Solving Unknown Structures Using NMR Organic Structure Analysis









![USING COSY DATA UNKNOWN H • • A compound shows an [M + H]+ USING COSY DATA UNKNOWN H • • A compound shows an [M + H]+](https://slidetodoc.com/presentation_image_h2/b3937f0e045dac5f434e79ccfb51d2c8/image-16.jpg)



- Slides: 73

Solving Unknown Structures Using NMR Organic Structure Analysis, Crews, Rodriguez and Jaspars

Six Simple Steps for Successful Structure Solution • Get molecular formula. Use combustion analysis, mass spectrum and/or 13 C NMR spectrum. Calculate double bond equivalents. • Determine functional groups from IR, 1 H and 13 C NMR • Compare 1 H integrals to number of H’s in the MF. • Determine coupling constants (J’s) for all multiplets. • Use information from 3. and 4. to construct spin systems (substructures) • Assemble substructures in all possible ways, taking account of dbe and functional groups. Make sure the integrals and coupling patterns agree with the proposed structure. Organic Structure Analysis, Crews, Rodriguez and Jaspars

USING 1 H NMR DATA UNKNOWN B • • • A compound shows an M+. in the EIMS at 154 m/z Further fragments are at 121, 93, 71, 55 and 39 m/z The IR shows bands at 3400 cm-1 (broad) & 1450 cm-1 Use the 1 H and 13 C data to determine the structure of the compound Organic Structure Analysis, Crews, Rodriguez and Jaspars

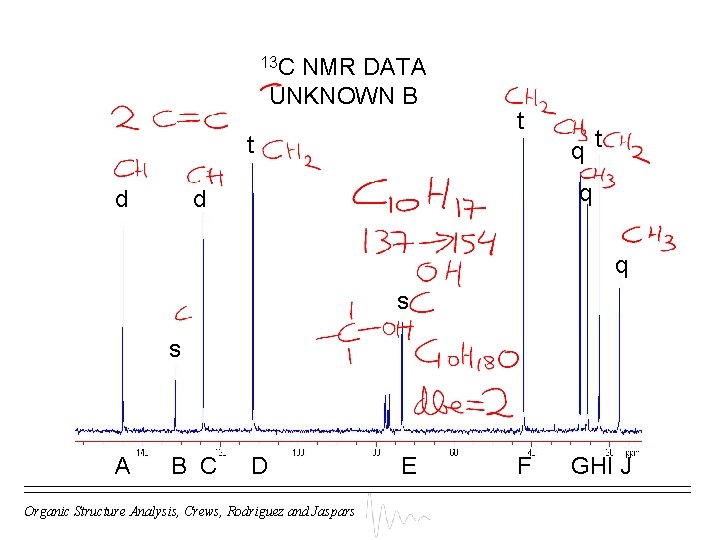
13 C NMR DATA UNKNOWN B t d t q d q s s A B C D Organic Structure Analysis, Crews, Rodriguez and Jaspars E F GHI J

MOLECULAR FORMULA DETERMINATION UNKNOWN B s d t q (C) + (CH 2) + (CH 3) = C H = Da The M+. appears at 154 m/z, so there is a mass difference of Da (= ) Therefore molecular formula = C H O ( dbe) Organic Structure Analysis, Crews, Rodriguez and Jaspars

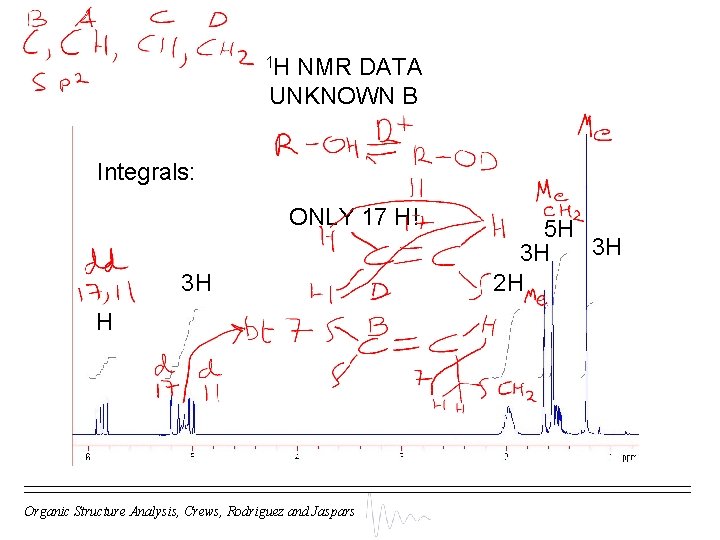
1 H NMR DATA UNKNOWN B Integrals: ONLY 17 H! 3 H H Organic Structure Analysis, Crews, Rodriguez and Jaspars 5 H 3 H 3 H 2 H

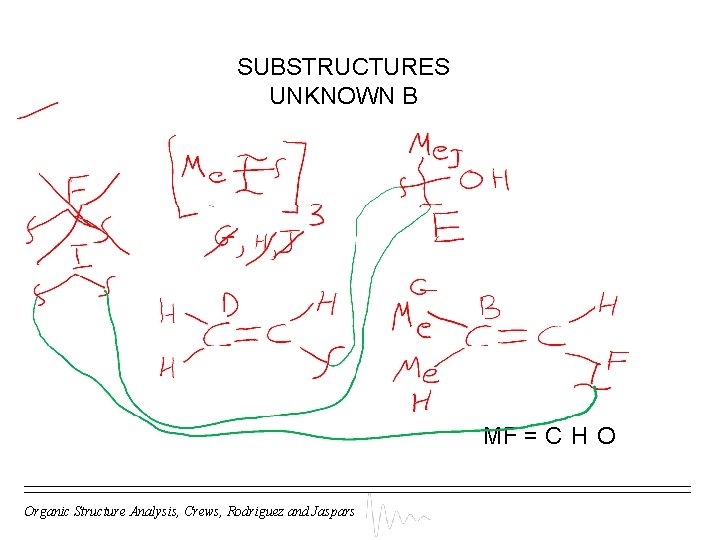
SUBSTRUCTURES UNKNOWN B MF = C H O Organic Structure Analysis, Crews, Rodriguez and Jaspars

WORKING STRUCTURES UNKNOWN B Organic Structure Analysis, Crews, Rodriguez and Jaspars

MASS SPECTRAL FRAGMENTATION UNKNOWN B Fragments at: 121, 93, 71, 55 and 39 m/z Organic Structure Analysis, Crews, Rodriguez and Jaspars

USING MASS SPECTRAL DATA UNKNOWN G • • • A compound shows an M+. in the EIMS at 128 m/z Further fragments are at 99, 83, 72 and 57 m/z The IR shows bands at 1680 cm-1 (strong) & bands at 1400 - 1500 cm-1 Use the 1 H and 13 C NMR and MS data to determine the structure of the compound Organic Structure Analysis, Crews, Rodriguez and Jaspars

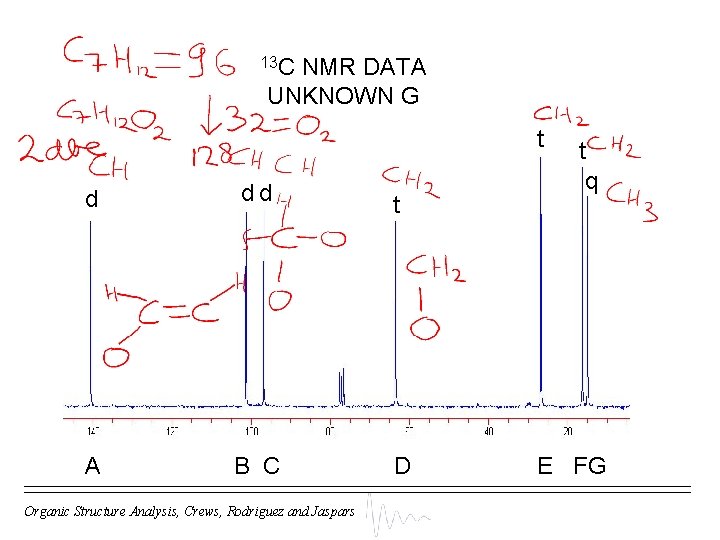
13 C NMR DATA UNKNOWN G t d dd t A B C D Organic Structure Analysis, Crews, Rodriguez and Jaspars t q E FG

MOLECULAR FORMULA DETERMINATION UNKNOWN G s d t q (C) + (CH 2) + (CH 3) = C H = Da The M+. appears at 128 m/z, so there is a mass difference of Da (= ) Therefore molecular formula = C H O ( dbe) Organic Structure Analysis, Crews, Rodriguez and Jaspars

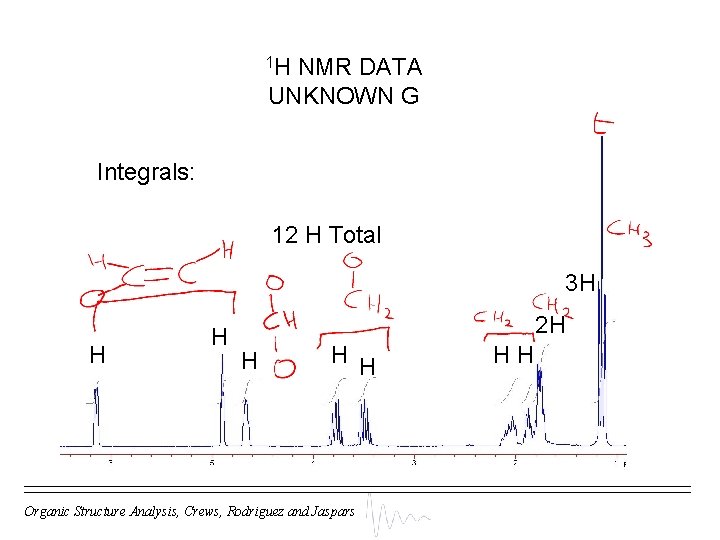
1 H NMR DATA UNKNOWN G Integrals: 12 H Total 3 H H H 2 H H Organic Structure Analysis, Crews, Rodriguez and Jaspars HH

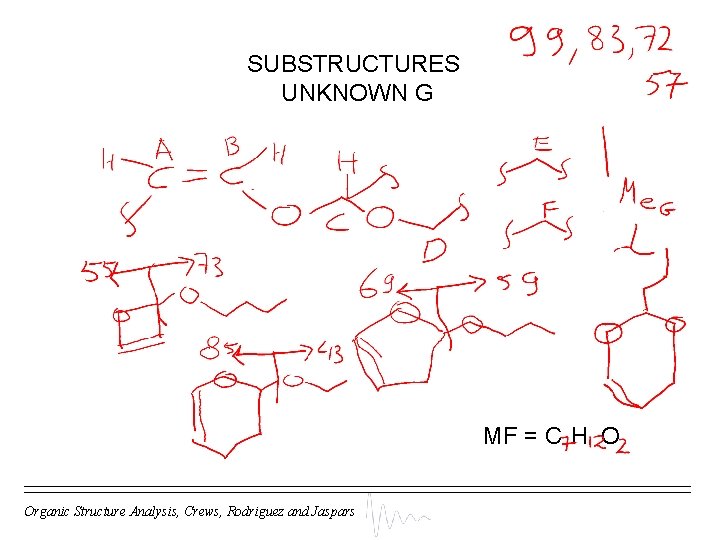
SUBSTRUCTURES UNKNOWN G MF = C H O Organic Structure Analysis, Crews, Rodriguez and Jaspars

