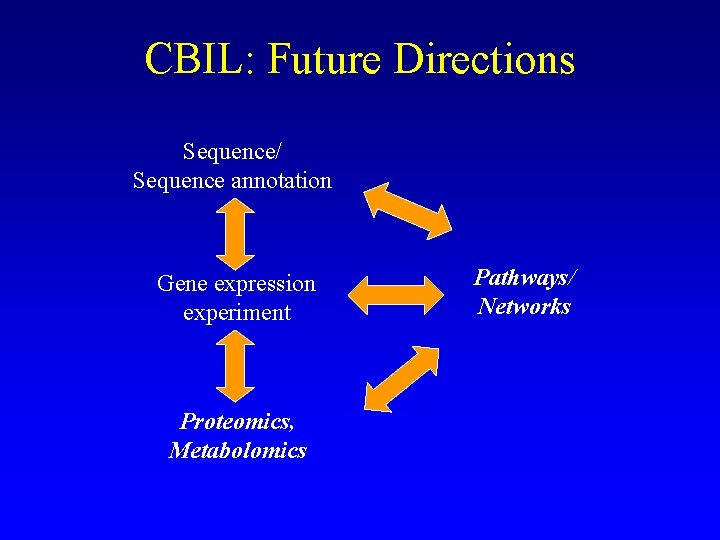
CBIL: Future Directions Sequence/ Sequence annotation Gene expression experiment Proteomics, Metabolomics Pathways/ Networks
- Slides: 27