Species from faeces predator scat metabarcoding in Tasmania

Species from faeces: predator scat metabarcoding in Tasmania Photo by Matt Young Photo DPIPWE Anna Mac. Donald, Dianne Gleeson & Stephen Sarre Institute for Applied Ecology, University of Canberra Twitter: @Dr_Anna. M

Feral cat with bandicoot Image by Guy Ballard, from Victorian camera trap study Changing predatorprey dynamics in Tasmania Introduced predators threaten native wildlife Fox in a Canberra nature reserve Photo by Matt Young
![Scats contain predator DNA and information about predator diet [Photos from DPIPWE] 3 Scats contain predator DNA and information about predator diet [Photos from DPIPWE] 3](http://slidetodoc.com/presentation_image_h2/6ac8e04ccd9f3a50cb0f934795830bef/image-3.jpg)
Scats contain predator DNA and information about predator diet [Photos from DPIPWE] 3
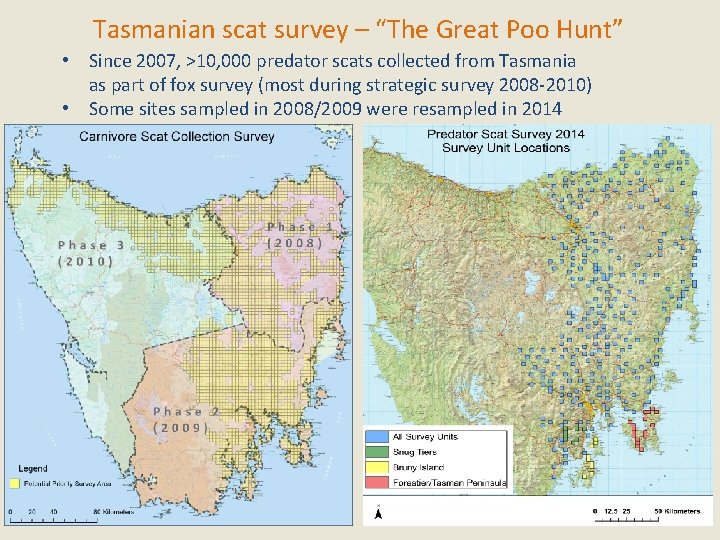
Tasmanian scat survey – “The Great Poo Hunt” • Since 2007, >10, 000 predator scats collected from Tasmania as part of fox survey (most during strategic survey 2008 -2010) • Some sites sampled in 2008/2009 were resampled in 2014 4

Scats could provide data on predator-prey interactions and species distributions • Fox / cat diets – potential impacts on wildlife / agriculture? • Diets of native vs introduced predators? • How are predators distributed in the landscape? • How are prey species distributed? • How have predator-prey interactions changed over time, in regions with / without perturbation?
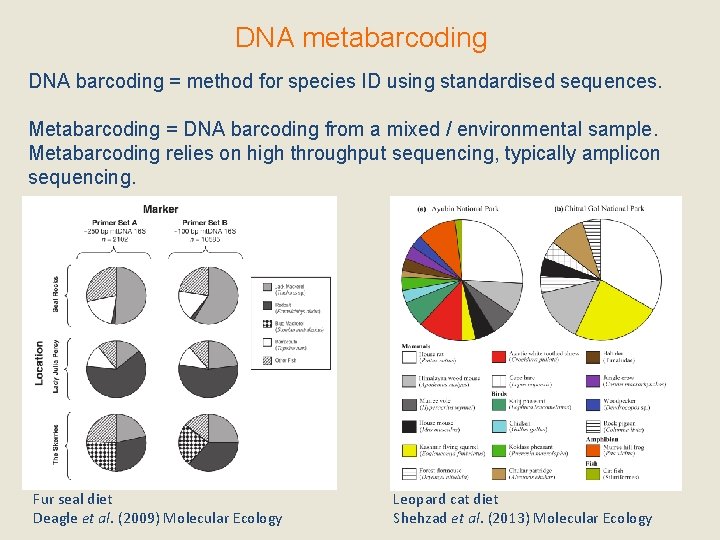
DNA metabarcoding DNA barcoding = method for species ID using standardised sequences. Metabarcoding = DNA barcoding from a mixed / environmental sample. Metabarcoding relies on high throughput sequencing, typically amplicon sequencing. Fur seal diet Deagle et al. (2009) Molecular Ecology Leopard cat diet Shehzad et al. (2013) Molecular Ecology

