Special Topics in Genomics Lecture 1 Introduction Instructor





![SNP array ACCGTGGA[C/T]CTGAACCG |||| | |||| TGGCACCT[G/A]GACTTGGC ACCGTGGA[G]CTGAACCGTGGA[C]CTGAACCGTGGA[T]CTGAACCGTGGA[A]CTGAACCG What will happen when the genotype SNP array ACCGTGGA[C/T]CTGAACCG |||| | |||| TGGCACCT[G/A]GACTTGGC ACCGTGGA[G]CTGAACCGTGGA[C]CTGAACCGTGGA[T]CTGAACCGTGGA[A]CTGAACCG What will happen when the genotype](https://slidetodoc.com/presentation_image_h2/a3e4cf37ac9855c00e765aab57217203/image-25.jpg)




- Slides: 36

Special Topics in Genomics Lecture 1: Introduction Instructor: Hongkai Ji Department of Biostatistics Email: hji@jhsph. edu

Outline of today’s lecture n Introduction to genome and genomics n Topics and tools n Relevance of statistics

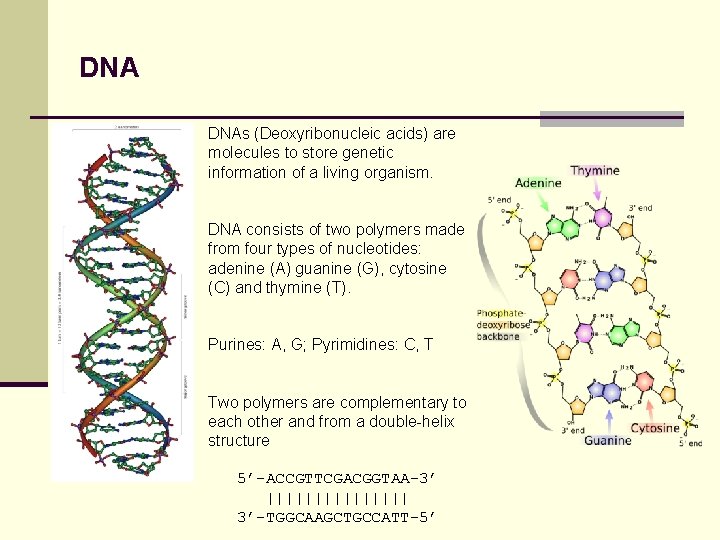
DNA DNAs (Deoxyribonucleic acids) are molecules to store genetic information of a living organism. DNA consists of two polymers made from four types of nucleotides: adenine (A) guanine (G), cytosine (C) and thymine (T). Purines: A, G; Pyrimidines: C, T Two polymers are complementary to each other and from a double-helix structure 5’-ACCGTTCGACGGTAA-3’ |||||||| 3’-TGGCAAGCTGCCATT-5’

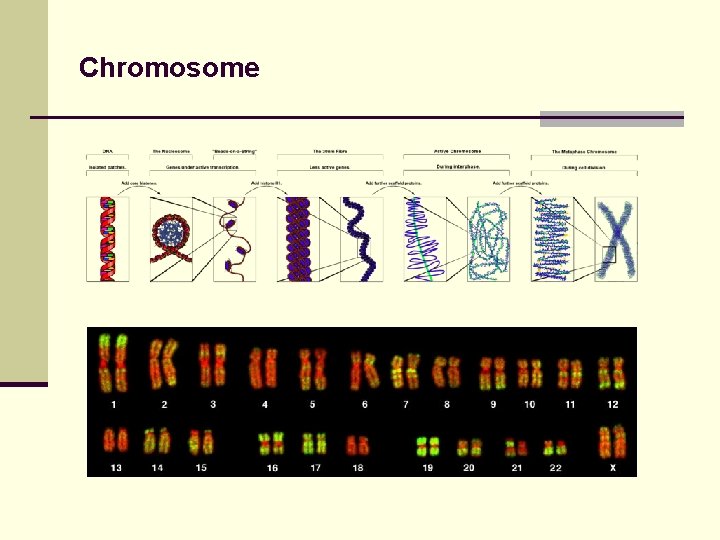
Chromosome

Genome TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCT CTCACACCTGACATGAAAAGGCACATGAGGATCCTCAAATACCCCGTGATCAGTCTCAGGGTAGCTCTCATAGCCTGGACA GGGCCCCCCTCGGGGGTTGCGCCCAGGTCCAGGCGGGGGATGCACAGCAACAGTCACCGAAGCAGAAGCCGTCACAGTGGT GATGGGCTGGCAGTAGCTGGGCACAGAGCTGCCCATGGCGGTGGACGTTGGGTTCCGAGGGTTGTGAGAACGGGCCCCACG GGGCCCTGAGCGGTCCCTATTGCTAGGGCCAGAATGCCCTTCAGTAGAAATTTCAAAAGCGTCTCTGCGCGGTCTGTAGGG GGGTGGCCGCAAGCCTTCTCTAGGGGGATCCCTTCGAGGCTGCTGGCCTTGCCGTCCAGGGGACAAGGAGCCAGAGTCCAG GTGGGGCTGTTGCCGAGGGGTCAAGGGAGGCTGATGTCTGGAGTCCGGATGGACCACCTGCAGAGGAGAGACATAGGTCAA CACAGGGAGGTAGGATGGTGGTGATGTTCCACAAAAGAAAACCTATTCCTTTAGAAACCTCCAGGATGTGAATCCTGCACAGCTGGAGGCATATAGCCACTGCCCATAGATCTCAACTTACCCTCACAACTGCCCCCAGG CCTAAGTTCTCTGCCTCAAAACTGCCAAGGCCTGGATAGCCAAGAGCCTGGGTGTCTTGGAAATATGCAACCATAAATAGT AGCTTTTAGAAGTATAAGGCTCCTGTTTCTGGGTCATATTAGTGTTGTTTTCACCTGTCCCCAGCCCTAAGCCAGGTGTGG CCAGAAGCAAATGTACTGTAAGAGCAAAAACTTCCACACAGATAGTTCTGTTAGGCAATACATCTCTGCCTGACTA TTAGGAATCTGGTTTCTGGGTCCTCTGTACAAAGCTCGGAGCAACACAGTGGCCACATCAAAAGGACCGTGACCAAC TTCAAAGTCGGTGAGCTTGTACCTATTTTTAGGCTCCTGCTGAACAGAACCAGATTCACACTACAGCTCAGCAGGGCATCG TCACGGGTGTGTGTGTGTGTGTGTGTTGGGGGGTGGACAGAGGACGGGGACACAATT CACTGGCCAGCCCTTCTCTCCTTCAAGGCTGCTCTAGCCTGGGACTGGAATACACATTTCCTGTAAACATGGTGGGG GCCTCAGGCAAGCCAGAGTTTTGGAGCCTTAACTCTTCAAGGTGAGCATCTTGACTTGGAGGGTGGGGGTGCGGGTA AGGAACCTGTGGACTCCTCCCTACAAGACAGAAAAGGAATAAGCCACGAAGACAATAACGATTTTTGTATCAAGCGT CCTCTCCCATTTCAGCTTACCTGACAATGAAATCAAATTCGGACCCTGCAAGCATCAGTACACCCAGCAGAGTGGACACAG CACCGTCCAGAACGGGAGCAAACATGTGCTCCAGAGCATAGCCCTGTGGTTCTTGTCCCCAATGGCTGTCAGAAAGG CCTGAACAAAGGAGAAAATTGACACGGTCACATTCTGGGTGTGGTAAAGTGCTCAGCTGTGTCTATACTTGGGTTTTGTAT… Total amount of DNA in human genome: 3 * 10 9 base pairs (bp)