13 C WORKING STRUCTURES UNKNOWN G Shift additivity data MS Fragmentation Retro Diels-Alder Fragments are at 99, 83, 72 and 57 m/z Organic Structure Analysis, Crews, Rodriguez and Jaspars
![USING COSY DATA UNKNOWN H A compound shows an M H USING COSY DATA UNKNOWN H • • A compound shows an [M + H]+](https://slidetodoc.com/presentation_image_h2/b3937f0e045dac5f434e79ccfb51d2c8/image-16.jpg)
USING COSY DATA UNKNOWN H • • A compound shows an [M + H]+ in the FAB MS at 132 m/z MW = 131 (Odd) therefore odd number of nitrogens A further fragment is at 86 m/z The IR shows bands at 3400 cm-1 (broad) & 1640 cm-1 (broad) Use the NMR data to determine the structure of the compound Organic Structure Analysis, Crews, Rodriguez and Jaspars

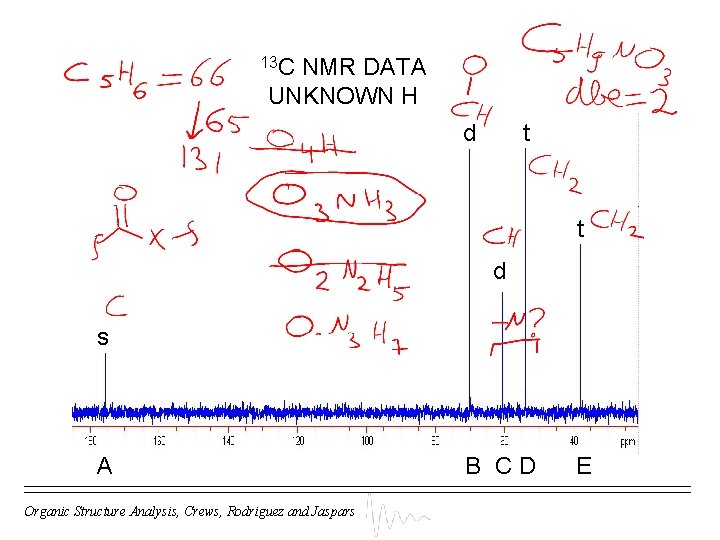
13 C NMR DATA UNKNOWN H d t t d s A Organic Structure Analysis, Crews, Rodriguez and Jaspars B CD E

MOLECULAR FORMULA DETERMINATION UNKNOWN H s d t q (C) + (CH 2) + (CH 3) = C H = Da The MW is 131, so there is a mass difference of Da (= ) Therefore molecular formula = C H N O ( dbe) Organic Structure Analysis, Crews, Rodriguez and Jaspars

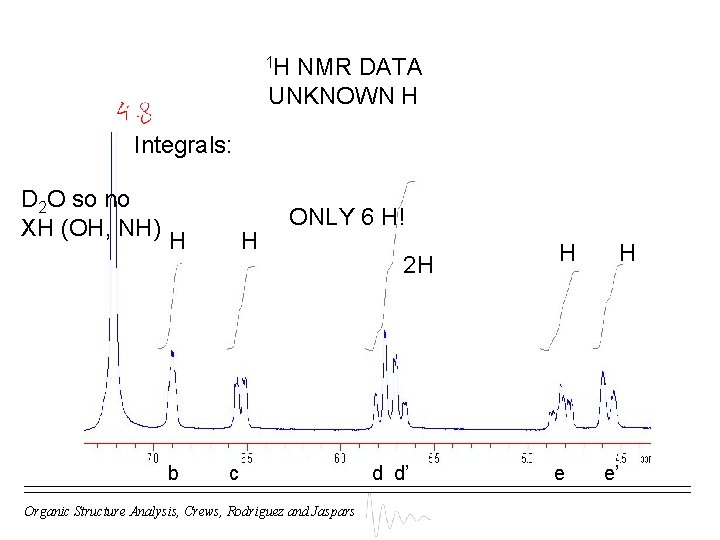
1 H NMR DATA UNKNOWN H Integrals: D 2 O so no XH (OH, NH) H b H ONLY 6 H! c Organic Structure Analysis, Crews, Rodriguez and Jaspars 2 H d d’ H e’

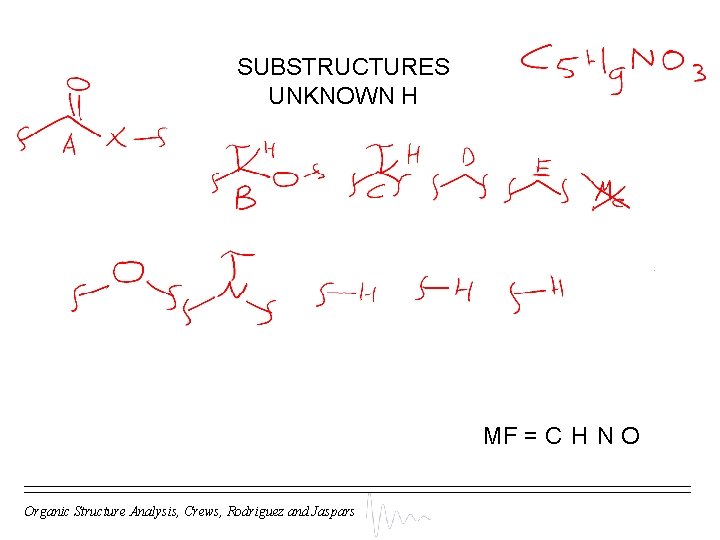
SUBSTRUCTURES UNKNOWN H MF = C H N O Organic Structure Analysis, Crews, Rodriguez and Jaspars

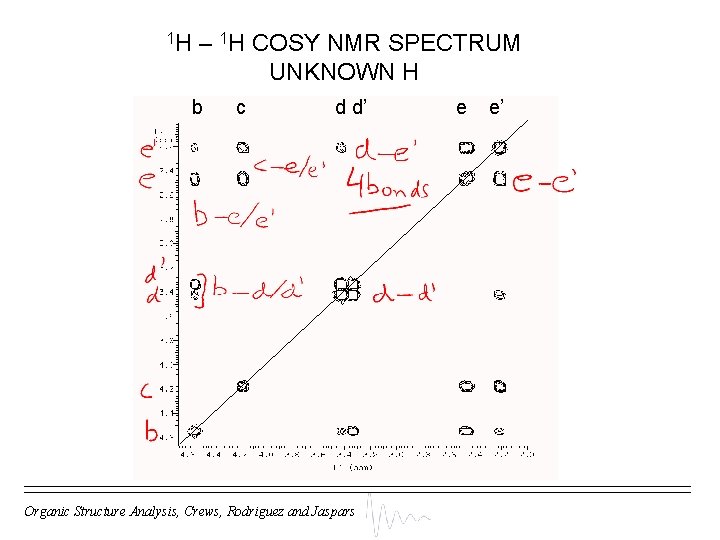
1 H – 1 H COSY NMR SPECTRUM UNKNOWN H b c d d’ Organic Structure Analysis, Crews, Rodriguez and Jaspars e e’

1 H – 1 H COSY NMR DATA UNKNOWN H Combine your substructures using COSY data Organic Structure Analysis, Crews, Rodriguez and Jaspars

SUBTRUCTURES UNKNOWN H Working structures Consider stereochemistry: MF = C H N O Organic Structure Analysis, Crews, Rodriguez and Jaspars

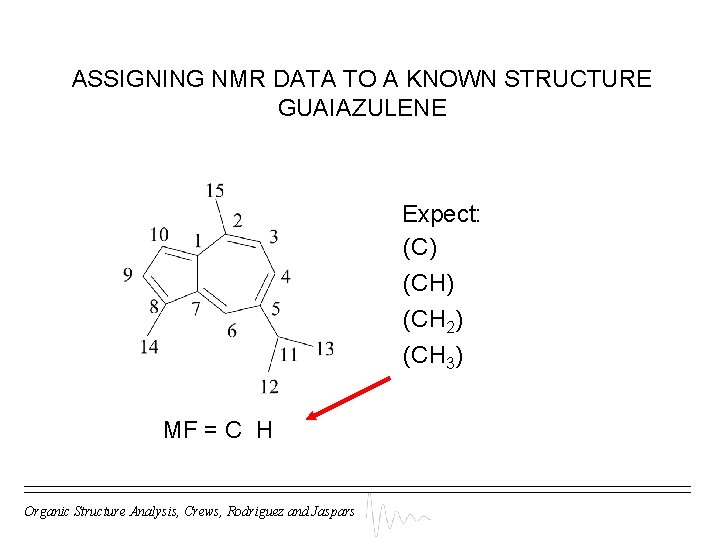
ASSIGNING NMR DATA TO A KNOWN STRUCTURE GUAIAZULENE Expect: (C) (CH 2) (CH 3) MF = C H Organic Structure Analysis, Crews, Rodriguez and Jaspars

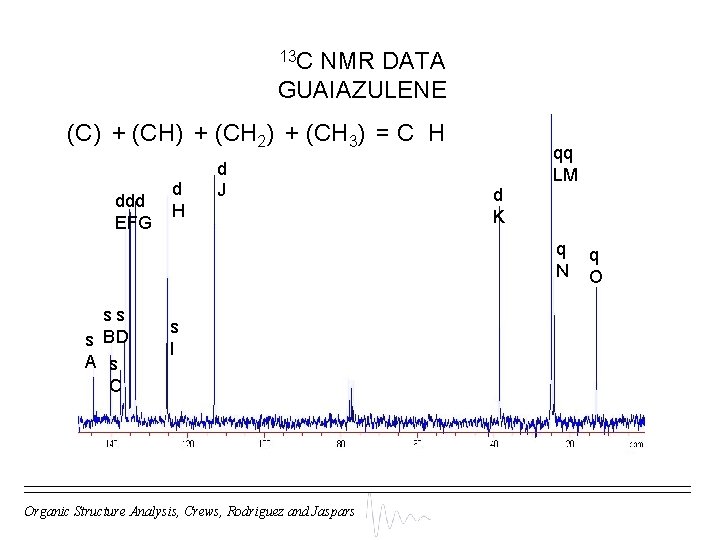
13 C NMR DATA GUAIAZULENE (C) + (CH 2) + (CH 3) = C H ddd EFG d H d J qq LM d K q N ss s BD A s C s I Organic Structure Analysis, Crews, Rodriguez and Jaspars q O

