Soft Computing Computational Intelligence Biologically inspired computing models

Soft Computing & Computational Intelligence • • Biologically inspired computing models Compatible with human expertise/reasoning Intensive numerical computations Data and goal driven Model-free learning Fault tolerant Real world/novel applications
Soft Computing & Computational Intelligence • • • Artificial Neural Networks (ANN) Fuzzy Logic Genetic Algorithms (GAs) Fractals/Chaos Artificial life Wavelets • Data mining FL ANNs GAs

Biological Neuron hair cell dendrites (sensory transducer) signal flow synapse axon hillock cell body axon synapse
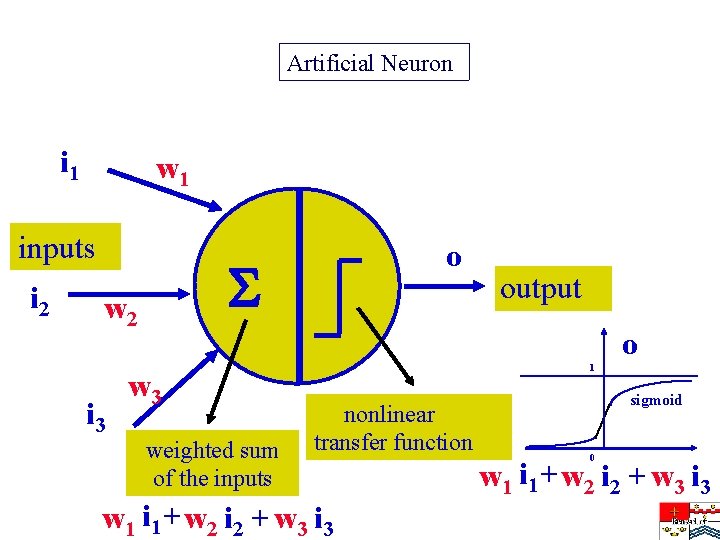
Artificial Neuron i 1 w 1 inputs i 2 w 2 i 3 o output o w 3 weighted sum of the inputs 1 nonlinear transfer function w 1 i 1 + w 2 i 2 + w 3 i 3 sigmoid 0 w 1 i 1 + w 2 i 2 + w 3 i 3
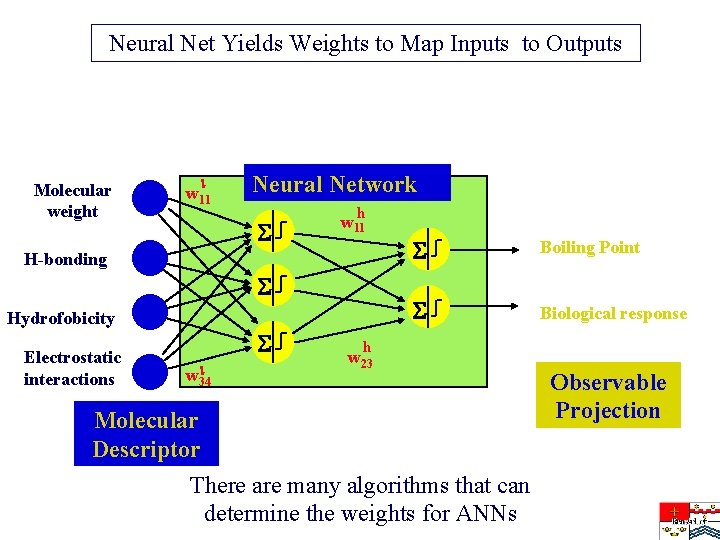
Neural Net Yields Weights to Map Inputs to Outputs Molecular weight w 11 Neural Network h w 11 H-bonding Hydrofobicity Electrostatic interactions w 34 Boiling Point Biological response h w 23 Molecular Descriptor There are many algorithms that can determine the weights for ANNs Observable Projection

Neural Networks in a Nutshell • A problem can be formulated and represented as a mapping problem from • Such a map can be realized by an ANN, which is a framework of basic building blocks of Mc. Culloch-Pitts neurons • The neural net can be trained to conform with the map based on samples of the map and will reasonably generalize to new cases it has not encountered before
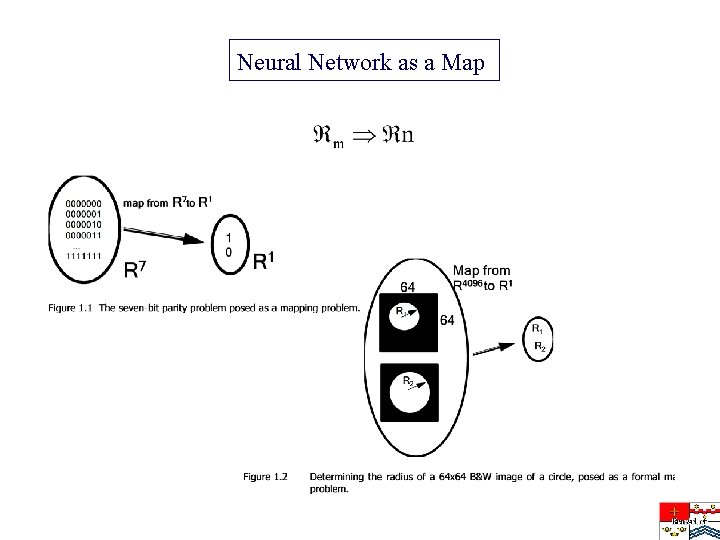
Neural Network as a Map

