Slicer 3 Training Compendium PETCT Analysis using 3









- Slides: 23

Slicer 3 Training Compendium PET/CT Analysis using 3 D Slicer Jeffrey Yap Ph. D Ron Kikinis MD Wendy Plesniak Ph. D Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -1 -

Study Data CT acquisition: (jeff, please fill in anything relevant about the dataset) PET acquisition: (jeff, please fill in anything relevant about the dataset, Isotope used, etc. ) Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -2 -

Clinical Context Clinical relevance: (jeff) -underlying diagnosis -why were scans done -what was radiologist’s interpretation -treatment? -followup plan. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -3 -

Clinical Context Tutorial context: -background: standard ways to evaluate response to treatment -how this tutorial addresses clinical needs -transition to specifics on SUV computation. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -4 -

Brief Description of Quantitative Measurement: Standardized Uptake Value SUV (time) = Radioactive Concentration x Weight Injected Activity • Under certain circumstances, 18 FDG SUV correlates with metabolic rate of glucose and/or the number of viable tumor cells • Simplified semi-quantitative measure that can be routinely performed in clinical PET studies • Adjusts for differences in patient size and injected activity Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -5 -

Making assessments from measurements: Response criteria • Complete response (CR): Complete resolution of all lesions • Partial Response (PR): >= 25% decrease in SUVmax • Stable Disease (SD): < 25% change • Progressive Disease (PD): >= 25% increase in SUVmax Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -6 -

Learning objective This tutorial demonstrates how a pre-treatment and followup study are analyzed. Following this simple tutorial, you’ll be able to use 3 D Slicer to: • Load a MRML scene description file, • Achieve combined display of PET and CT data, • Make quantitative measurements of Standardized Uptake Value (SUV) within one or more volumes of interest Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -7 -

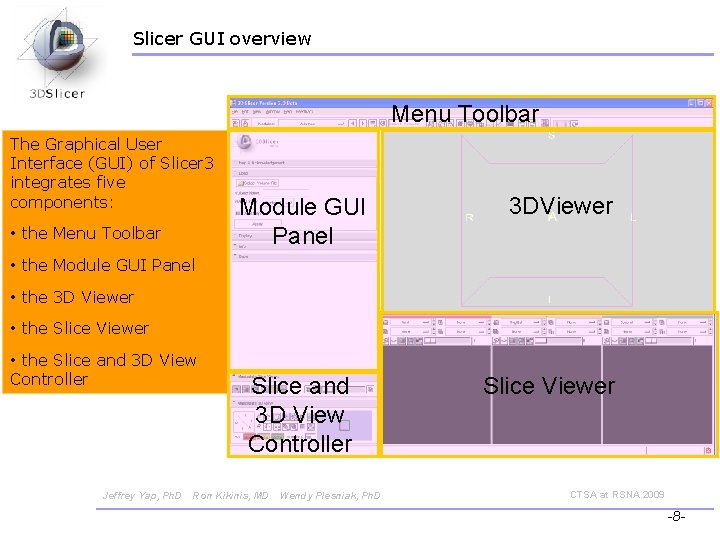
Slicer GUI overview Menu Toolbar The Graphical User Interface (GUI) of Slicer 3 integrates five components: • the Menu Toolbar Module GUI Panel 3 DViewer • the Module GUI Panel • the 3 D Viewer • the Slice and 3 D View Controller Jeffrey Yap, Ph. D Slice and 3 D View Controller Ron Kikinis, MD Wendy Plesniak, Ph. D Slice Viewer CTSA at RSNA 2009 -8 -