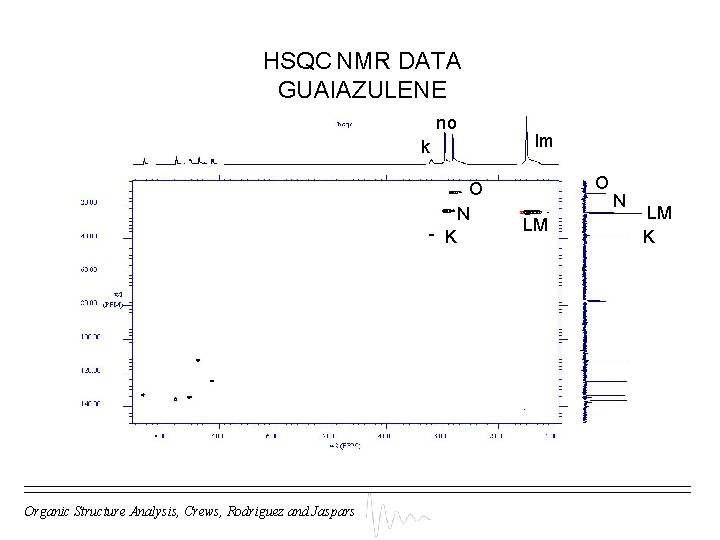
HSQC NMR DATA GUAIAZULENE no lm k O N K Organic Structure Analysis, Crews, Rodriguez and Jaspars O LM N LM K

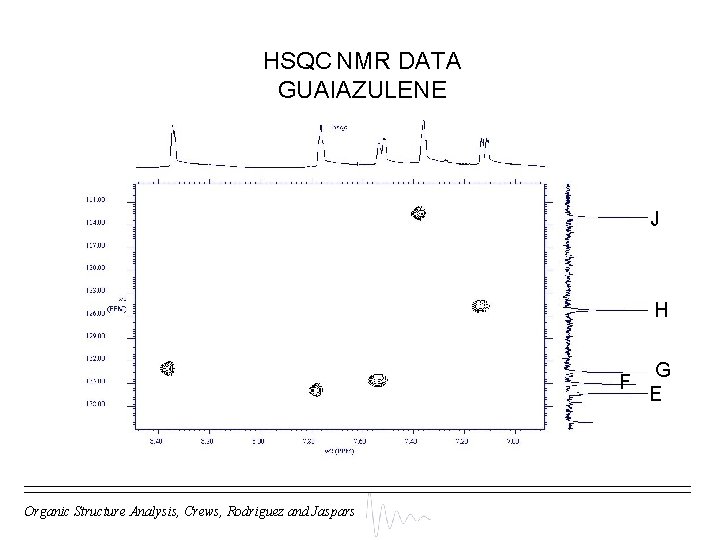
HSQC NMR DATA GUAIAZULENE J H F Organic Structure Analysis, Crews, Rodriguez and Jaspars G E

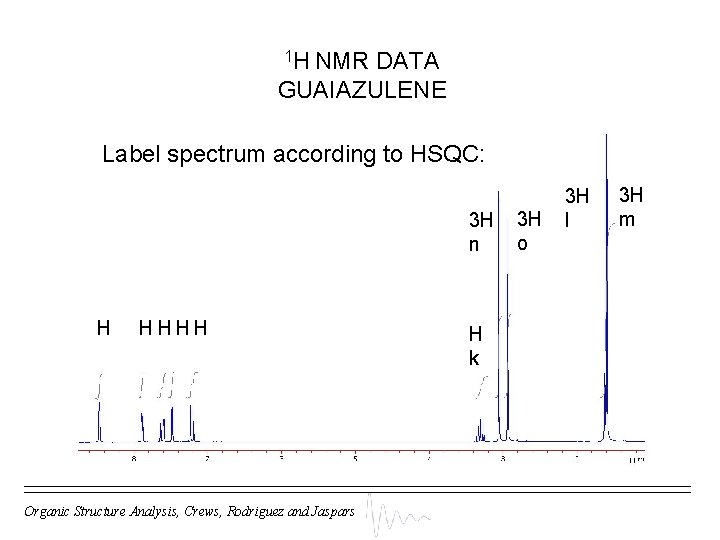
1 H NMR DATA GUAIAZULENE Label spectrum according to HSQC: 3 H n H HHHH Organic Structure Analysis, Crews, Rodriguez and Jaspars H k 3 H o 3 H l 3 H m

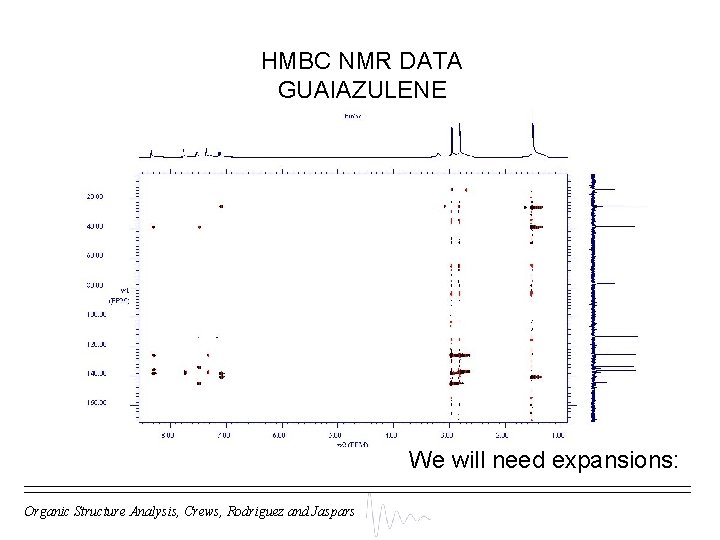
HMBC NMR DATA GUAIAZULENE We will need expansions: Organic Structure Analysis, Crews, Rodriguez and Jaspars

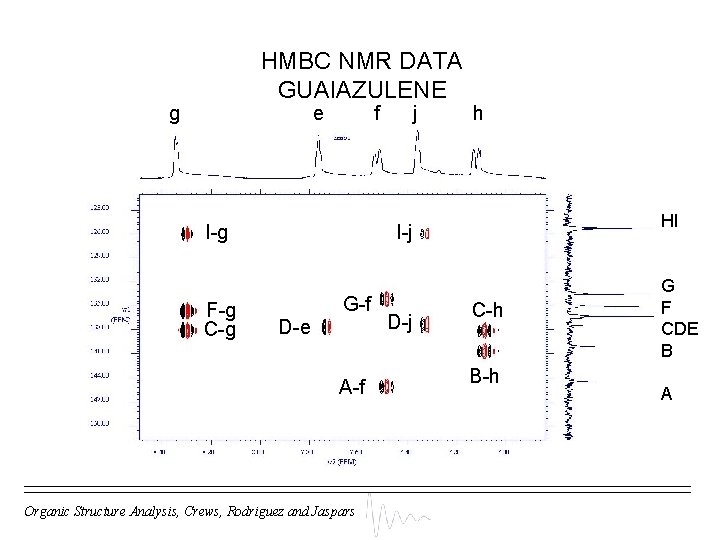
HMBC NMR DATA GUAIAZULENE g e f I-g F-g C-g j h HI I-j G-f D-e A-f Organic Structure Analysis, Crews, Rodriguez and Jaspars D-j C-h B-h G F CDE B A

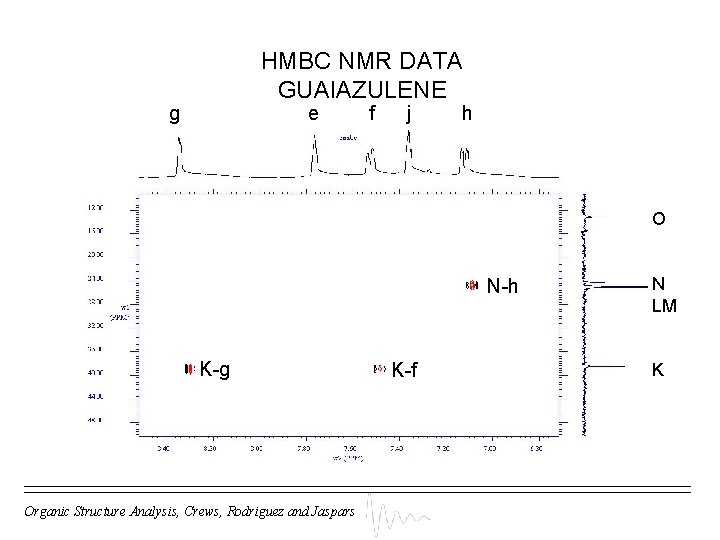
HMBC NMR DATA GUAIAZULENE g e f j h O N-h K-g Organic Structure Analysis, Crews, Rodriguez and Jaspars K-f N LM K

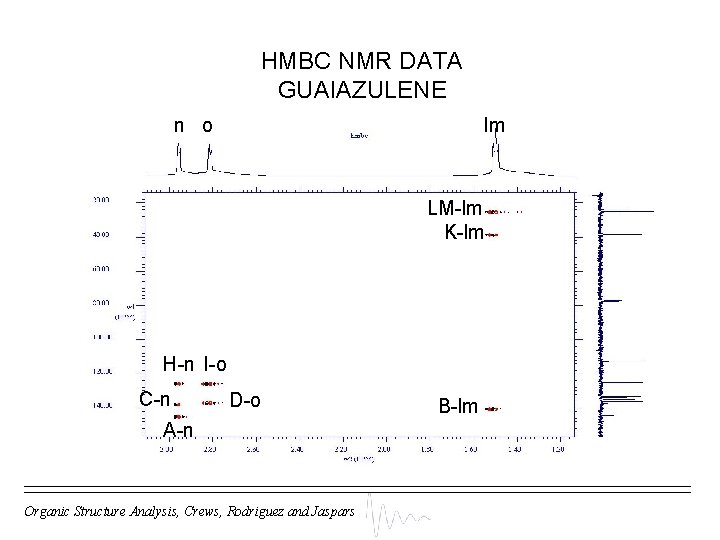
HMBC NMR DATA GUAIAZULENE n o lm LM-lm K-lm H-n I-o C-n D-o A-n Organic Structure Analysis, Crews, Rodriguez and Jaspars B-lm