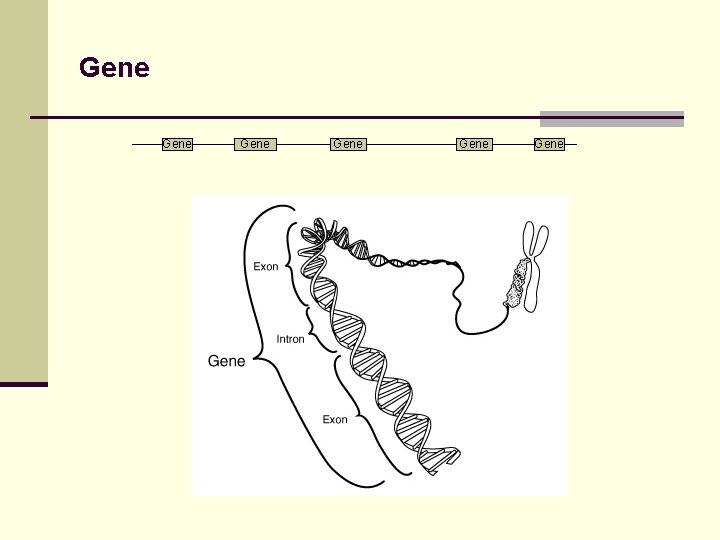
Gene Gene

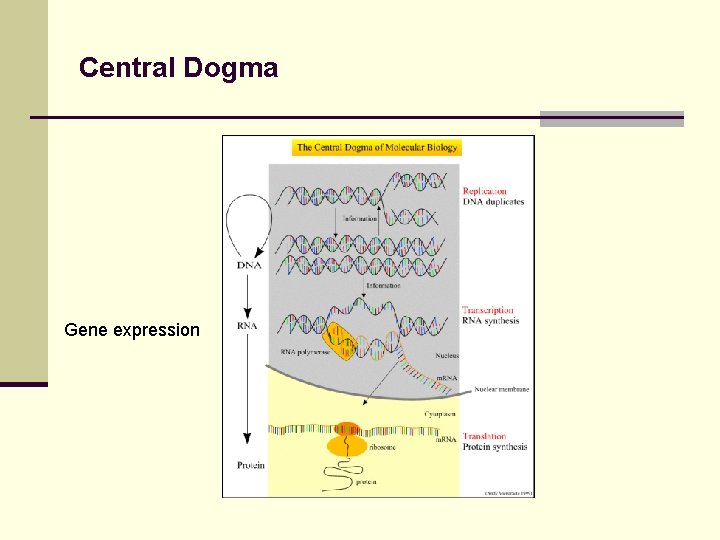
Central Dogma Gene expression

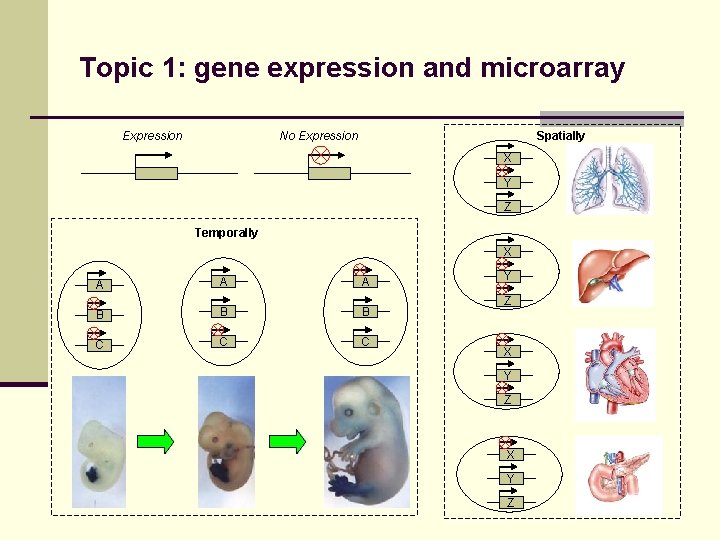
Topic 1: gene expression and microarray Expression No Expression Spatially X Y Z Temporally X A A A B B B C C C Y Z X Y Z