Metabarcoding from the Tasmanian scat collection A different kind of challenge: • Can we get sufficient DNA from old scats? • Can we ID unknown predators? • Can we detect prey DNA without predator blocking primers? • Which are appropriate markers for Australian fauna (no reference sequences for many taxa)? • How can we make this cost effective & practical for >10, 000 scats?
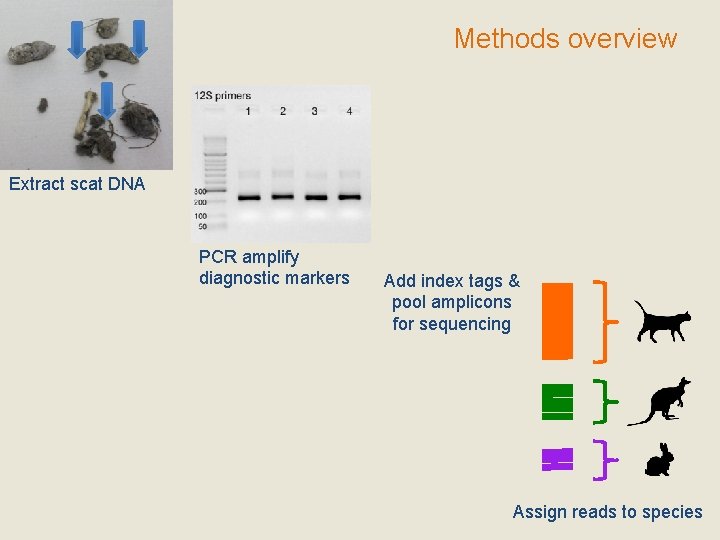
Methods overview Extract scat DNA PCR amplify diagnostic markers Add index tags & pool amplicons for sequencing Assign reads to species