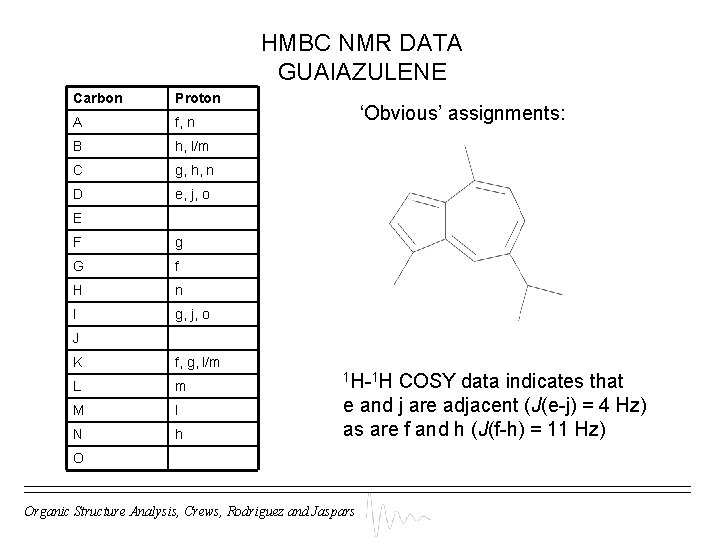
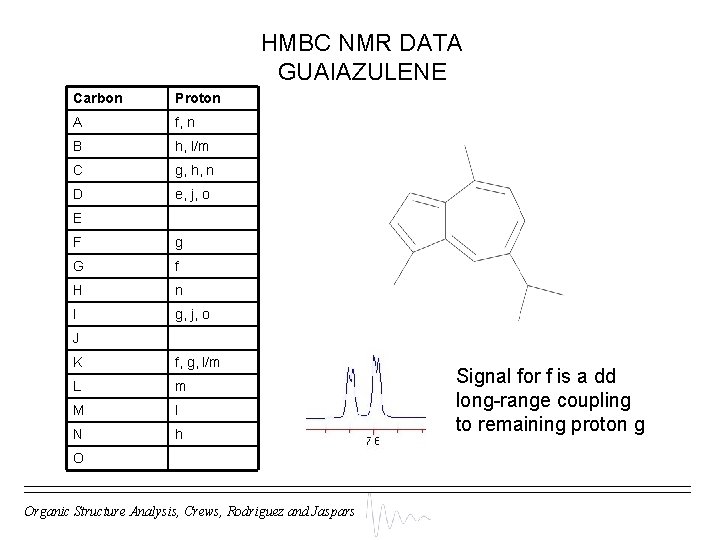
HMBC NMR DATA GUAIAZULENE Carbon Proton A f, n B h, l/m C g, h, n D e, j, o ‘Obvious’ assignments: E F g G f H n I g, j, o J K f, g, l/m L m M l N h 1 H-1 H COSY data indicates that e and j are adjacent (J(e-j) = 4 Hz) as are f and h (J(f-h) = 11 Hz) O Organic Structure Analysis, Crews, Rodriguez and Jaspars

HMBC NMR DATA GUAIAZULENE Carbon Proton A f, n B h, l/m C g, h, n D e, j, o E F g G f H n I g, j, o J K f, g, l/m L m M l N h O Organic Structure Analysis, Crews, Rodriguez and Jaspars Signal for f is a dd long-range coupling to remaining proton g

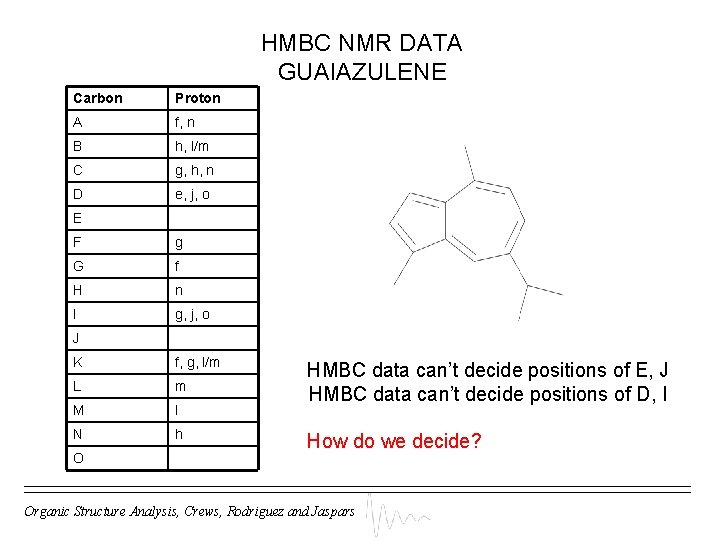
HMBC NMR DATA GUAIAZULENE Carbon Proton A f, n B h, l/m C g, h, n D e, j, o E F g G f H n I g, j, o J K f, g, l/m L m M l N h O HMBC data can’t decide positions of E, J HMBC data can’t decide positions of D, I How do we decide? Organic Structure Analysis, Crews, Rodriguez and Jaspars

FINALISING THE ASSIGNMENTS GUAIAZULENE Organic Structure Analysis, Crews, Rodriguez and Jaspars

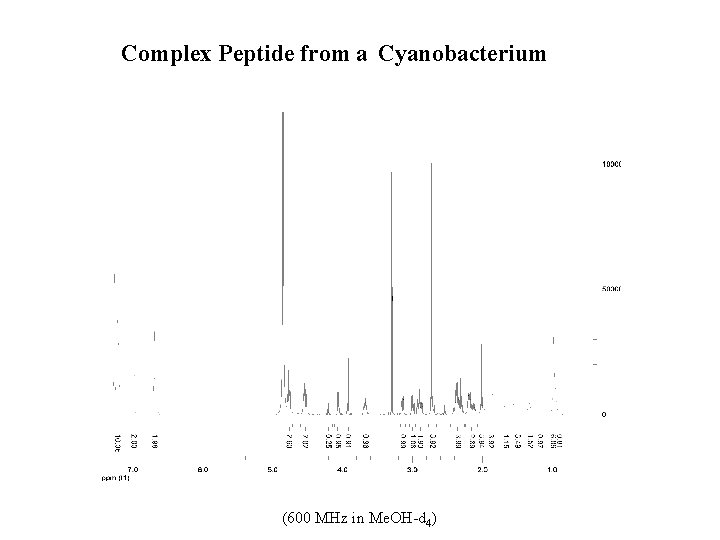
Complex Peptide from a Cyanobacterium (600 MHz in Me. OH-d 4)

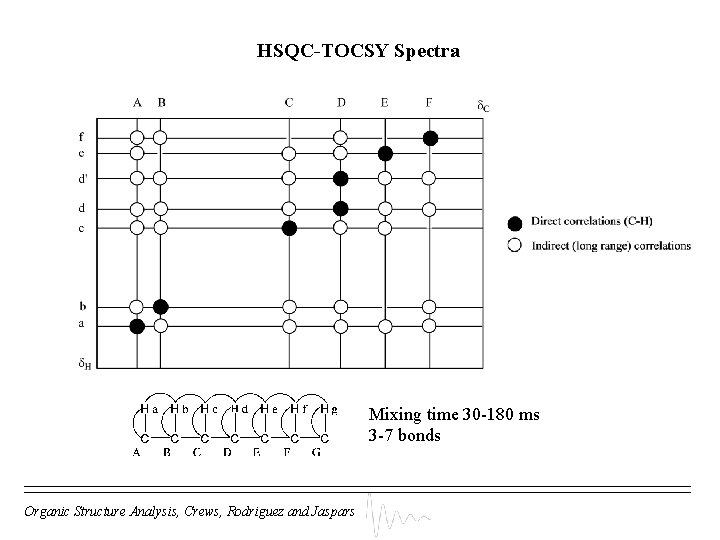
HSQC-TOCSY Spectra Mixing time 30 -180 ms 3 -7 bonds Organic Structure Analysis, Crews, Rodriguez and Jaspars

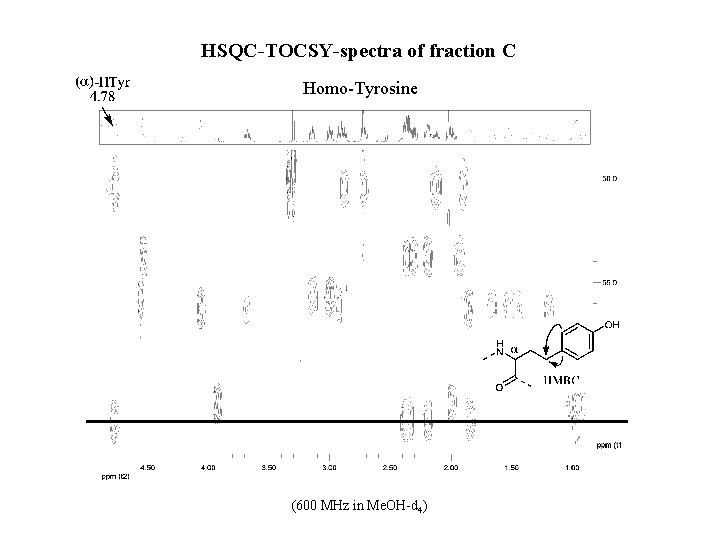
HSQC-TOCSY-spectra of fraction C Homo-Tyrosine (600 MHz in Me. OH-d 4)

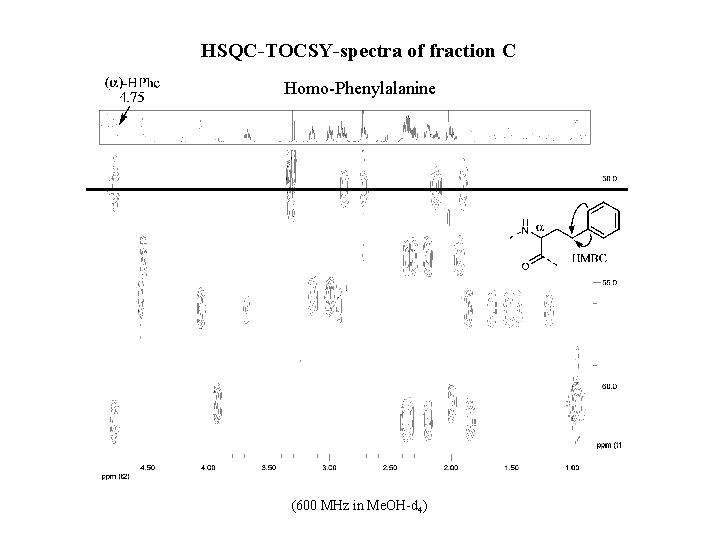
HSQC-TOCSY-spectra of fraction C Homo-Phenylalanine (600 MHz in Me. OH-d 4)

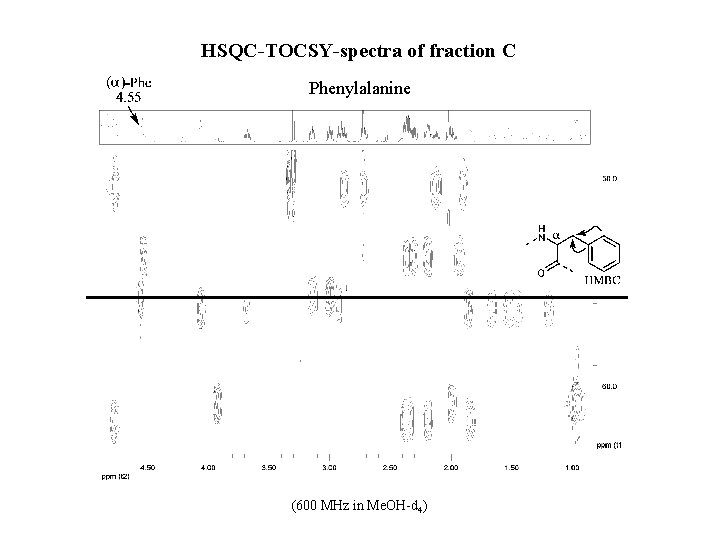
HSQC-TOCSY-spectra of fraction C Phenylalanine (600 MHz in Me. OH-d 4)