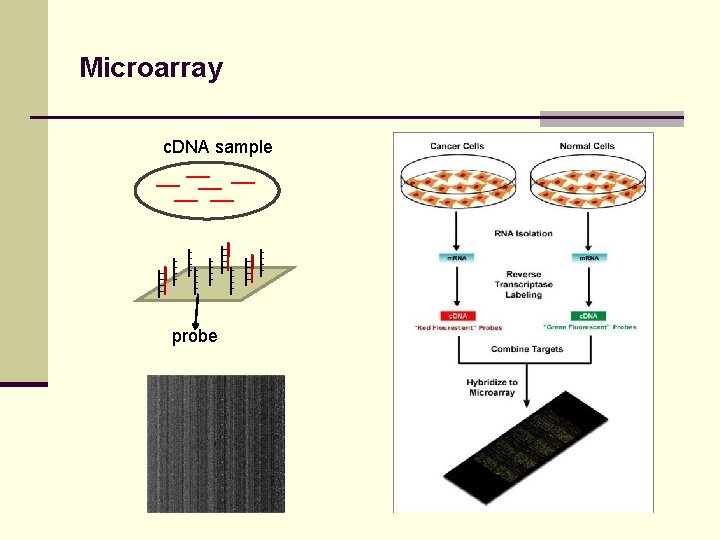
Microarray c. DNA sample probe

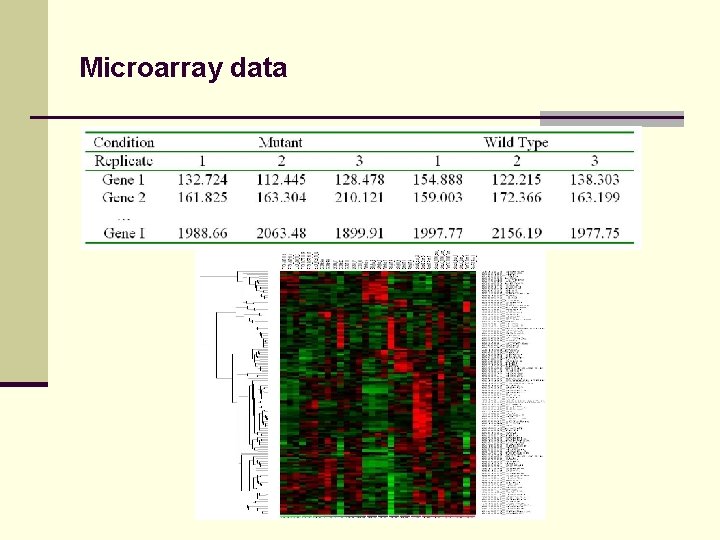
Microarray data

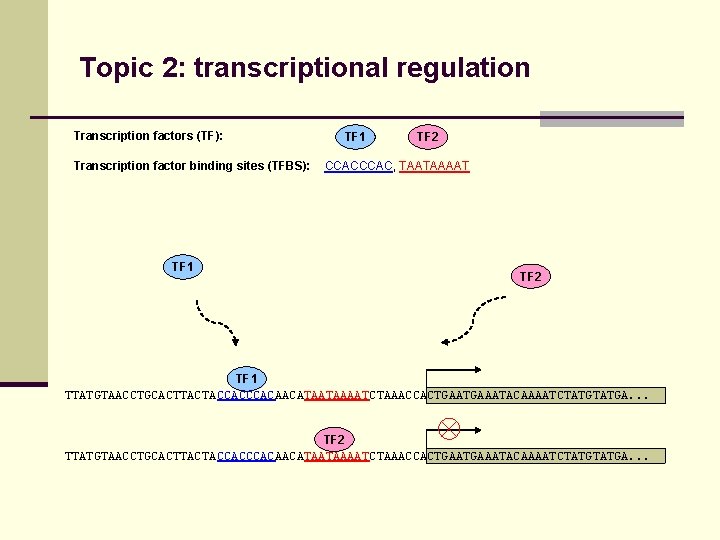
Topic 2: transcriptional regulation Transcription factors (TF): Transcription factor binding sites (TFBS): TF 1 TF 2 CCAC, TAATAAAAT TF 2 TF 1 TTATGTAACCTGCACTTACTACCACAACATAATAAAATCTAAACCACTGAAATACAAAATCTATGA. . . TF 2 TTATGTAACCTGCACTTACTACCACAACATAATAAAATCTAAACCACTGAAATACAAAATCTATGA. . .

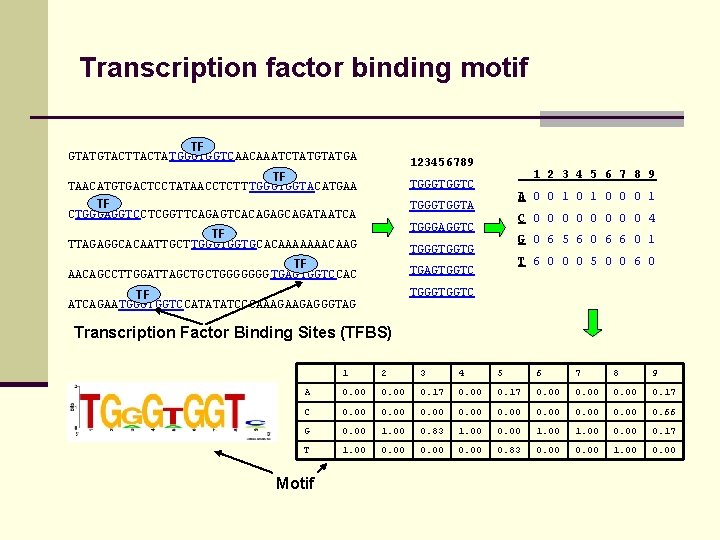
Transcription factor binding motif TF GTATGTACTATGGGTGGTCAACAAATCTATGA 123456789 TF TAACATGTGACTCCTATAACCTCTT TGGGTGGTACATGAA TF CTGGGAGGTCCTCGGTTCAGAGTCACAGAGCAGATAATCA 1 2 3 4 5 6 7 8 9 TGGGTGGTC A 0 0 1 0 0 0 1 TGGGTGGTA C 0 0 0 0 4 TGGGAGGTC TF TTAGAGGCACAATTGCTTGGGTGGTGCACAAAAAAACAAG G 0 6 5 6 0 6 6 0 1 TGGGTGGTG TF AACAGCCTTGGATTAGCTGCTGGGGGGG TGAGTGGTCCAC T 6 0 0 0 5 0 0 6 0 TGAGTGGTC TGGGTGGTC TF ATCAGAATGGGTGGTCCATATATCCCAAAGAAGAGGGTAG Transcription Factor Binding Sites (TFBS) 1 2 3 4 5 6 7 8 9 A 0. 00 0. 17 C 0. 00 0. 66 G 0. 00 1. 00 0. 83 1. 00 0. 00 1. 00 0. 17 T 1. 00 0. 83 0. 00 1. 00 0. 00 Motif