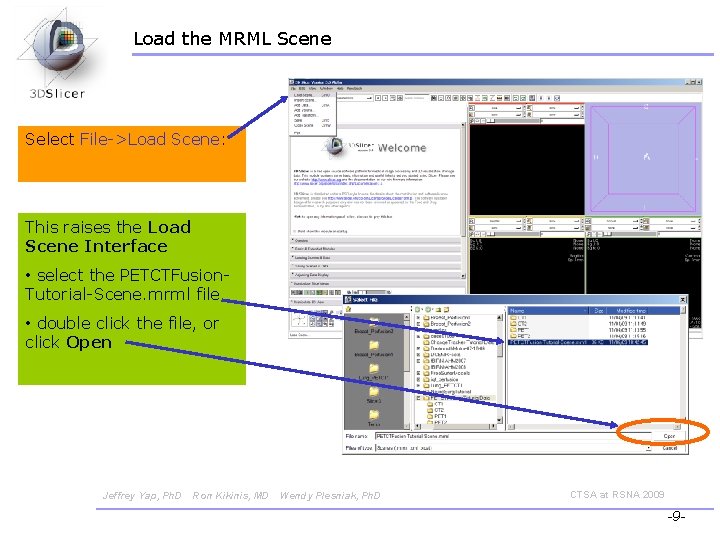
Load the MRML Scene Select File->Load Scene: This raises the Load Scene Interface • select the PETCTFusion. Tutorial-Scene. mrml file • double click the file, or click Open Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -9 -

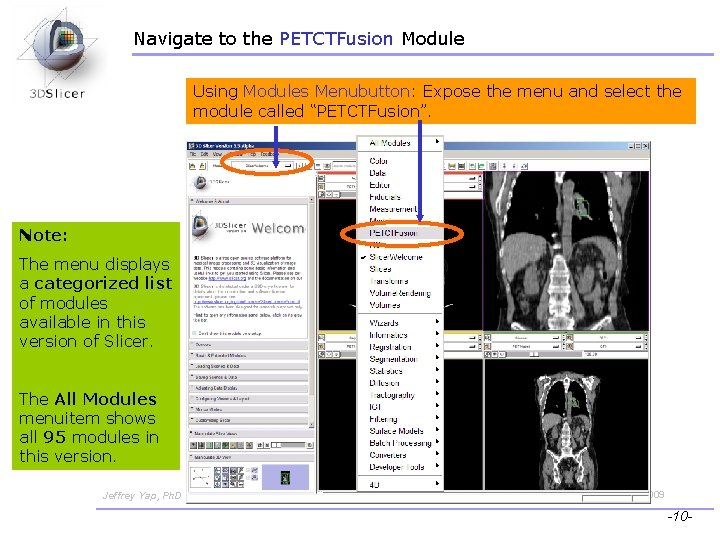
Navigate to the PETCTFusion Module Using Modules Menubutton: Expose the menu and select the module called “PETCTFusion”. Note: The menu displays a categorized list of modules available in this version of Slicer. The All Modules menuitem shows all 95 modules in this version. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -10 -

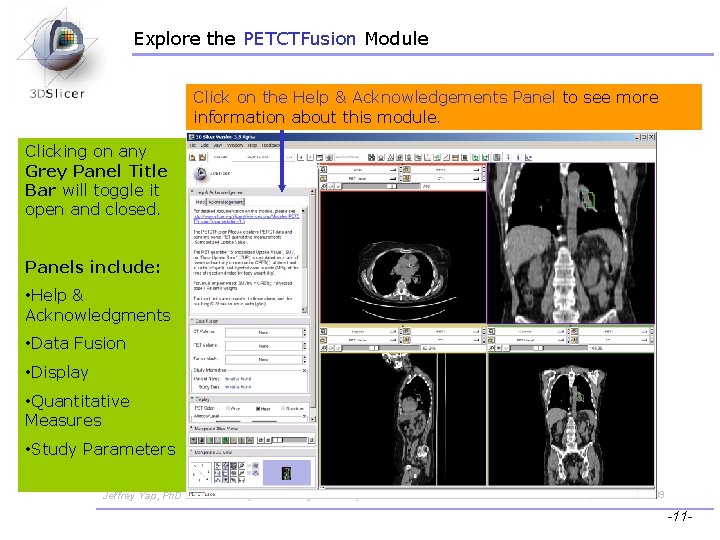
Explore the PETCTFusion Module Click on the Help & Acknowledgements Panel to see more information about this module. Clicking on any Grey Panel Title Bar will toggle it open and closed. Panels include: • Help & Acknowledgments • Data Fusion • Display • Quantitative Measures • Study Parameters Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -11 -