Poisonous/Edible Mushroom Classification Problem 1. cap-shape: bell=b, conical=c, convex=x, flat=f, knobbed=k, sunken=s 2. cap-surface: fibrous=f, grooves=g, scaly=y, smooth=s 3. cap-color: brown=n, buff=b, cinnamon=c, gray=g, green=r, pink=p, purple=u, red=e, white=w, yellow=y 4. bruises? : bruises=t, no=f 5. odor: almond=a, anise=l, creosote=c, fishy=y, foul=f, musty=m, none=n, pungent=p, spicy=s 6. gill-attachment: attached=a, descending=d, free=f, notched=n 7. gill-spacing: close=c, crowded=w, distant=d 8. gill-size: broad=b, narrow=n 9. gill-color: black=k, brown=n, buff=b, chocolate=h, gray=g, green=r, orange=o, pink=p, purple=u, red=e, white=w, yellow=y 10. stalk-shape: enlarging=e, tapering=t 11. stalk-root: bulbous=b, club=c, cup=u, equal=e, rhizomorphs=z, rooted=r, missing=? 12. stalk-surface-above-ring: ibrous=f, scaly=y, silky=k, smooth=s 13. stalk-surface-below-ring: ibrous=f, scaly=y, silky=k, smooth=s 14. stalk-color-above-ring: brown=n, buff=b, cinnamon=c, gray=g, orange=o, pink=p, red=e, white=w, yellow=y 15. stalk-color-below-ring: brown=n, buff=b, cinnamon=c, gray=g, orange=o, pink=p, red=e, white=w, yellow=y 16. veil-type: partial=p, universal=u 17. veil-color: brown=n, orange=o, white=w, yellow=y 18. ring-number: none=n, one=o, two=t 19. ring-type: cobwebby=c, evanescent=e, flaring=f, large=l, none=n, pendant=p, sheathing=s, zone=z 20. spore-print-color: black=k, brown=n, buff=b, chocolate=h, green=r, orange=o, purple=u, white=w, yellow=y 21. population: abundant=a, clustered=c, numerous=n, scattered=s, several=v, solitary=y 22. habitat: grasses=g, leaves=l, meadows=m, paths=p, urban=u, waste=w, woods=d Relevant Information: This data set includes descriptions of hypothetical samples corresponding to 23 