Motifs are regulatory codes in the genome TCAGTTGGAGCTGCTCCCCCACGGCCTCTCCTCACATTCCACGTCCTGTAGCTCTATGACCTCCACCTTTGAGTCCCTCCT CTCACACCACCCATGTTTATGAGGATCCTCAAATACCCCGTGATCAGTCTCAGGGTAGCTCTCATAGCCTGGACAG GGCCCCCCTCGGGGGTTGCGCCCAGGTCCAGGCGGGGGATGCACAGCAACAGTCACCGAAGCAGAAGCCGTCACAGTGGTG ATGGGCTGGCAGTAGCTGGGCACAGAGCTGCCCATGGCGGTGGACGTTGGGTTCCGAGGGTTGTGAGAACGGGCCCCACGG GGCCCTGAGCGGTCCCTATTGCTAGGGCCAGAATGCCCTTCAGTAGAAATTTCAAAAGCGTCTCTGCGCGGTCTGTAGGGG GGTGGCCGCAAGCCTTCTCTAGGGGGATC CCTTCGTTGCTGCTGGCCTTGCCGTCCAGGGGACAAGGAGCCAGAGTCCAGG TGGGGCTGTTGCCGAGGGGTCAAGGGAGGCTGATGTCTGGAGTCCGGATGGACCACCTGCAGAGGAGAGACATAGGTCAAC ACAGGGAGGTAGGATGGTGGTGATGTTCCACAAAAGAAAACCTATTCCTTTAGAAACCTCCAGGATGTGAATCCTGCACAGCTGGAGGCATATAGCCACTGCCCATAGATCTCAACTTACCCTCACAACTGCCCCCAGGC CTAAGTTCTCTGCCTCAAAACTGCCAAGGCCTGGATAGCCAAGAGCCTGGGTGTCTTGGAAATATGCAACCATAAATAGTA GCTTTTAGAAGTATAAGGCTCCTGTTTCTGGGTCATATTAG TTTTGTTTTCACCTGTCCCCATAAGCCAGGTGTGGC CAGAAGCAAATGTACTGTAAGAGCAAAAACTTCCACACAGATAGTTCTGTTAGGCAATACATCTCTGCCTGACTAT TAGGAATCTGGTTTCTGGGTCCTCTGTACAAAGCTCGGAGCAACACAGTGGCCACATCAAAAGGACCGTGACCAACT TCAAAGTCGGTGAGCTTGTACCTATTTTTAGGCTCCTGCTGAACAGAACCAGATTCACACTACAGCTCAGCAGGGCATCGT CACGGGTGTGTGTGTGTGTGTGTGTTGGGGGGTGGACAGAGGACGGGGACACAATTC ACTGGCCAGCCCTTCTCTCCTTCAAGGCTGCTCTAGCCTGGGACTGGAATACACATTTCCTGTAAACATGGTGGGGG CCTCAGGCAAGCCAGAGTTTTGGAG CCTTAACTCTTCAAGGTGAGCATCTTGACTTGGAGGGTGGGGGTGCGGGTAA GGAACCTGTGGACTCCACCCAACAAGACAGAAAAGGAATAAGCCACGAAGACAATAACGATTTTTGTATCAAGCGTC CTCTCCCATTTCAGCTTACCTGACAATGAAATCAAATTCGGACCCTGCAAGCATCAGTACACCCAGCAGAGTGGACACAGC ACCGTCCAGAACGGGAGCAAACATGTGCTCCAGAGCATAGCCCTGTGGTTCTTGTCCCCAATGGCTGTCAGAAAGGC CTGAACAAAGGAGAAAATTGACACGGTCACATTCTGGGTGTGGTAAAGTGCTCAGCTGTGTCTATACTTGGGTTTTGTAT Transcription Factor Binding Sites (TFBS) Gene

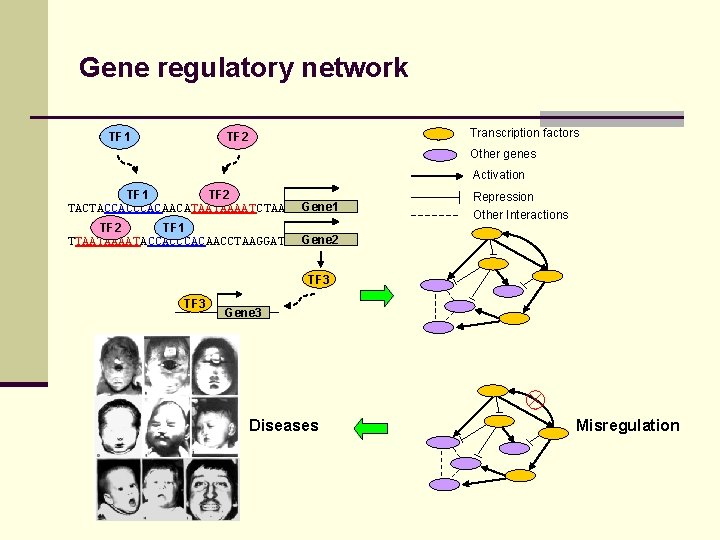
Gene regulatory network TF 1 Transcription factors TF 2 Other genes Activation TF 1 TF 2 TACTACCACAACATAATAAAATCTAA Gene 1 TF 2 TF 1 TTAATAAAATACCACAACCTAAGGAT Gene 2 Repression Other Interactions TF 3 Gene 3 Diseases Misregulation

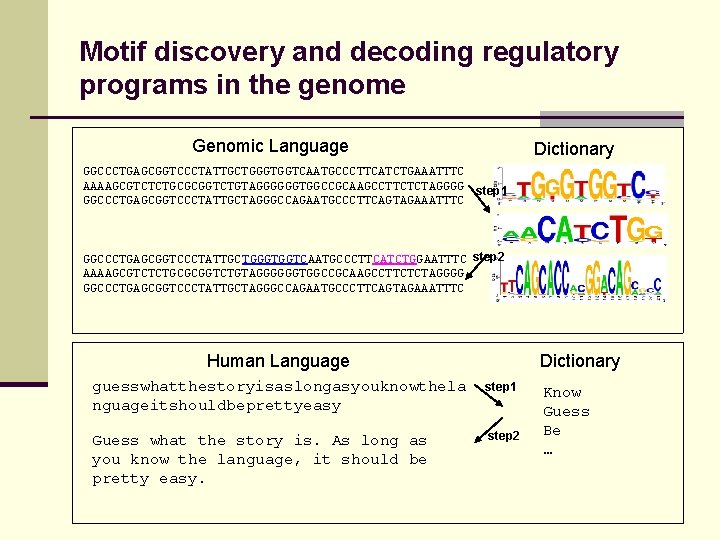
Motif discovery and decoding regulatory programs in the genome Genomic Language Dictionary GGCCCTGAGCGGTCCCTATTGCTGGGTGGTCAATGCCCTTCATCTGAAATTTC AAAAGCGTCTCTGCGCGGTCTGTAGGGGGGTGGCCGCAAGCCTTCTCTAGGGG step 1 GGCCCTGAGCGGTCCCTATTGCTAGGGCCAGAATGCCCTTCAGTAGAAATTTC GGCCCTGAGCGGTCCCTATTGCTGGGTGGTCAATGCCCTTCATCTGGAATTTC step 2 AAAAGCGTCTCTGCGCGGTCTGTAGGGGGGTGGCCGCAAGCCTTCTCTAGGGG GGCCCTGAGCGGTCCCTATTGCTAGGGCCAGAATGCCCTTCAGTAGAAATTTC Human Language Dictionary guesswhatthestoryisaslongasyouknowthela nguageitshouldbeprettyeasy step 1 Guess what the story is. As long as you know the language, it should be pretty easy. step 2 Know Guess Be …

