SNPs and CNPs By David Wendel What are

SNPs and CNPs By: David Wendel
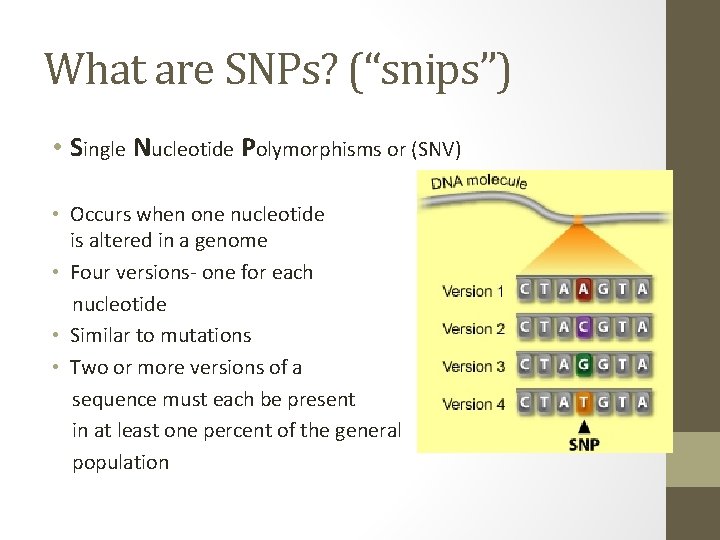
What are SNPs? (“snips”) • Single Nucleotide Polymorphisms or (SNV) • Occurs when one nucleotide is altered in a genome • Four versions- one for each nucleotide • Similar to mutations • Two or more versions of a sequence must each be present in at least one percent of the general population

What are SNPs? (cont. ) • Many have no effect on cell function • Do not directly cause disease • Most found outside of protein sequences • Likely to alter protein functions if inside • Help pinpoint disease on human genome map • Evolutionarily stable
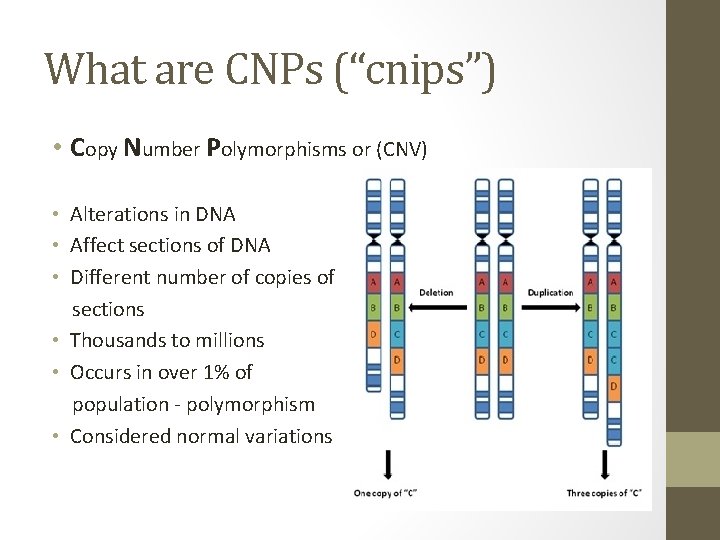
What are CNPs (“cnips”) • Copy Number Polymorphisms or (CNV) • Alterations in DNA • Affect sections of DNA • Different number of copies of sections • Thousands to millions • Occurs in over 1% of population - polymorphism • Considered normal variations

SNPs and CNPs in Humans • Human DNA is very similar • 99. 6% of human DNA is identical to all other humans • Other 0. 4% is made up of CNPs and SNPs • SNPs- 80% of the 0. 4% Ø Small scale variant • CNPs- 20% of the 0. 4% Ø Large scale variant

How do we find SNPs? • Two main ways scientists identify, categorize, and catalog SNPs q. Genomic Approaches q. Functional Approaches • Primer Extension is also used

Genomic Approaches • Scientists who want to see the big picture • Hundreds of scientists involved • Genomes of numerous individuals are compared • Computer-powered data analysis • Results available to anyone on internet