species of gilled mushrooms in the Agaricus and Lepiota Family (pp. 500 -525). Each species is identified as definitely edible, definitely poisonous, or of unknown edibility and not recommended. This latter class was combined with the poisonous one. The Guide clearly states that there is no simple rule for determining the edibility of a mushroom; no rule like ``leaflets three, let it be'' for Poisonous Oak and Ivy. Sources: (a) Mushroom records drawn from The Audubon Society Field Guide to North American Mushrooms (1981). G. H. Lincoff (Pres. ), New York: Alfred A. Knopf (b) Donor: Jeff Schlimmer (Jeffrey. Schlimmer@a. gp. cs. cmu. edu) (c) Date: 27 April 1987 Number of Instances: 8124; Number of Attributes: 22 (all nominally valued) Mushroom: original data were alphanumeric. replace alphanumeric attributes in order mentioned by 1, 2, 3 etc
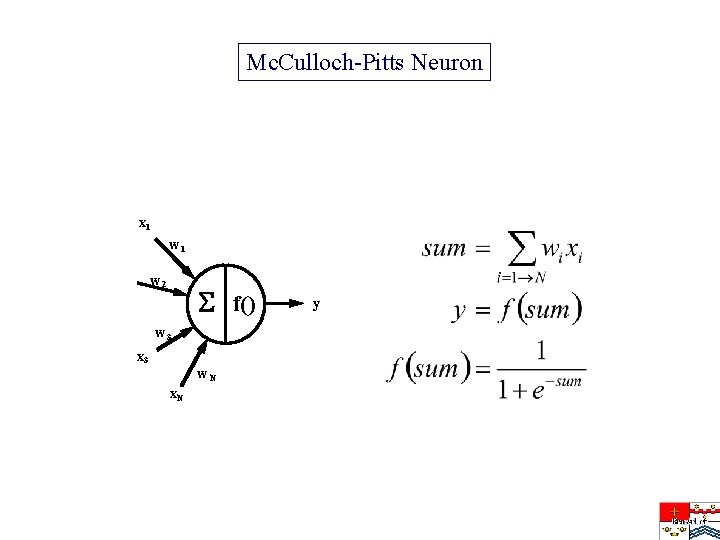
Mc. Culloch-Pitts Neuron x 1 w 2 f() w 3 x 3 w. N x. N y
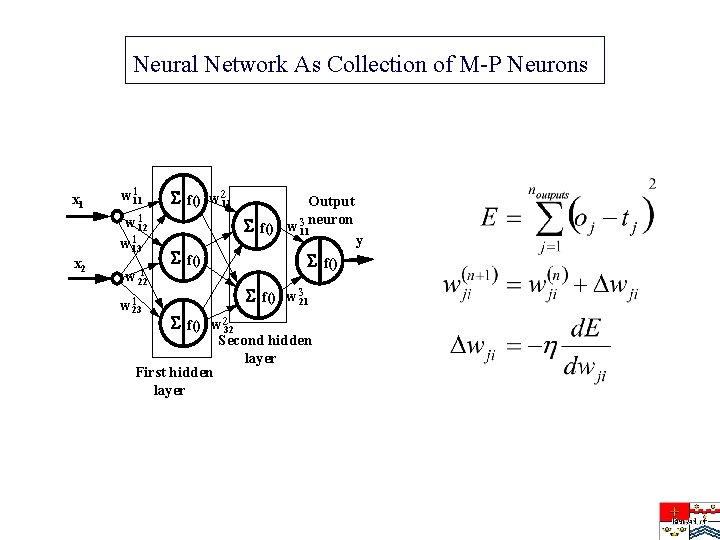
Neural Network As Collection of M-P Neurons x 1 x 2 w 111 w 112 w 113 1 w 22 w 123 f() w 211 f() Output w 3 neuron 11 f() w 321 f() w 232 First hidden layer Second hidden layer y
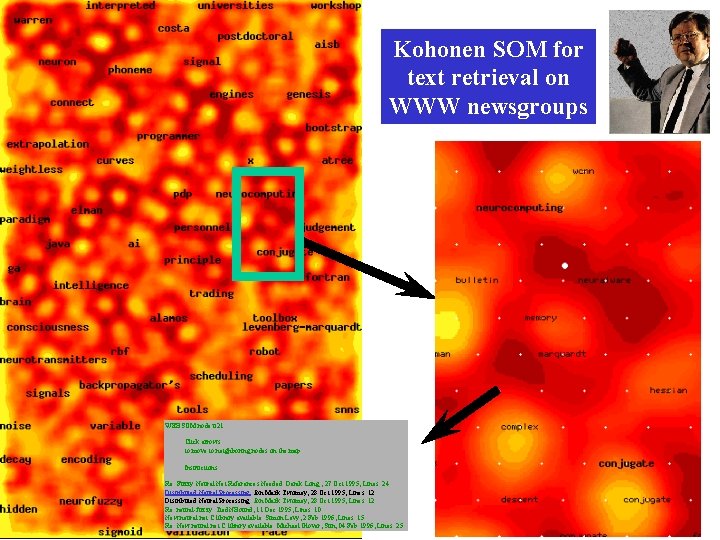
Kohonen SOM for text retrieval on WWW newsgroups WEBSOM node u 21 Click arrows to move to neighboring nodes on the map. Instructions Re: Fuzzy Neural Net References Needed Derek Long , 27 Oct 1995, Lines: 24. Distributed Neural Processing Jon Mark Twomey, 28 Oct 1995, Lines: 12. Re: neural-fuzzy Tied. NBound, 11 Dec 1995, Lines: 10. New neural net C library available Simon Levy, 2 Feb 1996, Lines: 15. Re: New neural net C library available Michael Glover, Sun, 04 Feb 1996, Lines: 25.