Finding motifs from co-regulated genes (Roth et al. , 1998; Hughes et al. , 2000; etc. ) GTATGTACTATGGGTGGTCAACAAATCTATGA Gene 1 GTATGTACTATGGGTGGTCAACAAATCTATGA CTGGGAGGTCCTCGGTTCAGAGTCACAGAGCAGATAATCA Gene 2 CTGGGAGGTCCTCGGTTCAGAGTCACAGAGCAGATAATCA TAACATGTGACTCCTATAACCTCTTTGGGTGGTACATGAA Gene 3 TAACATGTGACTCCTATAACCTCTTTGGGTGGTACATGAA Condition 1 Gene 2 Gene 3 … Gene N Condition 2

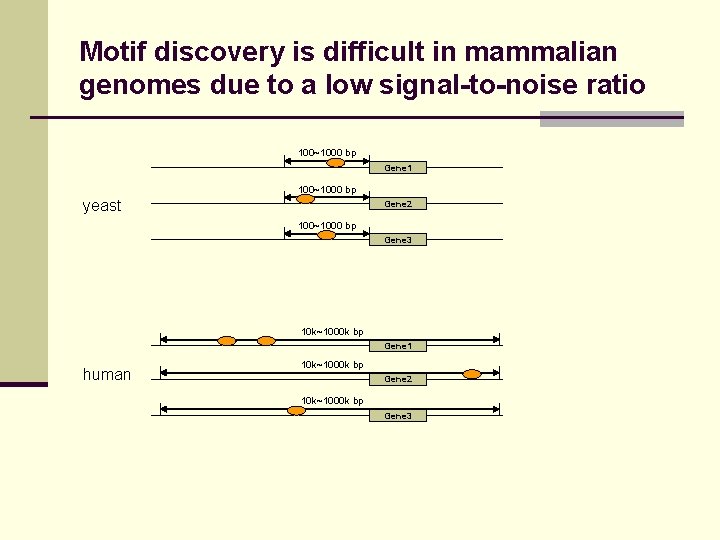
Motif discovery is difficult in mammalian genomes due to a low signal-to-noise ratio 100~1000 bp Gene 1 100~1000 bp yeast Gene 2 100~1000 bp Gene 3 10 k~1000 k bp Gene 1 human 10 k~1000 k bp Gene 2 10 k~1000 k bp Gene 3

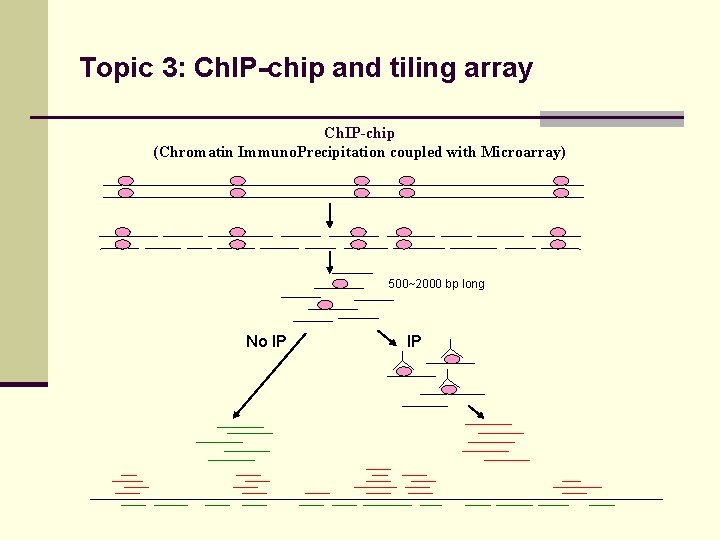
Topic 3: Ch. IP-chip and tiling array Ch. IP-chip (Chromatin Immuno. Precipitation coupled with Microarray) 500~2000 bp long No IP IP

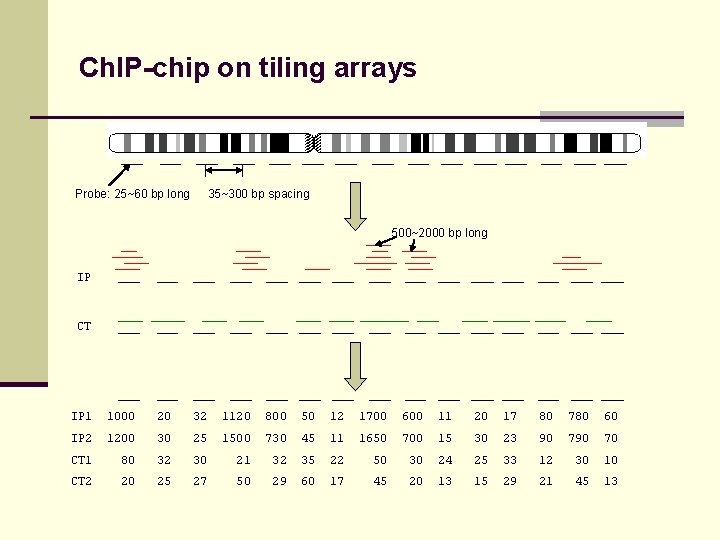
Ch. IP-chip on tiling arrays Probe: 25~60 bp long 35~300 bp spacing 500~2000 bp long IP CT IP 1 1000 20 32 1120 800 50 12 1700 600 11 20 17 80 780 60 IP 2 1200 30 25 1500 730 45 11 1650 700 15 30 23 90 70 CT 1 80 32 30 21 32 35 22 50 30 24 25 33 12 30 10 CT 2 20 25 27 50 29 60 17 45 20 13 15 29 21 45 13