Functional Approaches • Scientists interested in particular diseases or drug responses 1. Select genes known in a process 2. Examine the genes in people with and without a disease 3. Identify SNPs that correspond with particular response
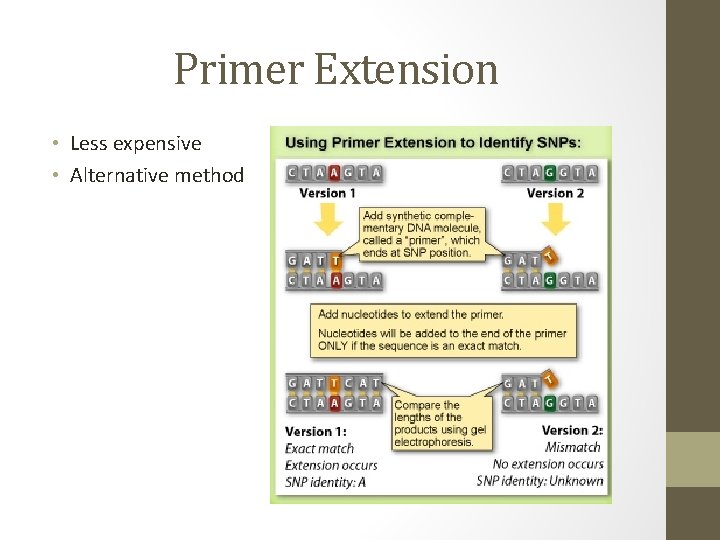
Primer Extension • Less expensive • Alternative method

Haplotypes • A set of single nucleotide polymorphisms • Set is on a single chromosome of a chromosome pair that is statistically associated • Associations can identify other polymorphic sites in the region • SNPs close together on the chromosome • Valuable for investigating genetics behind diseases
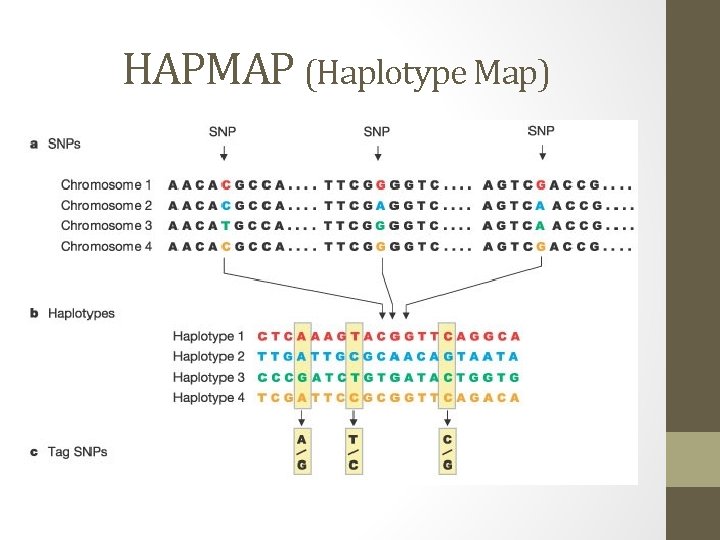
HAPMAP (Haplotype Map)

HAPMAP (Haplotype Map) cont. • Hapmap- a catalog of common genetic variants that occur in human beings o What the variants are o Where the variants occur in our DNA o How they are distributed among populations • Able to see how SNPs are organized on chromosomes • Construction of Hapmap occurs in three steps: 1. Single Nucleotide Polymorphisms 2. Adjacent SNPs inherited together compiled into haplotypes 3. “Tag” SNPs in haplotypes identified that uniquely identify those haplotypes.

Hapmap I 2005 • This was the first phase of the project • The project was launched in October 2002 • Original goal to be finished was September 2005 • Finished first draft in February 2005 • Contained 1, 000 most common SNPs

Hapmap II 2006 • Results from Phase II were released in August 2006 • These results were published in February 2007 • 10, 000 SNPs were included in this Hapmap

Hapmap III 2010 • The third phase was published in 2010 • Data from 1, 184 individuals • 11 global populations • 1, 410, 616 SNPs were released for detailed studies • Ten 100 -kilobase ENCODE regions were sequenced in 692 of these individuals • Allowed integration of SNPs and CNPs

Average Person and SNPs • Density of SNPs per base pair varies with every person • SNPs occur about one in every 500 - 1000 base pairs • About 3, 750, 000 total
- Slides: 16