Illumina Mi. Seq pilot: 5 primer sets from 4 mt. DNA genes 12 known-origin scats (Some knowledge of diets) Fox, cat, dog, devil, eastern quoll, spotted-tail quoll 17 Tasmanian fox-positive scats Collected in Tasmania 2008 -2010 Tested fox-positive using cytb PCR and sequencing 14 scats genotyped using 8 fox microsatellites (all genotypes unique) 18 Tasmanian fox-negative scats Most collected in Tasmania 2008 -2011 Mammal DNA detected using 12 S primers, but no fox DNA detected One scat collected Maria Island 2013 - devil scat from insurance population 4 DNA mixtures Good quality DNA from tissue samples Includes mammal, reptile, bird and invertebrate DNA, no predator DNA
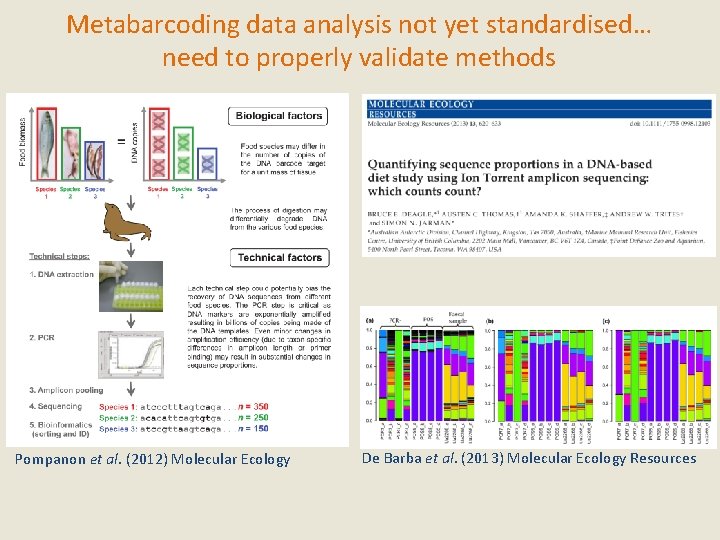
Metabarcoding data analysis not yet standardised… need to properly validate methods Pompanon et al. (2012) Molecular Ecology De Barba et al. (2013) Molecular Ecology Resources
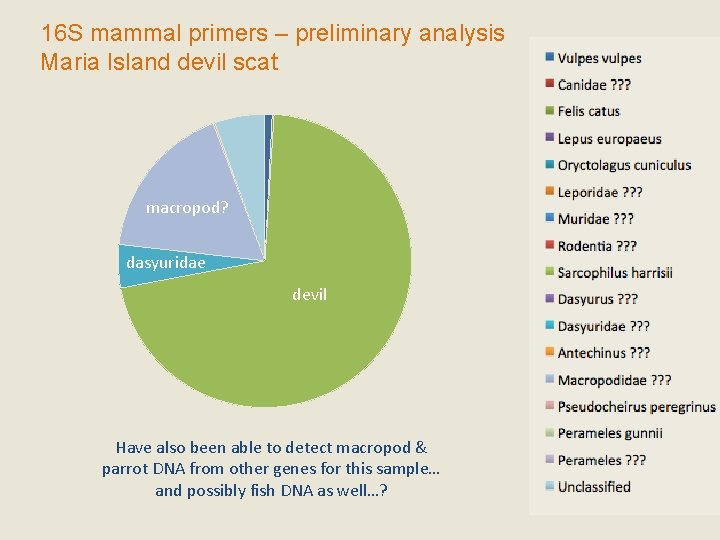
16 S mammal primers – preliminary analysis Maria Island devil scat macropod? dasyuridae devil Have also been able to detect macropod & parrot DNA from other genes for this sample… and possibly fish DNA as well…?
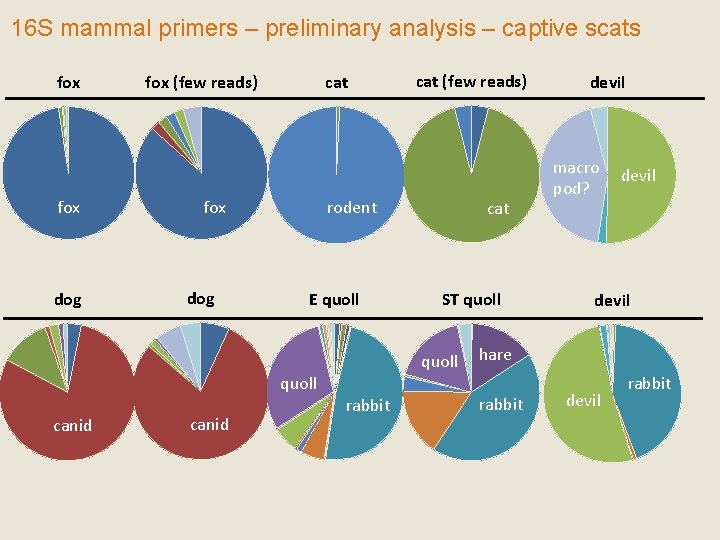
16 S mammal primers – preliminary analysis – captive scats fox dog fox (few reads) cat fox dog cat (few reads) rodent E quoll cat ST quoll canid rabbit devil macro pod? devil hare rabbit devil rabbit
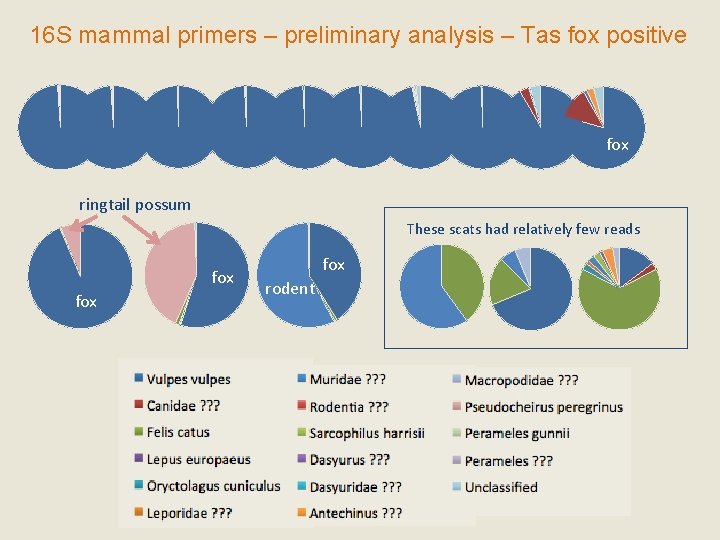
16 S mammal primers – preliminary analysis – Tas fox positive fox ringtail possum These scats had relatively few reads fox fox rodent