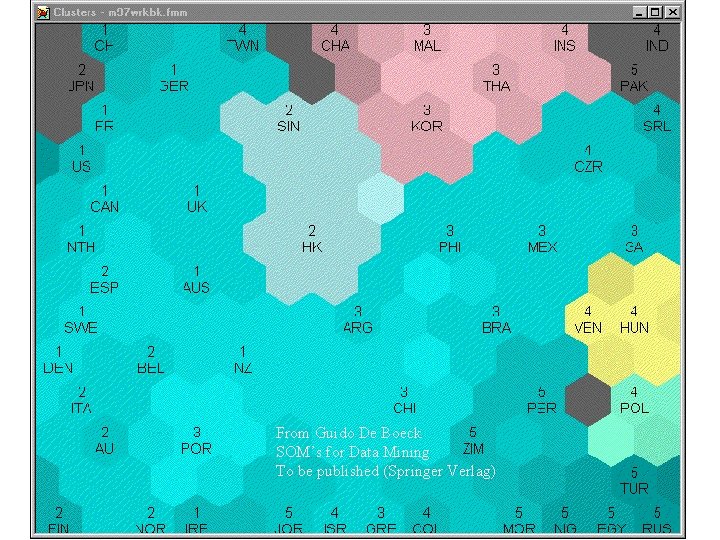
From Guido De Boeck SOM’s for Data Mining To be published (Springer Verlag)
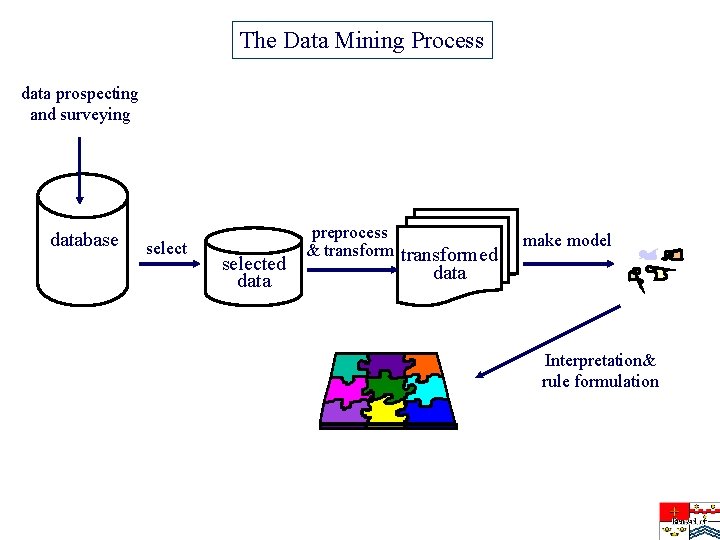
The Data Mining Process data prospecting and surveying database selected data preprocess & transformed make model data Interpretation& rule formulation
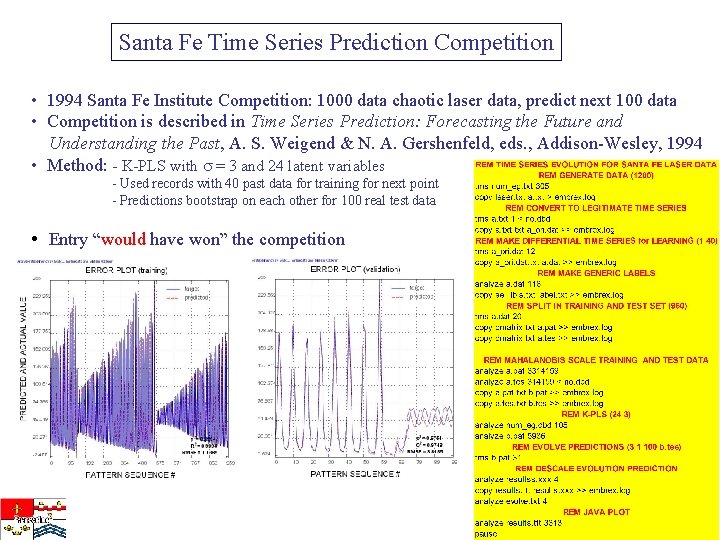
Santa Fe Time Series Prediction Competition • 1994 Santa Fe Institute Competition: 1000 data chaotic laser data, predict next 100 data • Competition is described in Time Series Prediction: Forecasting the Future and Understanding the Past, A. S. Weigend & N. A. Gershenfeld, eds. , Addison-Wesley, 1994 • Method: - K-PLS with = 3 and 24 latent variables - Used records with 40 past data for training for next point - Predictions bootstrap on each other for 100 real test data • Entry “would have won” the competition