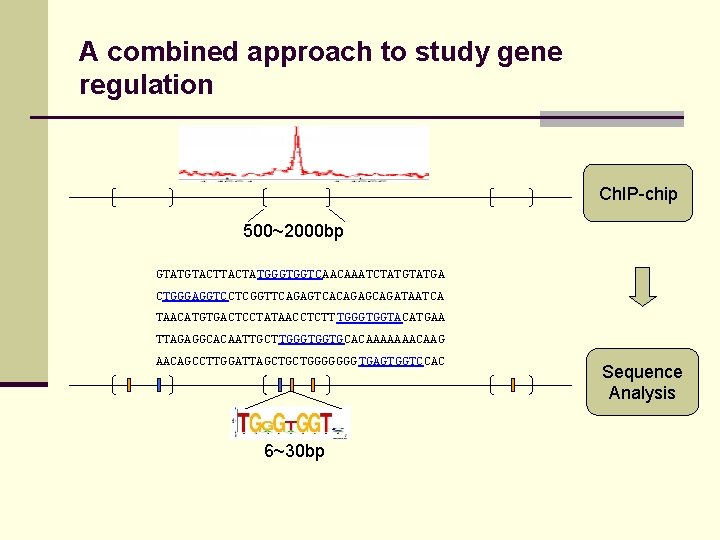
A combined approach to study gene regulation Ch. IP-chip 500~2000 bp GTATGTACTATGGGTGGTCAACAAATCTATGA CTGGGAGGTCCTCGGTTCAGAGTCACAGAGCAGATAATCA TAACATGTGACTCCTATAACCTCTT TGGGTGGTACATGAA TTAGAGGCACAATTGCTTGGGTGGTGCACAAAAAAACAAG AACAGCCTTGGATTAGCTGCTGGGGGGG TGAGTGGTCCAC 6~30 bp Sequence Analysis

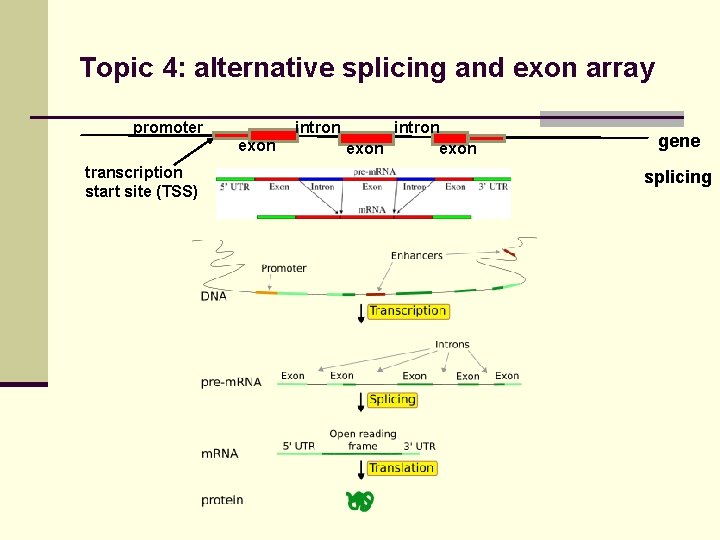
Topic 4: alternative splicing and exon array promoter intron exon transcription start site (TSS) intron exon gene splicing

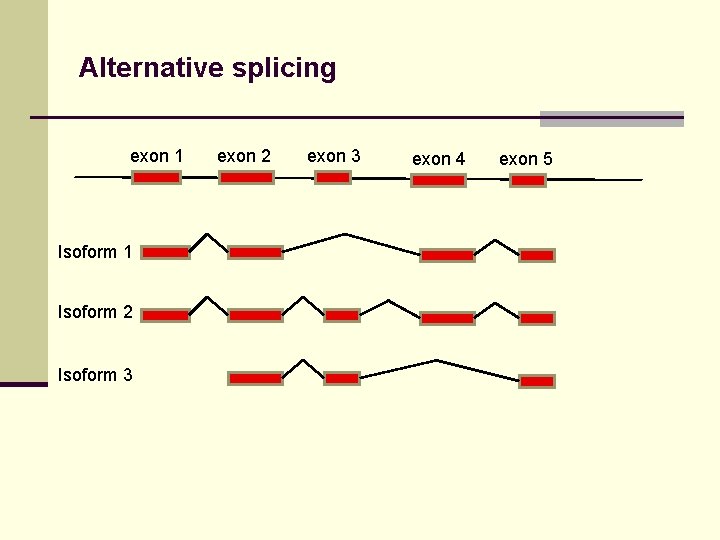
Alternative splicing exon 1 Isoform 2 Isoform 3 exon 2 exon 3 exon 4 exon 5