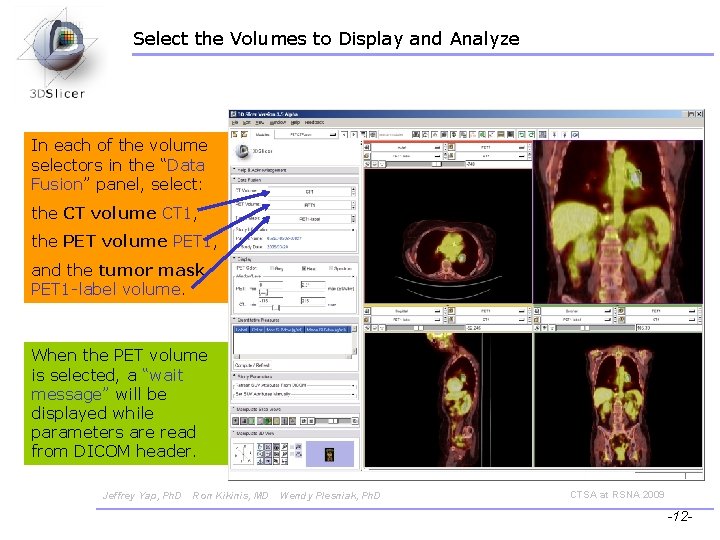
Select the Volumes to Display and Analyze In each of the volume selectors in the “Data Fusion” panel, select: the CT volume CT 1, the PET volume PET 1, and the tumor mask PET 1 -label volume. When the PET volume is selected, a “wait message” will be displayed while parameters are read from DICOM header. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -12 -

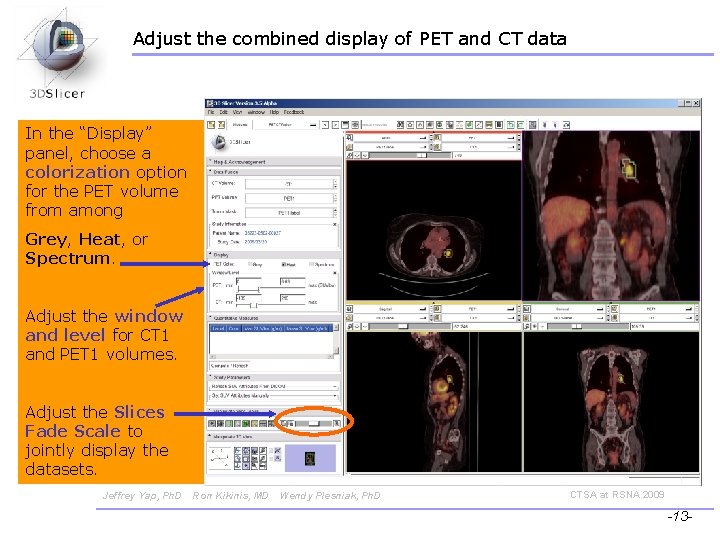
Adjust the combined display of PET and CT data In the “Display” panel, choose a colorization option for the PET volume from among Grey, Heat, or Spectrum. Adjust the window and level for CT 1 and PET 1 volumes. Adjust the Slices Fade Scale to jointly display the datasets. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -13 -

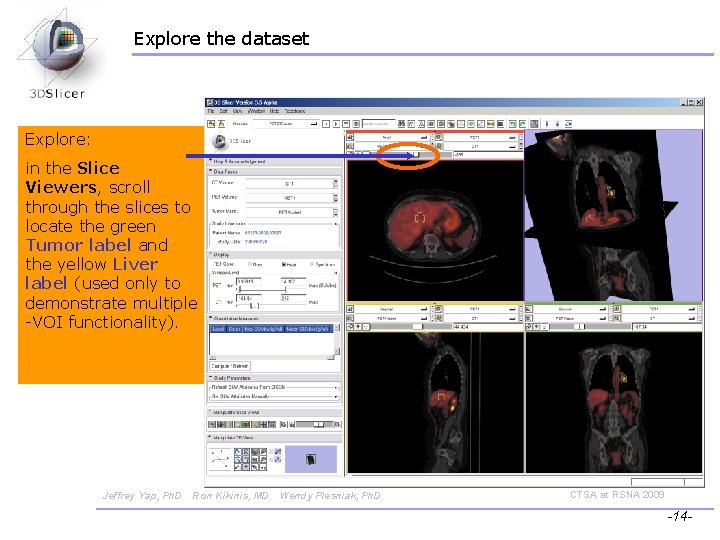
Explore the dataset Explore: in the Slice Viewers, scroll through the slices to locate the green Tumor label and the yellow Liver label (used only to demonstrate multiple -VOI functionality). Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -14 -