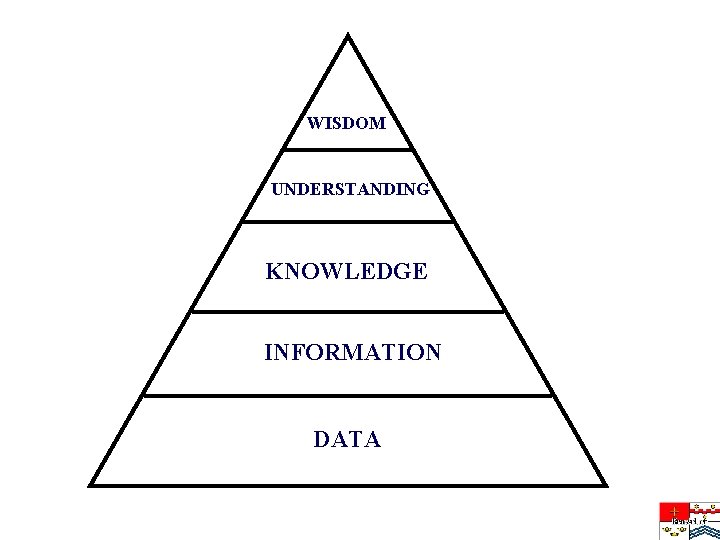
WISDOM UNDERSTANDING KNOWLEDGE INFORMATION DATA

Docking Ligands is a Nonlinear Problem
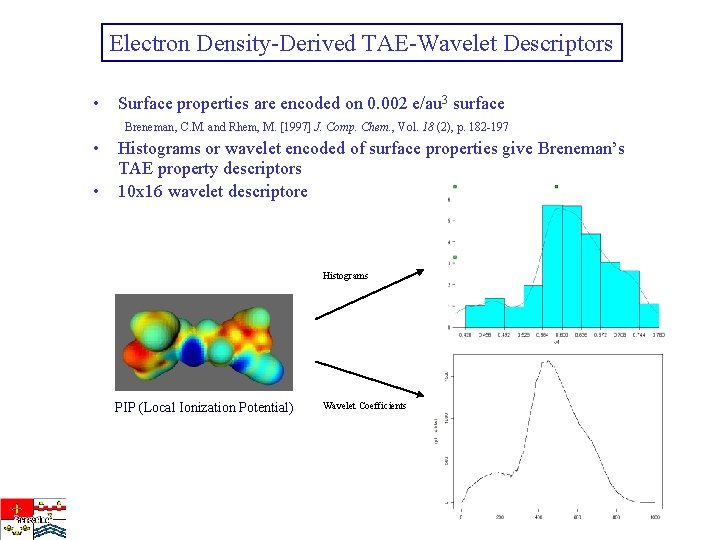
Electron Density-Derived TAE-Wavelet Descriptors • Surface properties are encoded on 0. 002 e/au 3 surface Breneman, C. M. and Rhem, M. [1997] J. Comp. Chem. , Vol. 18 (2), p. 182 -197 • Histograms or wavelet encoded of surface properties give Breneman’s TAE property descriptors • 10 x 16 wavelet descriptore Histograms PIP (Local Ionization Potential) Wavelet Coefficients
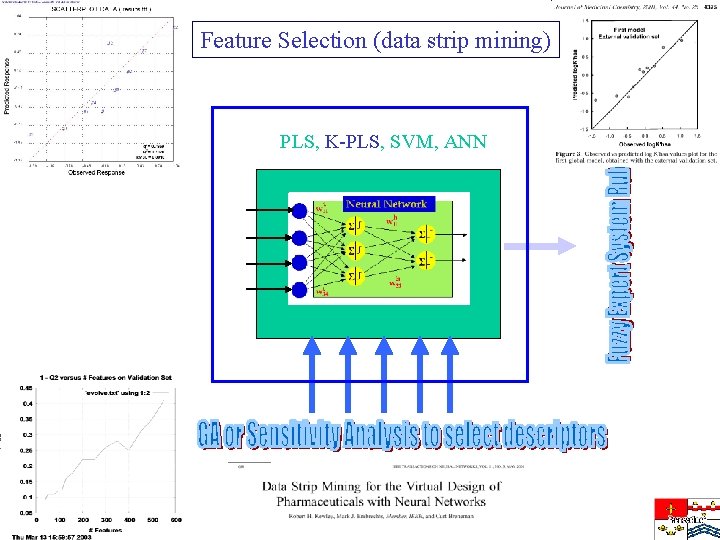
Feature Selection (data strip mining) PLS, K-PLS, SVM, ANN
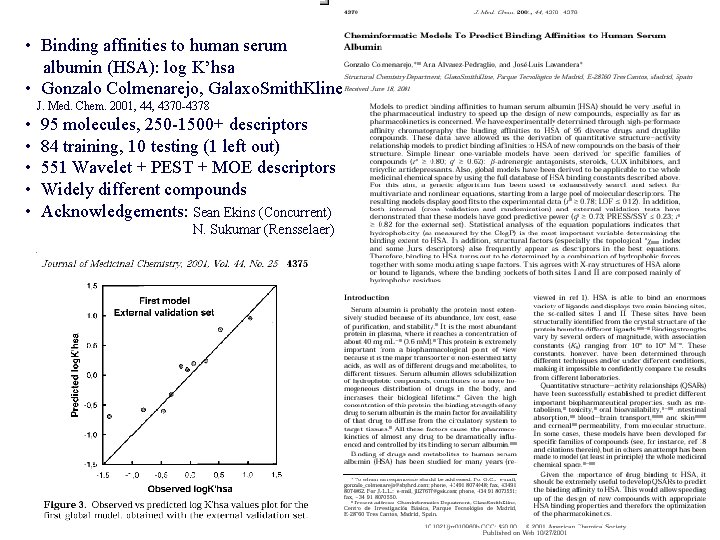