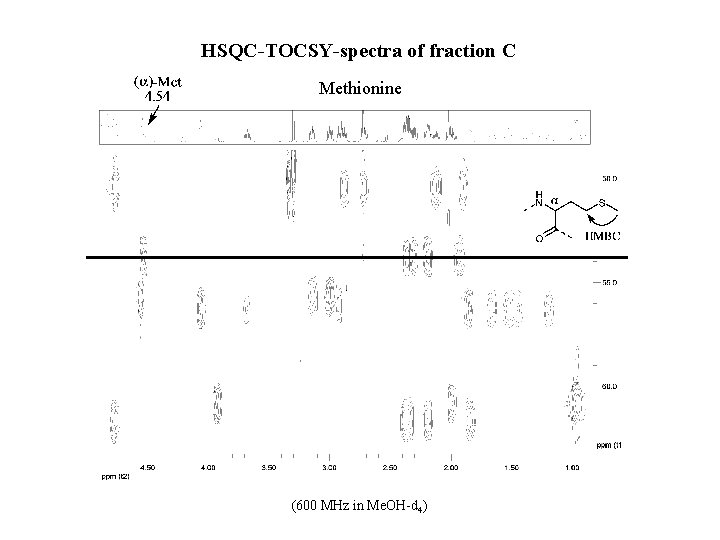
HSQC-TOCSY-spectra of fraction C Methionine (600 MHz in Me. OH-d 4)

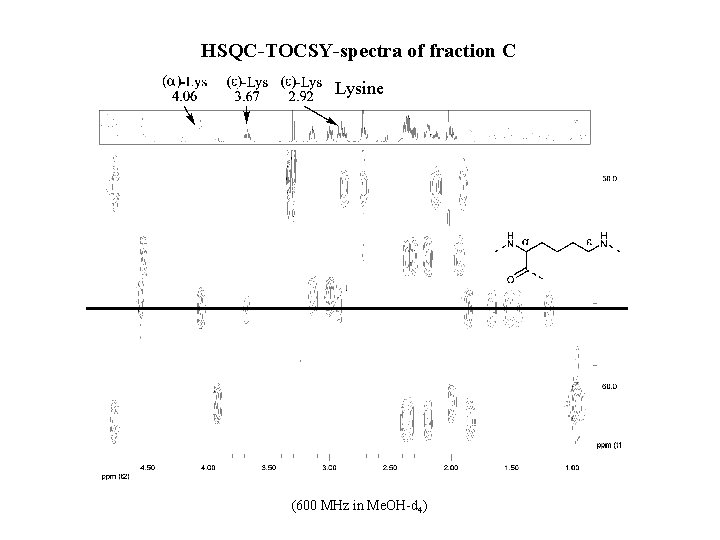
HSQC-TOCSY-spectra of fraction C Lysine (600 MHz in Me. OH-d 4)

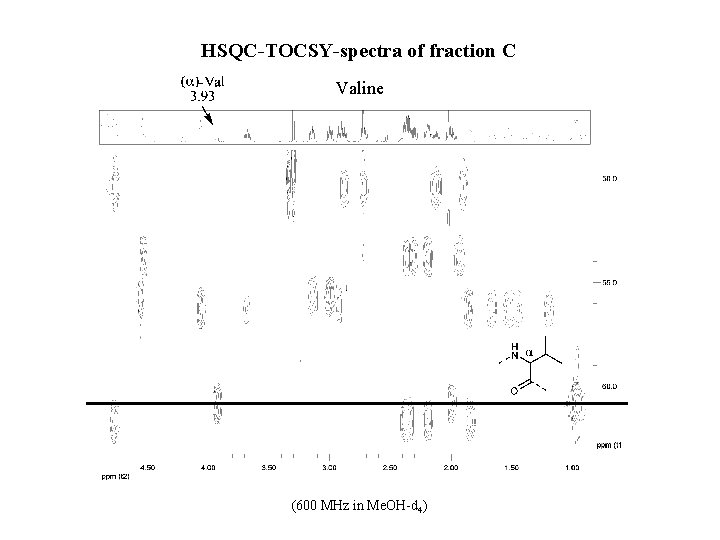
HSQC-TOCSY-spectra of fraction C Valine (600 MHz in Me. OH-d 4)

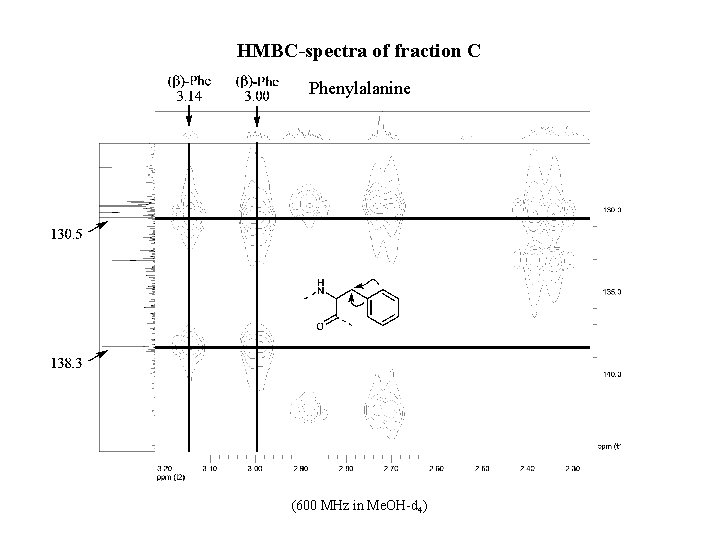
HMBC-spectra of fraction C Phenylalanine (600 MHz in Me. OH-d 4)

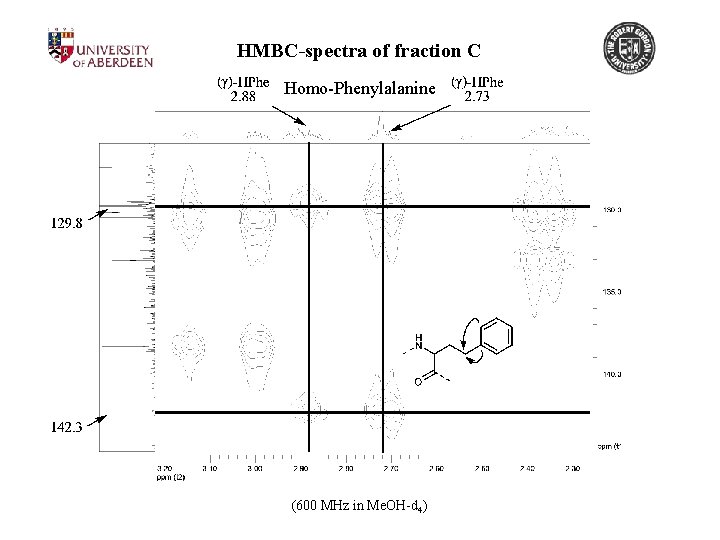
HMBC-spectra of fraction C Homo-Phenylalanine (600 MHz in Me. OH-d 4)

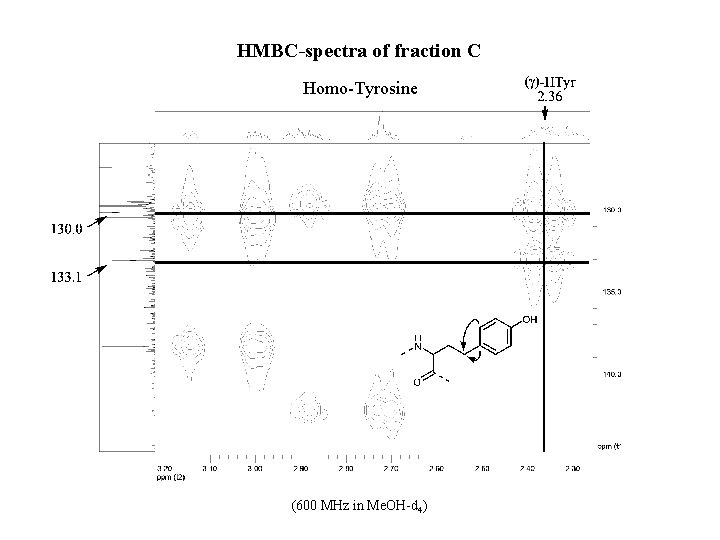
HMBC-spectra of fraction C Homo-Tyrosine (600 MHz in Me. OH-d 4)

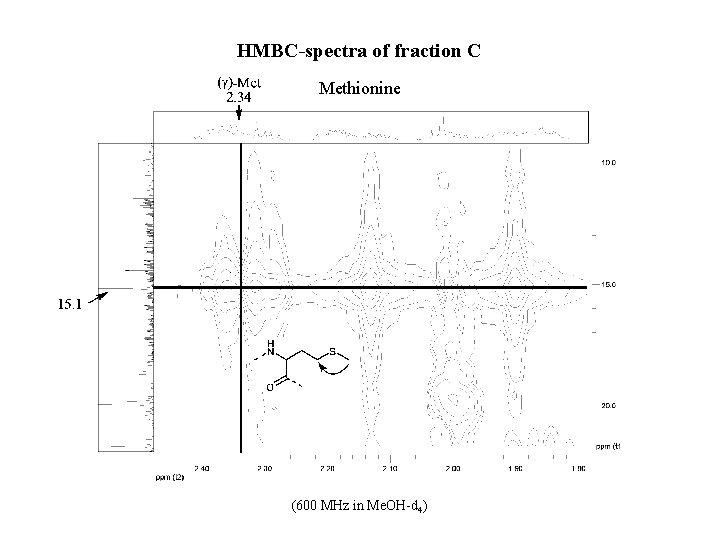
HMBC-spectra of fraction C Methionine (600 MHz in Me. OH-d 4)

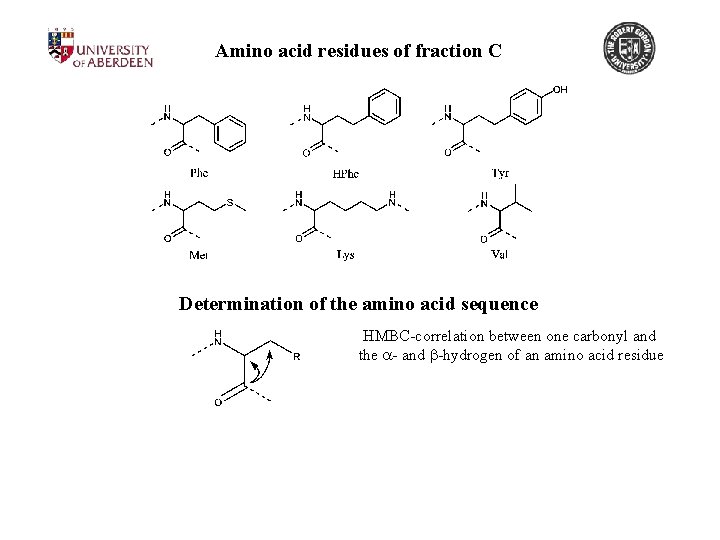
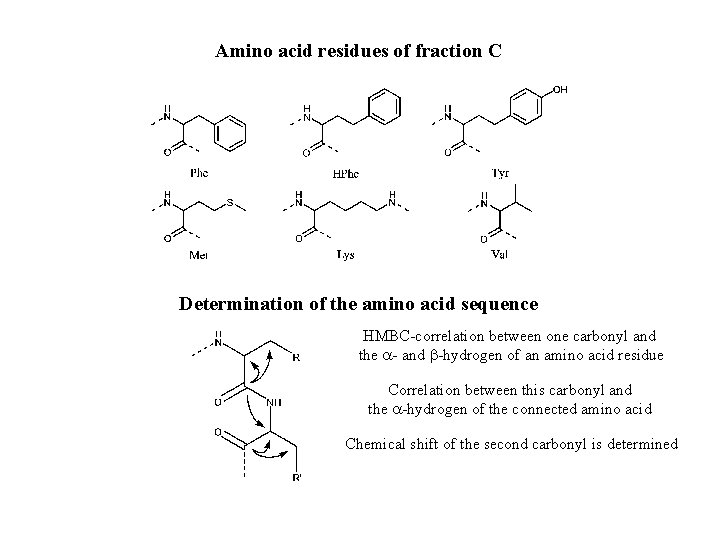
Amino acid residues of fraction C Determination of the amino acid sequence HMBC-correlation between one carbonyl and the a- and b-hydrogen of an amino acid residue