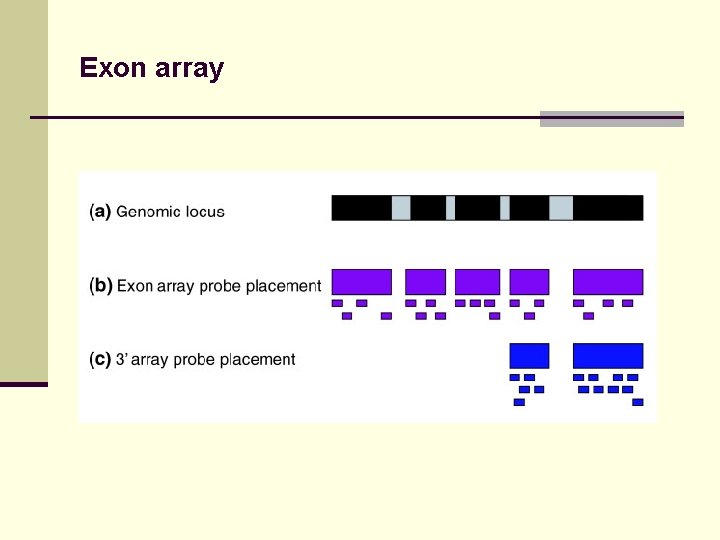
Exon array

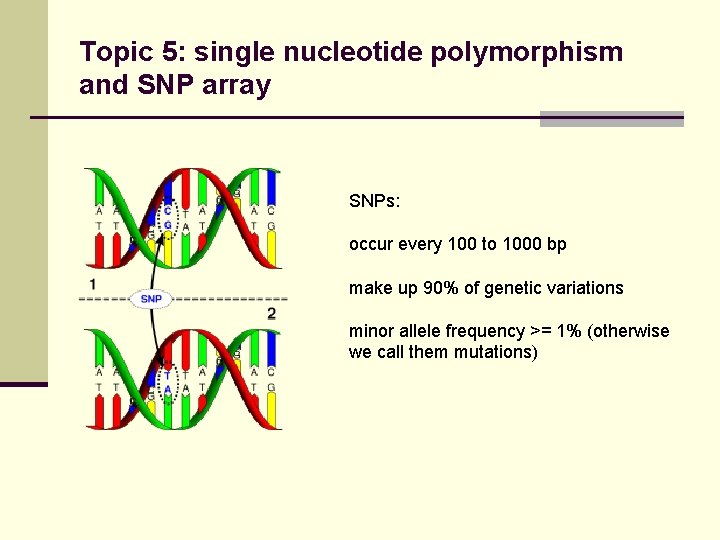
Topic 5: single nucleotide polymorphism and SNP array SNPs: occur every 100 to 1000 bp make up 90% of genetic variations minor allele frequency >= 1% (otherwise we call them mutations)
![SNP array ACCGTGGACTCTGAACCG TGGCACCTGAGACTTGGC ACCGTGGAGCTGAACCGTGGACCTGAACCGTGGATCTGAACCGTGGAACTGAACCG What will happen when the genotype SNP array ACCGTGGA[C/T]CTGAACCG |||| | |||| TGGCACCT[G/A]GACTTGGC ACCGTGGA[G]CTGAACCGTGGA[C]CTGAACCGTGGA[T]CTGAACCGTGGA[A]CTGAACCG What will happen when the genotype](https://slidetodoc.com/presentation_image_h2/a3e4cf37ac9855c00e765aab57217203/image-25.jpg)
SNP array ACCGTGGA[C/T]CTGAACCG |||| | |||| TGGCACCT[G/A]GACTTGGC ACCGTGGA[G]CTGAACCGTGGA[C]CTGAACCGTGGA[T]CTGAACCGTGGA[A]CTGAACCG What will happen when the genotype is CC? CT? TT? Applications: 1. Genotyping & genome-wide association study 2. Copy number variations and loss of heterozygosity 3. Allele specific expression …

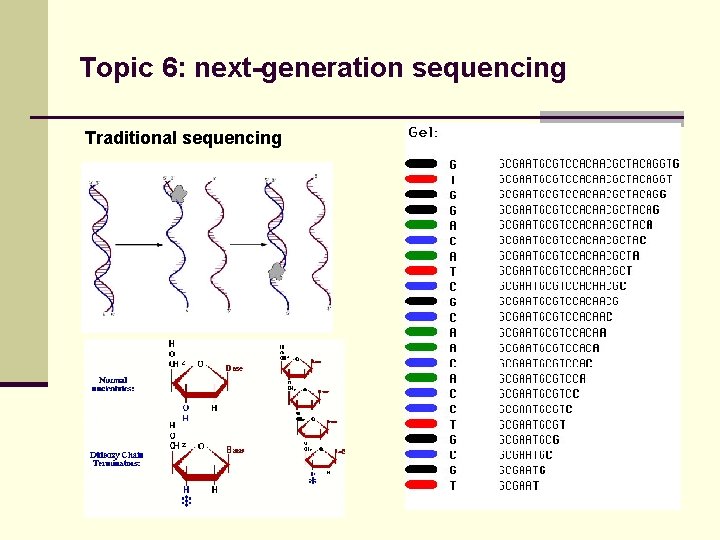
Topic 6: next-generation sequencing Traditional sequencing

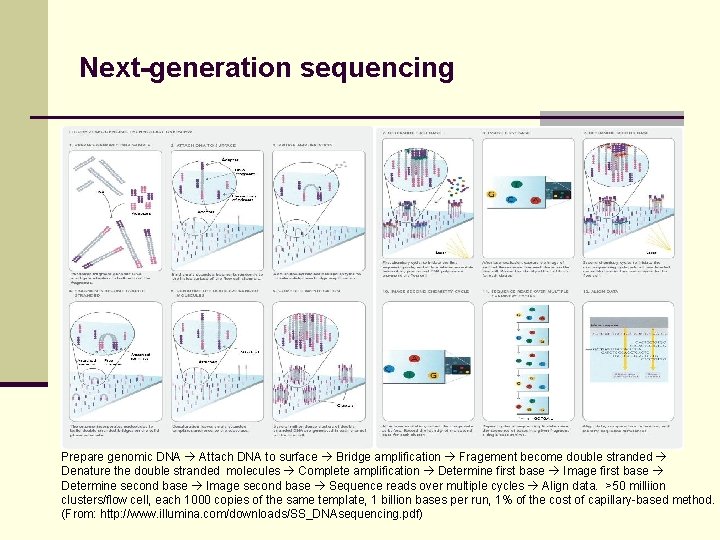
Next-generation sequencing Prepare genomic DNA Attach DNA to surface Bridge amplification Fragement become double stranded Denature the double stranded molecules Complete amplification Determine first base Image first base Determine second base Image second base Sequence reads over multiple cycles Align data. >50 milliion clusters/flow cell, each 1000 copies of the same template, 1 billion bases per run, 1% of the cost of capillary-based method. (From: http: //www. illumina. com/downloads/SS_DNAsequencing. pdf)

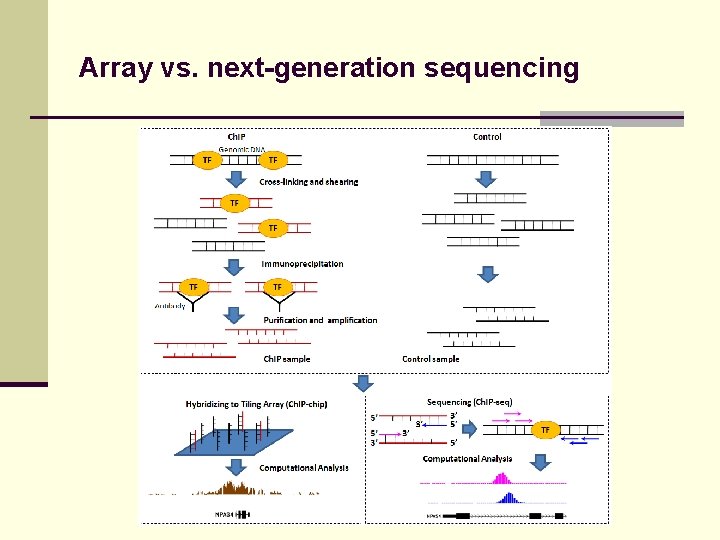
Array vs. next-generation sequencing

Array vs. next-generation sequencing Microarray, Exon array RNA-seq Ch. IP-chip Ch. IP-seq SNP array SNP/mutation detection by sequencing … …