• Binding affinities to human serum albumin (HSA): log K’hsa • Gonzalo Colmenarejo, Galaxo. Smith. Kline J. Med. Chem. 2001, 44, 4370 -4378 • • • 95 molecules, 250 -1500+ descriptors 84 training, 10 testing (1 left out) 551 Wavelet + PEST + MOE descriptors Widely different compounds Acknowledgements: Sean Ekins (Concurrent) N. Sukumar (Rensselaer)
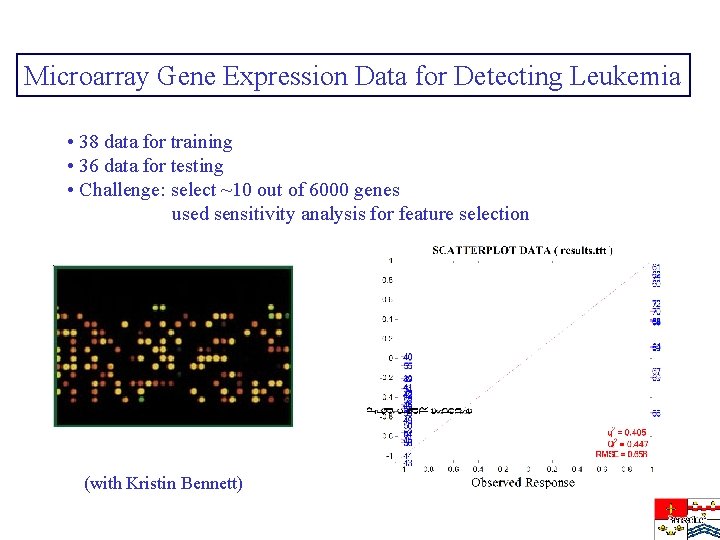
Microarray Gene Expression Data for Detecting Leukemia • 38 data for training • 36 data for testing • Challenge: select ~10 out of 6000 genes used sensitivity analysis for feature selection (with Kristin Bennett)

WORK IN PROGRESS GATCAATGAGGTGGACACCAGAGGCGGGGACTTGTAAATAACACTGGGCTGTAGGAGTGA TGGGGTTCACCTCTAATTCTAAGATGGCTAGATAATGCATCTTTCAGGGTTGTGCTTCTAGAAGGTAGAGCTGTGGTCGTTCAATAAAAGTCCTCAAGAGGTTAATACGCAT GTTTAATAGTACAGTATGGTGACTATAGTCAACAATAATTTATTGTACATTTTTAAATAG