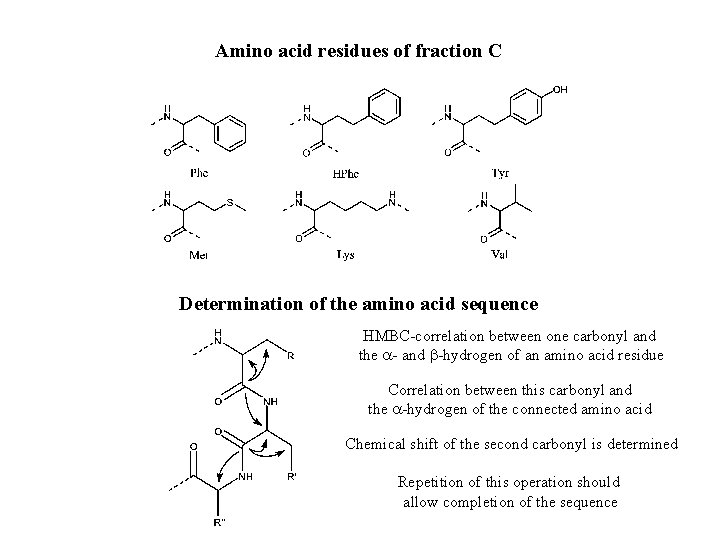
Amino acid residues of fraction C Determination of the amino acid sequence HMBC-correlation between one carbonyl and the a- and b-hydrogen of an amino acid residue Correlation between this carbonyl and the a-hydrogen of the connected amino acid Chemical shift of the second carbonyl is determined

Amino acid residues of fraction C Determination of the amino acid sequence HMBC-correlation between one carbonyl and the a- and b-hydrogen of an amino acid residue Correlation between this carbonyl and the a-hydrogen of the connected amino acid Chemical shift of the second carbonyl is determined Repetition of this operation should allow completion of the sequence

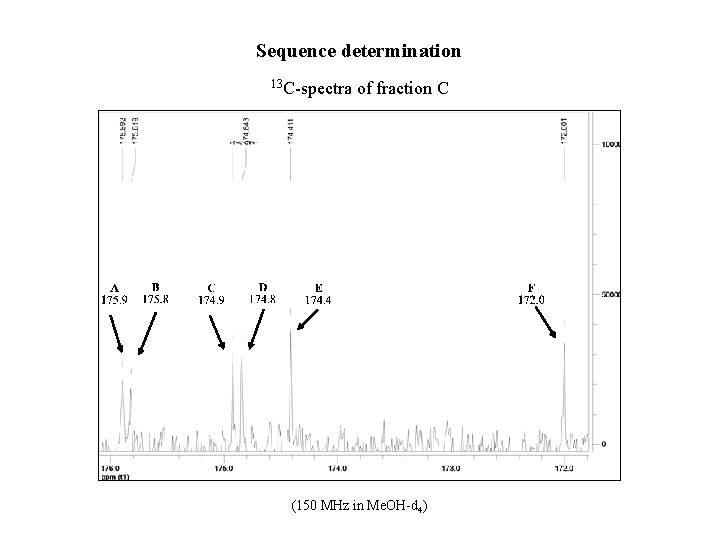
Sequence determination 13 C-spectra of fraction C (150 MHz in Me. OH-d 4)

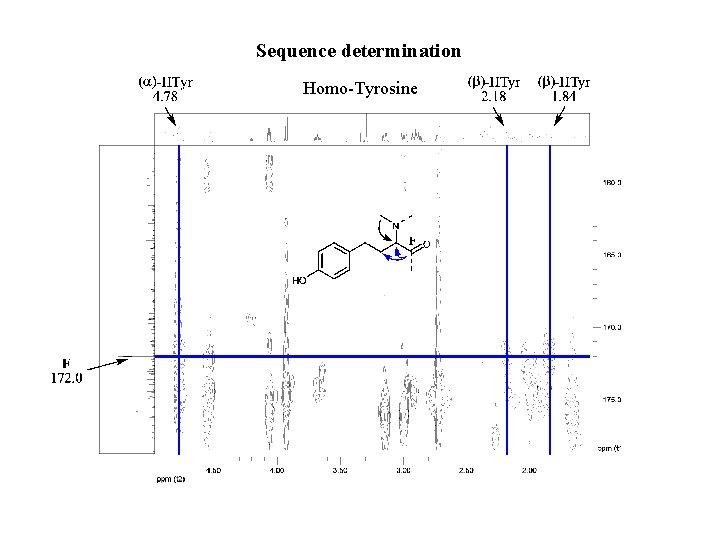
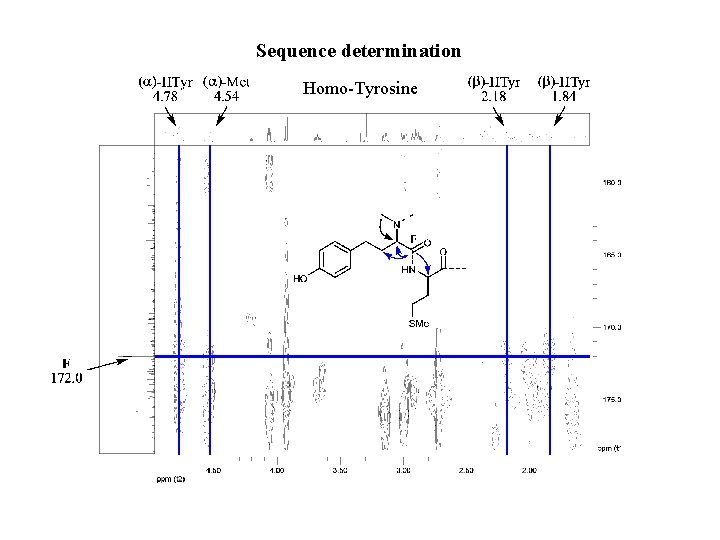
Sequence determination Homo-Tyrosine

Sequence determination Homo-Tyrosine

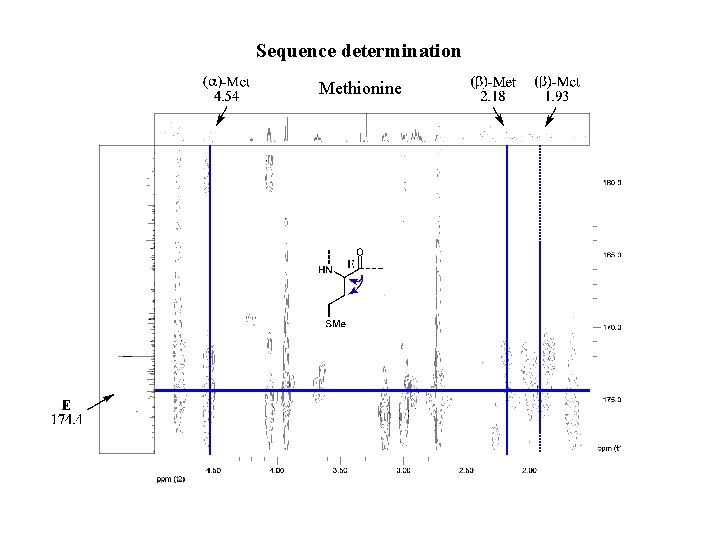
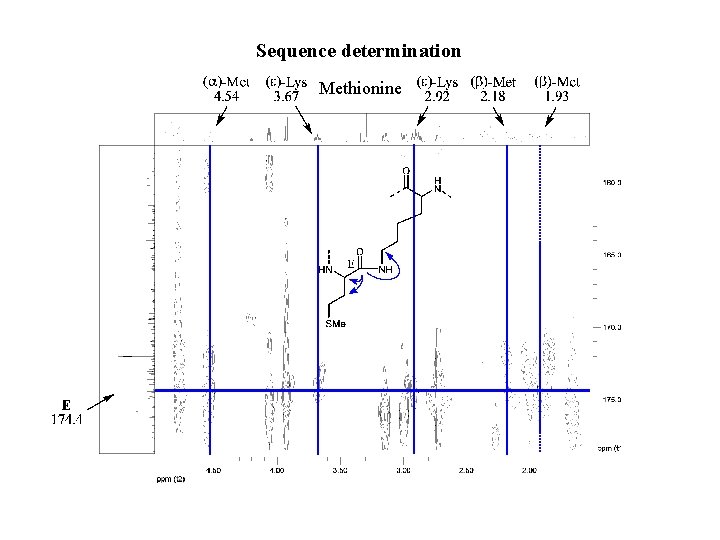
Sequence determination Methionine