Other topics n Epigenomics n Transposon n mi. RNA

Relevance of statistics Need new statistical theories and tools Genomics Statistics Guide development of efficient data analysis strategies

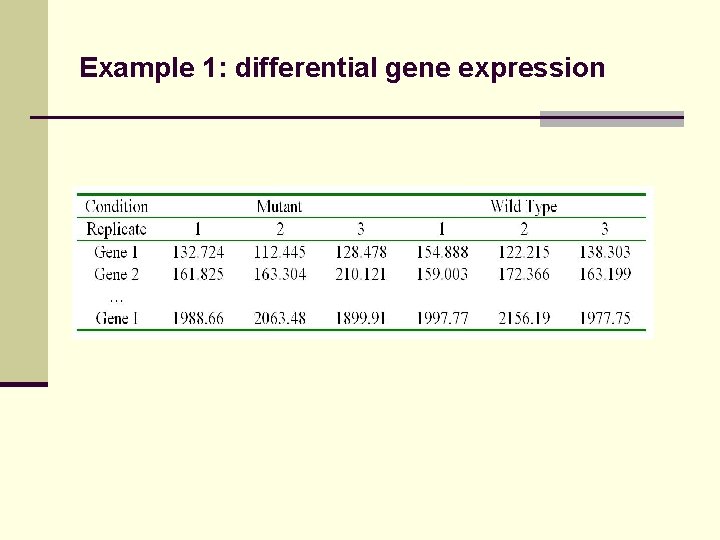
Example 1: differential gene expression

Example 1: multiple testing Gene t-statistic p-value i=1 1. 2 0. 30 i=2 6. 7 0. 001 i=3 5. 1 0. 002 … … … i=I -0. 56 Bonferroni adjustment Rejections … Multiplicity needs to be adjusted in order to determine statistical significance Bonferroni adjustment too stringent False discovery rate

False discovery rate (FDR) False discovery rate (FDR, Benjamini & Hochberg, 1995) Accept Reject Total True H 0 U V m 0 True H 1 T S m-m 0 m-R R m FDR = E(V/R) = Pr(R>0)E(V/R|R>0) FWER = Pr(V ≥ 1)

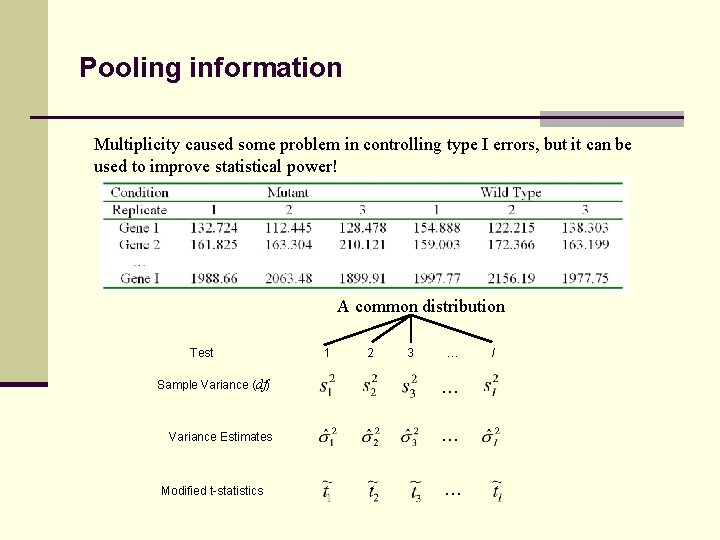
Pooling information Multiplicity caused some problem in controlling type I errors, but it can be used to improve statistical power! A common distribution Test 1 2 3 … Sample Variance (df) … Variance Estimates … Modified t-statistics … I

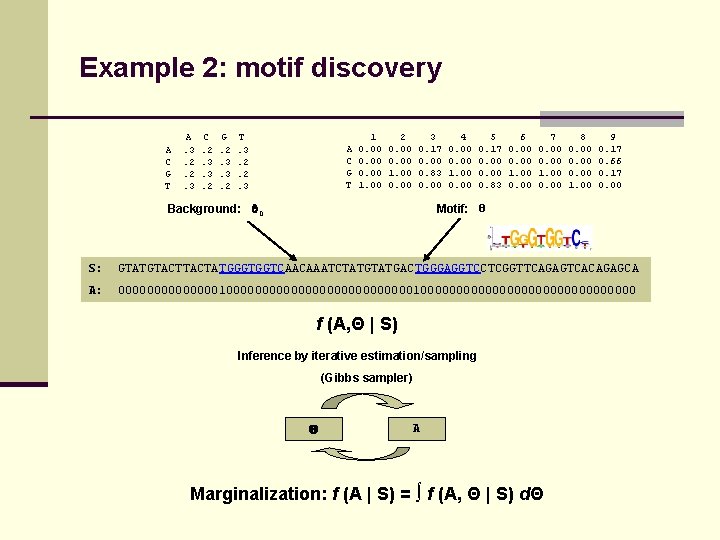
Example 2: motif discovery A C G T A. 3. 2. 2. 3 C. 2. 3. 3. 2 G. 2. 3. 3. 2 T. 3. 2. 2. 3 A C G T 1 0. 00 1. 00 2 0. 00 1. 00 0. 00 3 0. 17 0. 00 0. 83 0. 00 Background: 0 4 0. 00 1. 00 0. 00 5 0. 17 0. 00 0. 83 6 0. 00 1. 00 0. 00 7 0. 00 1. 00 0. 00 8 0. 00 1. 00 9 0. 17 0. 66 0. 17 0. 00 Motif: Θ S: GTATGTACTATGGGTGGTCAACAAATCTATGACTGGGAGGTCCTCGGTTCAGAGTCACAGAGCA A: 000000000000001000000000000000000000 f (A, Θ | S) Inference by iterative estimation/sampling (Gibbs sampler) A Marginalization: f (A | S) = ∫ f (A, Θ | S) dΘ