CTAGAAGAAAAGCATTGGGAAGTTTCCAACATGAAGAAAAGATAAATGGTCAAGGGAATG GATATCCTAATTACCCTGATTTGATCATTATGCATTATATACATGAATCAAAATATCACA CATACCTTCAAACTATGTACAAATATTATATACCAATAAAAAATCATCATCTCC ATCATCACCACCCTCCTCCTCATCACCACCAGCATCACCACCATCATCACCACCACCATC ATCACCACTGCCATCATCATCACCACCACTG TCATTATCACCACCACCATCATCACCAACACCACTGCCATCGTCATCACCACCACTGTCA TTATCACCACCACCATCACCAACATCACCACCACCATTATCACCACCATCAACACCACCA CCCCCATCATCACTACTACCATCATTACCAGCACCACTATCACCACCACAATCACCACTATCATCAACATCATCACTACCACCATCACCAACACCA CCATCATTATCACCACCATCACCAACATCACCACCATCATCATCACCACCATCA CCAAGACCATCATCATCACCACCAACATCACCACCATCACCAACACCACCATCA CCACCACCATCATCACCACCATCATCATCACCACCACCGCCATCATCA TCGCCACCACCATGACCACCACCATCACAACCATCACCACCATCACAACCACCATCATCA CTATCGCTATCACCACCATTACCACCACCATTACTACAACCATGACCATCACCACCACCATCACAACGATCACCATCACAGCCACCATCATCACCACCATCAAACCATCGGCATTATTATTTTTTTAGAATTTTGTTGGGATTCAGT