Sequence determination Methionine

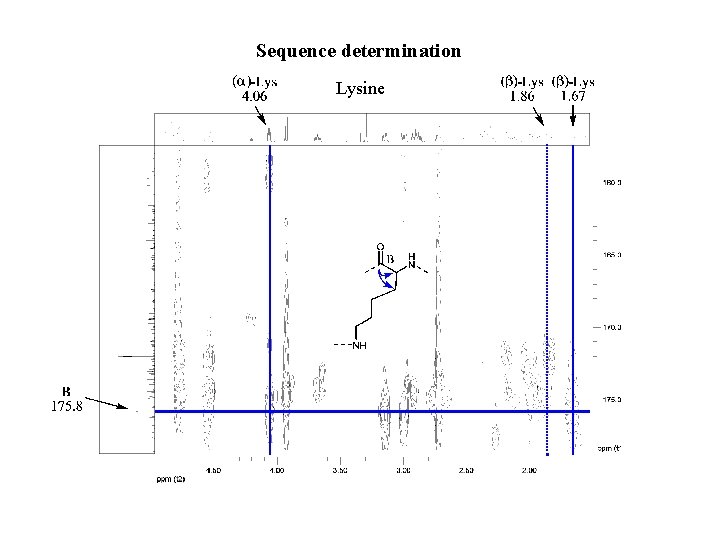
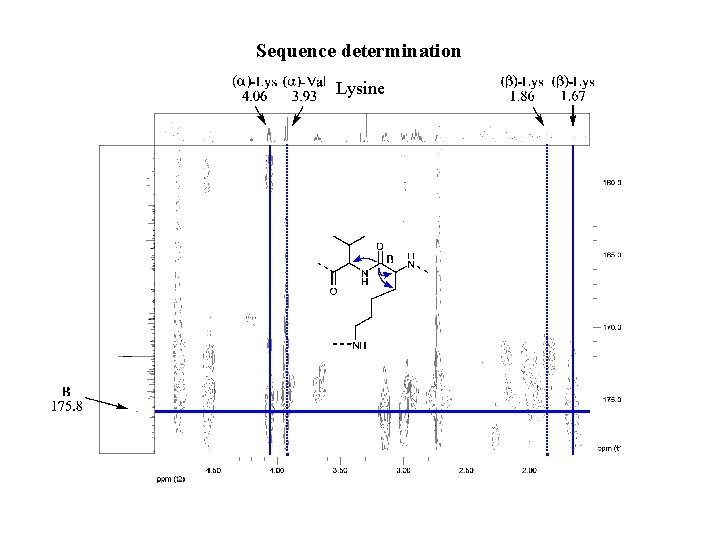
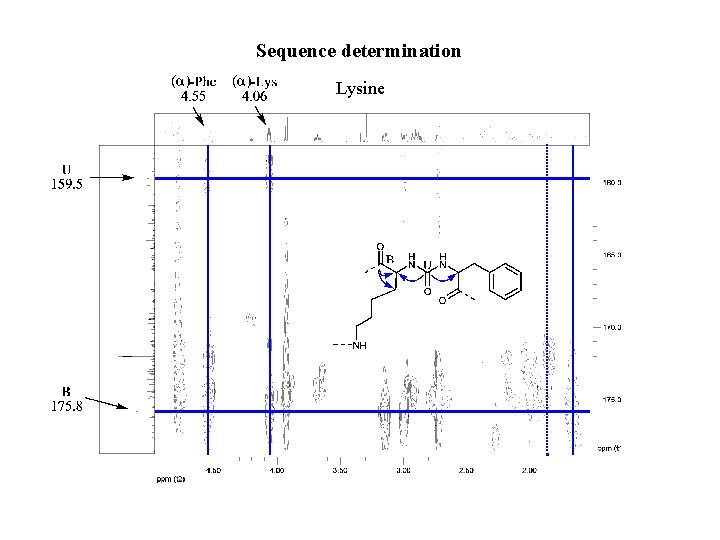
Sequence determination Lysine

Sequence determination Lysine

Sequence determination Lysine

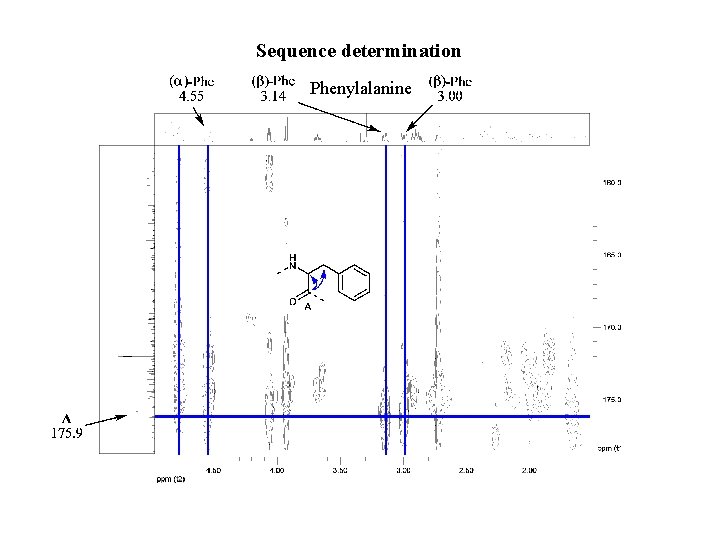
Sequence determination Phenylalanine

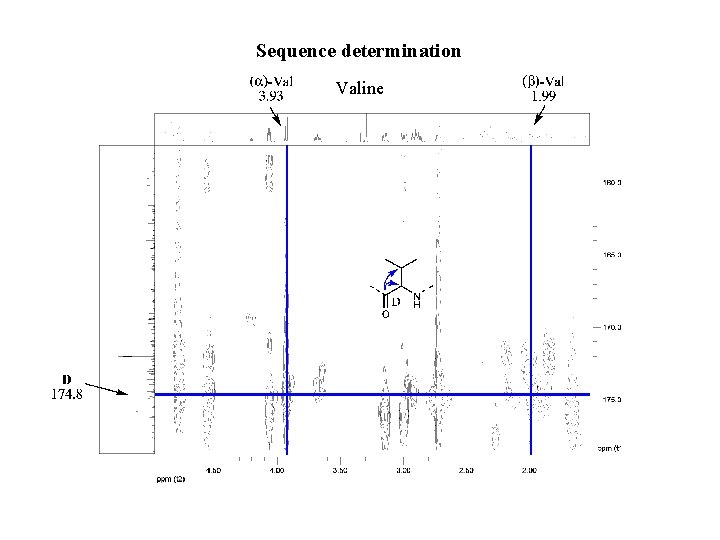
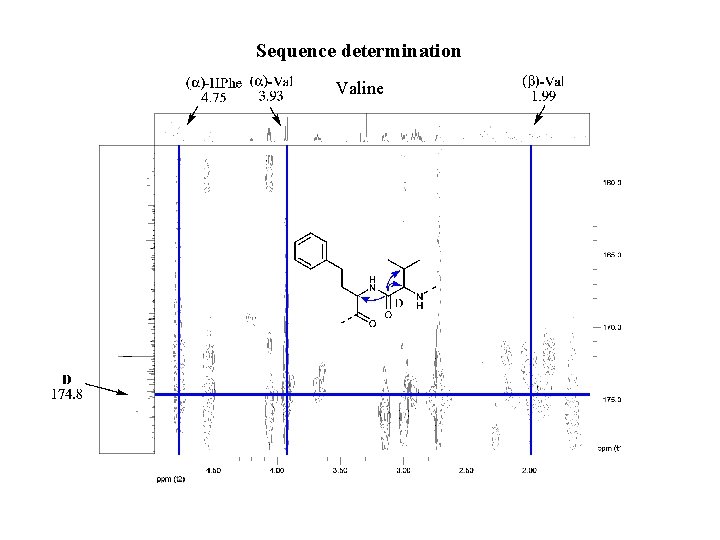
Sequence determination Valine

Sequence determination Valine

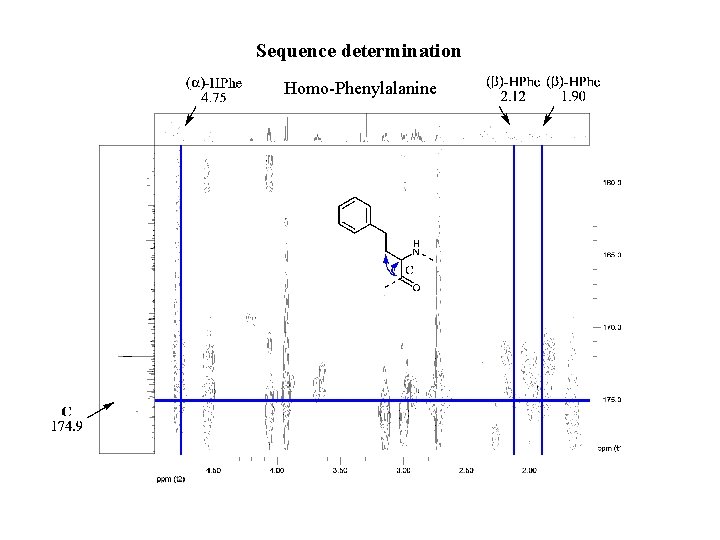
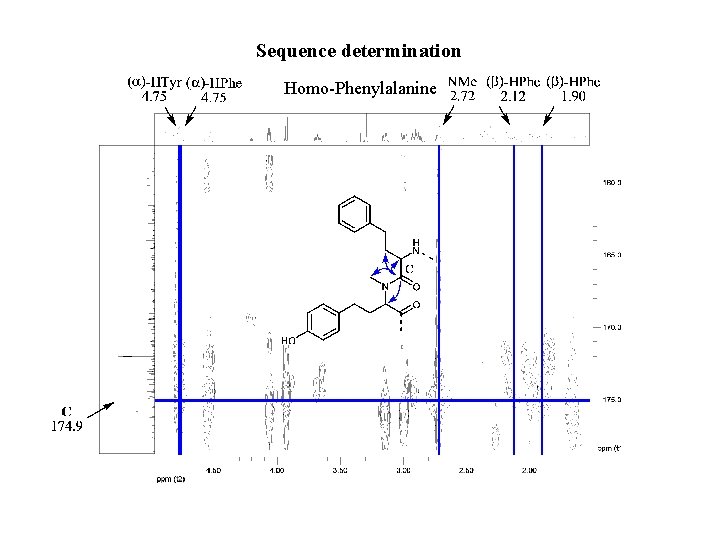
Sequence determination Homo-Phenylalanine

Sequence determination Homo-Phenylalanine

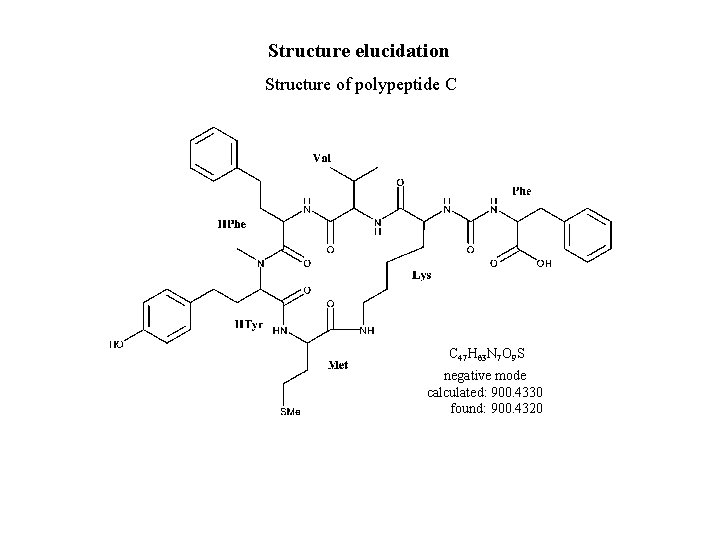
Structure elucidation Structure of polypeptide C C 47 H 63 N 7 O 9 S negative mode calculated: 900. 4330 found: 900. 4320

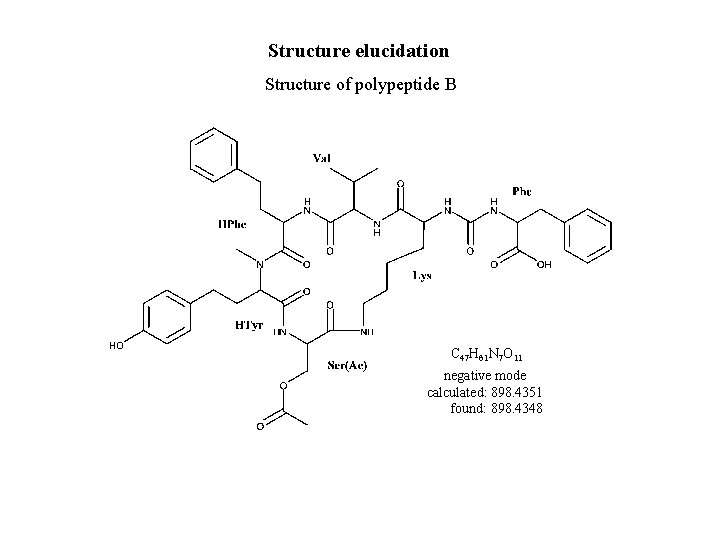
Structure elucidation Structure of polypeptide B C 47 H 61 N 7 O 11 negative mode calculated: 898. 4351 found: 898. 4348