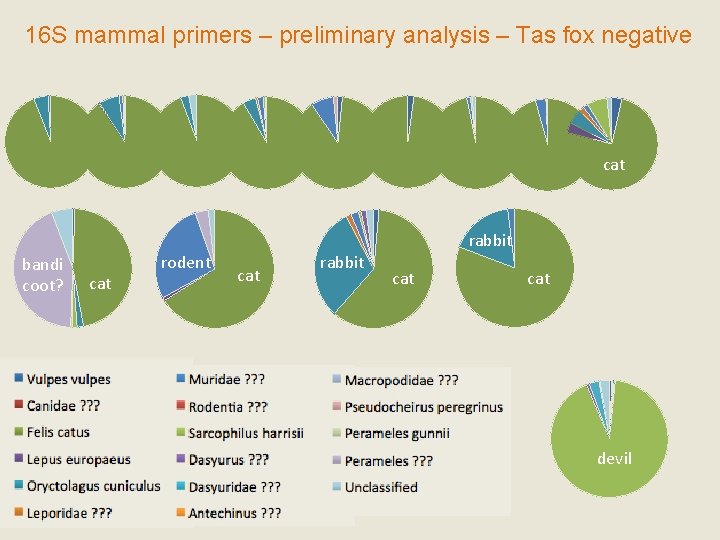
16 S mammal primers – preliminary analysis – Tas fox negative cat bandi coot? cat rodent rabbit cat cat devil
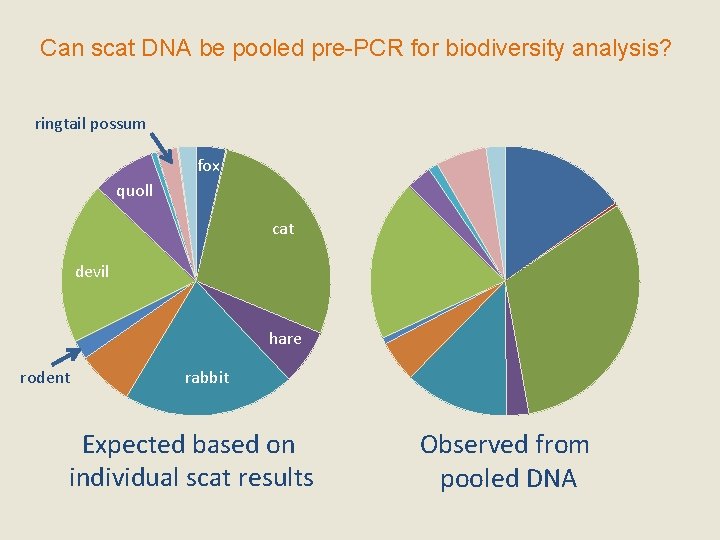
Can scat DNA be pooled pre-PCR for biodiversity analysis? ringtail possum fox quoll cat devil hare rodent rabbit Expected based on individual scat results Observed from pooled DNA

Metabarcoding from the Tasmanian scat collection Challenges: • Can we get sufficient DNA from old scats? – Yes, 46/48 scats successfully amplified & most sequenced • Can we ID unknown predators? – Yes, predator DNA recovered from most scats • Can we detect prey DNA without predator blocking primers? – Yes, but need to work on limits of detection • Which are appropriate markers for Australian fauna (no reference sequences for many taxa)? – Work in progress – depends on question & context – In process of generating more reference sequences • How much can we multiplex – to make this affordable as a management tool? – Work in progress – again depends on questions

Acknowledgements UC • • Catriona Campbell Richard Duncan Bernd Gruber Elodie Modave • • Max Powell Sumaiya Quasim Kristin Stewardson Kejun Wei Matt Young Rachel Annetts Pia Salgado Collections • ANWC • QVMAG • SA Museum • TMAG Illumina Mi. Seq Pilot Project funding & AGRF DPIPWE • Candida Barclay • Peter Cremasco • Elise Dewar • Simon Fearn • • Stephen Harris Dean Heinz Tom Priestley FEP staff CSIRO • Oliver Berry • Chris Hardy • Leo Joseph • Matt Morgan • Robert Palmer
- Slides: 17