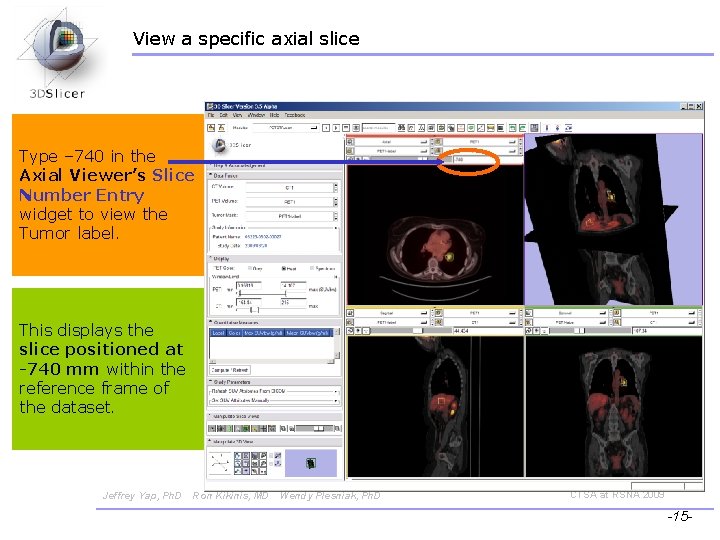
View a specific axial slice Type – 740 in the Axial Viewer’s Slice Number Entry widget to view the Tumor label. This displays the slice positioned at -740 mm within the reference frame of the dataset. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -15 -

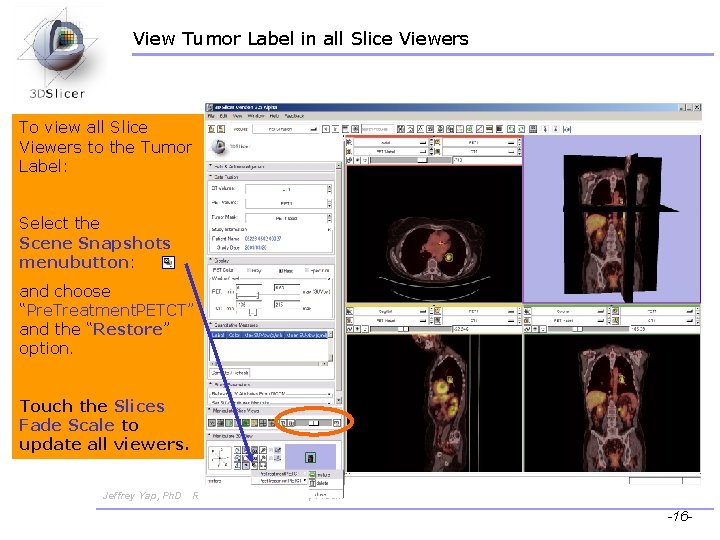
View Tumor Label in all Slice Viewers To view all Slice Viewers to the Tumor Label: Select the Scene Snapshots menubutton: and choose “Pre. Treatment. PETCT” and the “Restore” option. Touch the Slices Fade Scale to update all viewers. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -16 -