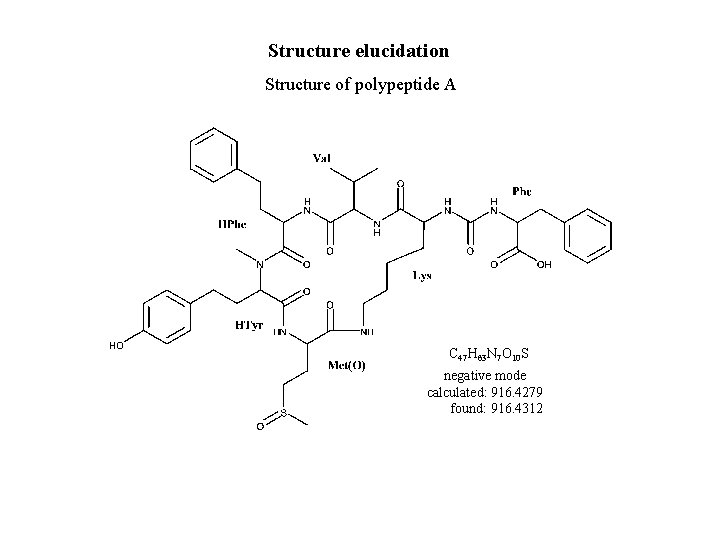
Structure elucidation Structure of polypeptide A C 47 H 63 N 7 O 10 S negative mode calculated: 916. 4279 found: 916. 4312

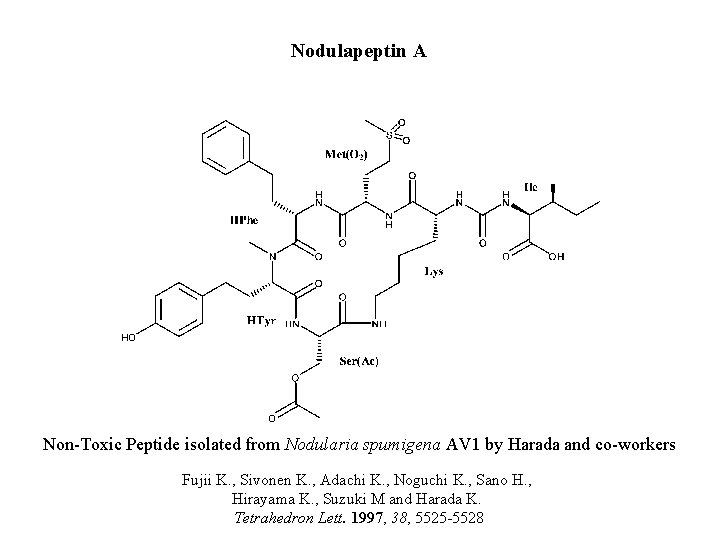
Nodulapeptin A Non-Toxic Peptide isolated from Nodularia spumigena AV 1 by Harada and co-workers Fujii K. , Sivonen K. , Adachi K. , Noguchi K. , Sano H. , Hirayama K. , Suzuki M and Harada K. Tetrahedron Lett. 1997, 38, 5525 -5528

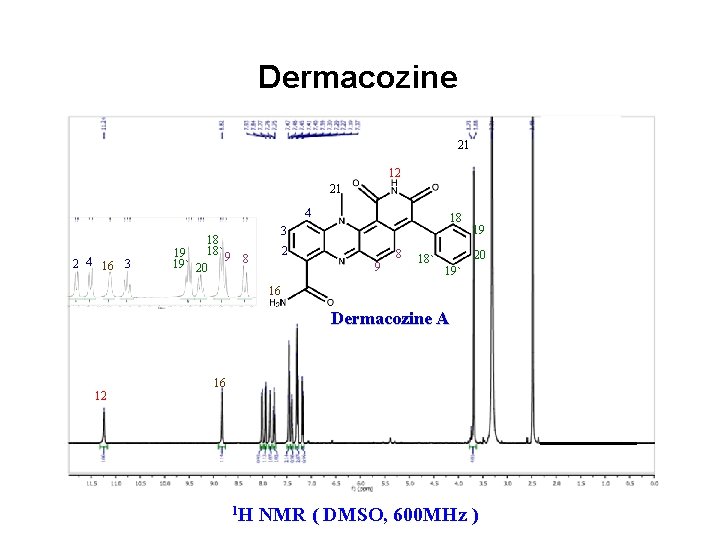
Dermacozine 21 12 21 4 2 4 16 3 18 18` 9 19 8 19` 20 2 9 8 18` 19 20 19` 16 Dermacozine A 12 16 1 H NMR ( DMSO, 600 MHz )

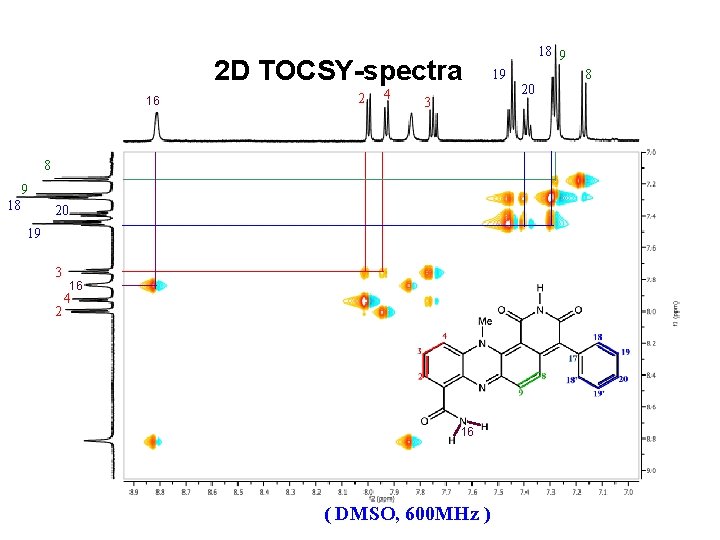
2 D TOCSY-spectra 16 2 4 3 8 9 18 20 19 3 2 16 4 16 ( DMSO, 600 MHz ) 18 9 19 20 8

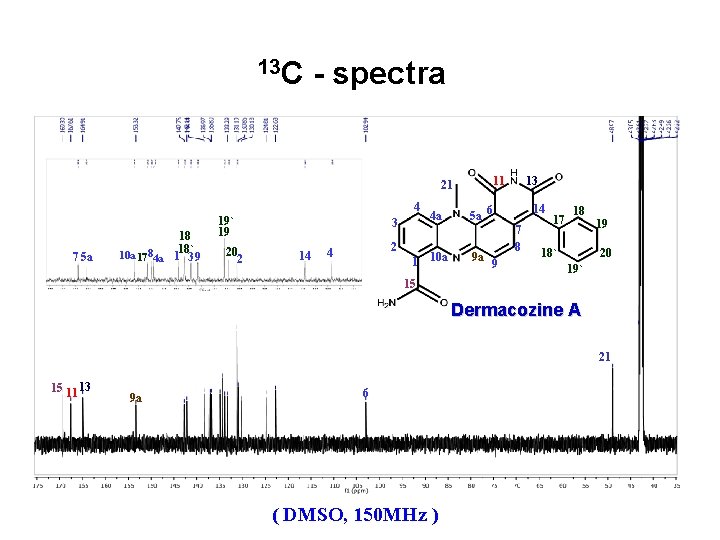
13 C - spectra 4 7 5 a 18 18` 10 a 17 84 a 1 3 9 19` 19 20 2 3 14 2 4 1 4 a 10 a 13 11 21 14 5 a 6 9 a 7 8 9 17 18 19 18` 20 19` 15 Dermacozine A 21 15 11 13 9 a 6 ( DMSO, 150 MHz )

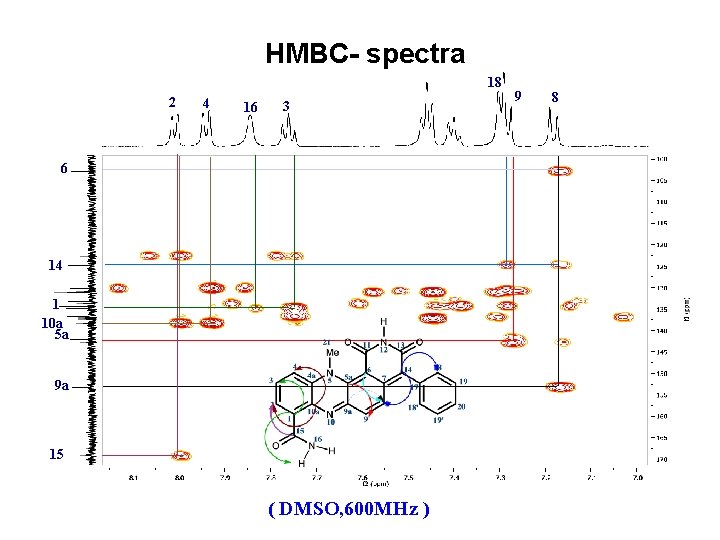
HMBC- spectra 18 2 4 16 3 6 14 1 10 a 5 a 9 a 15 ( DMSO, 600 MHz ) 9 8

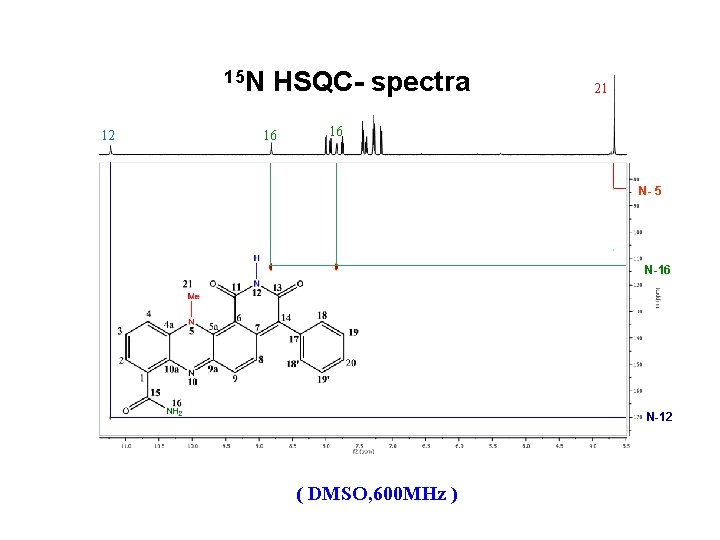
15 N 12 HSQC- spectra 16 21 16 N- 5 N-16 N-12 ( DMSO, 600 MHz )