ATCTGCCAAGATACCCATTCTTAAAACATGAAAAAGCAGCTGACCCTCCTGTGGCCCCCT TTTTGGGCAGTCATTGCAGGACCTCATCCCCAAGCAGCAGCTCTGGTGGCATACAGGCAA CCCACCACCAAGGTAGAGGGTAATTGAGCAGAAAAGCCACTTCCTCCAGCAGTTCCCTGT
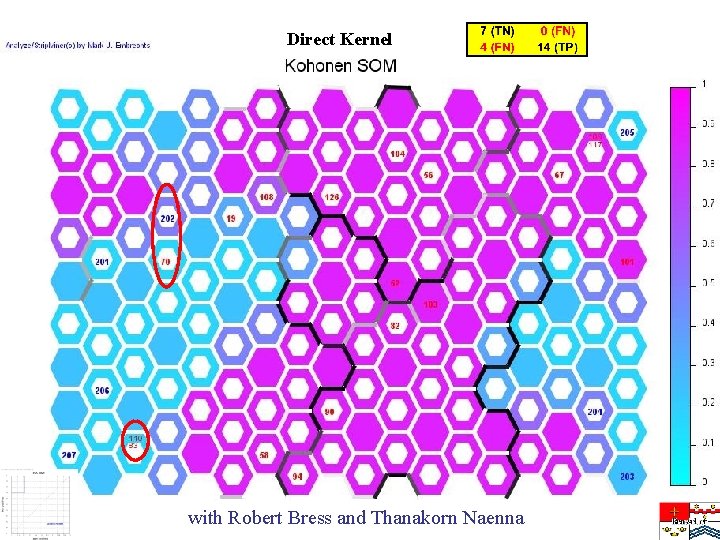
Direct Kernel with Robert Bress and Thanakorn Naenna
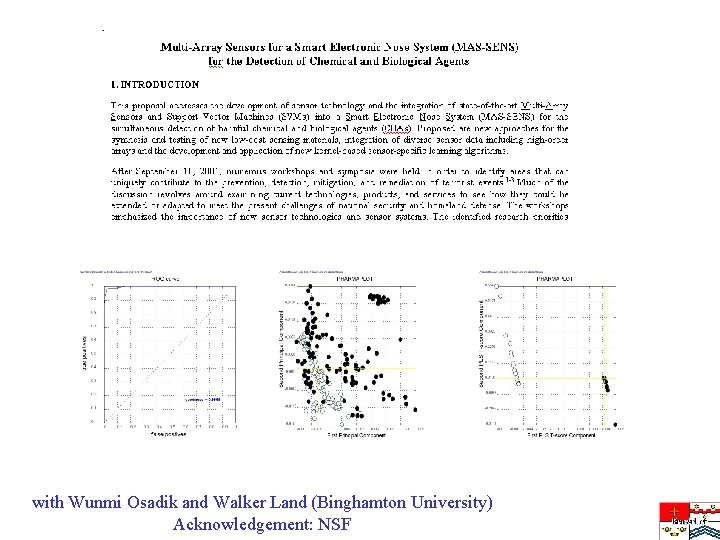
with Wunmi Osadik and Walker Land (Binghamton University) Acknowledgement: NSF

Magneto-cardiogram Data with Karsten Sternickel (Cardiomag Inc. ) and Boleslaw Szymanski (Rensselaer) Acknowledgemnent: NSF SBIR phase I project

Direct Kernel PLS with 3 Latent Variables
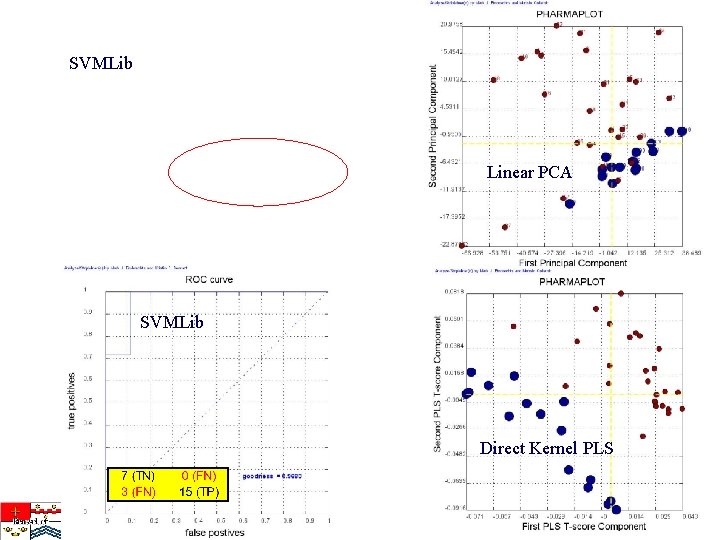
SVMLib Linear PCA SVMLib Direct Kernel PLS

www. drugmining. com Kristin Bennett and Mark Embrechts
- Slides: 27