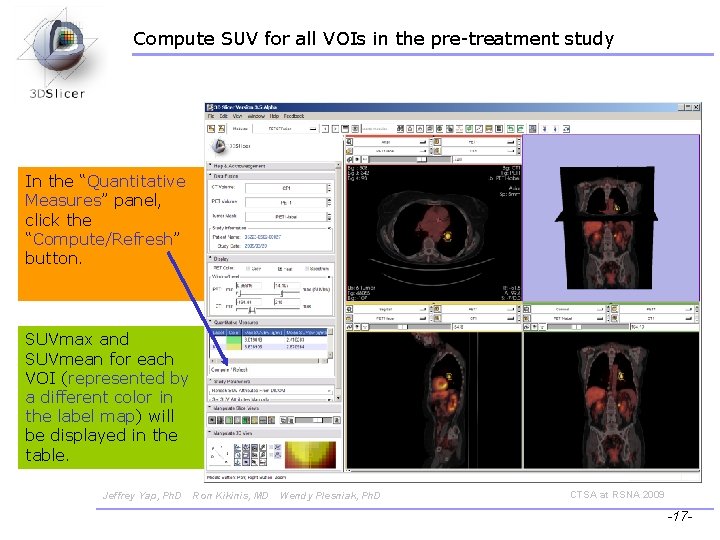
Compute SUV for all VOIs in the pre-treatment study In the “Quantitative Measures” panel, click the “Compute/Refresh” button. SUVmax and SUVmean for each VOI (represented by a different color in the label map) will be displayed in the table. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -17 -

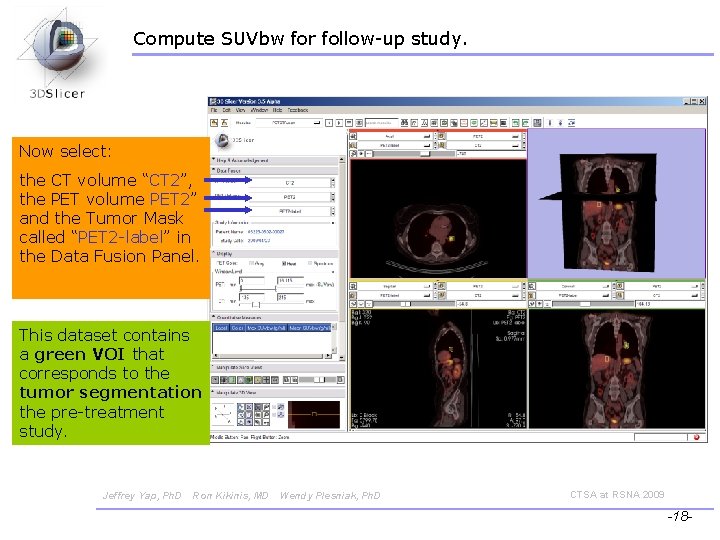
Compute SUVbw for follow-up study. Now select: the CT volume “CT 2”, the PET volume PET 2” and the Tumor Mask called “PET 2 -label” in the Data Fusion Panel. This dataset contains a green VOI that corresponds to the tumor segmentation the pre-treatment study. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -18 -

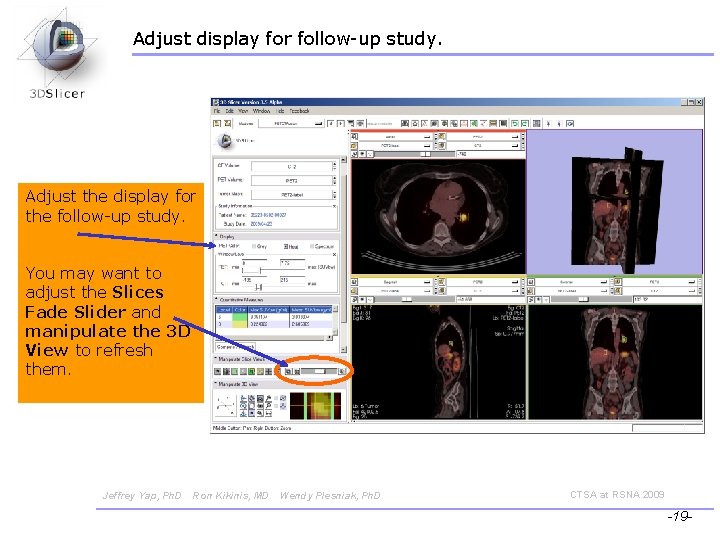
Adjust display for follow-up study. Adjust the display for the follow-up study. You may want to adjust the Slices Fade Slider and manipulate the 3 D View to refresh them. Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -19 -

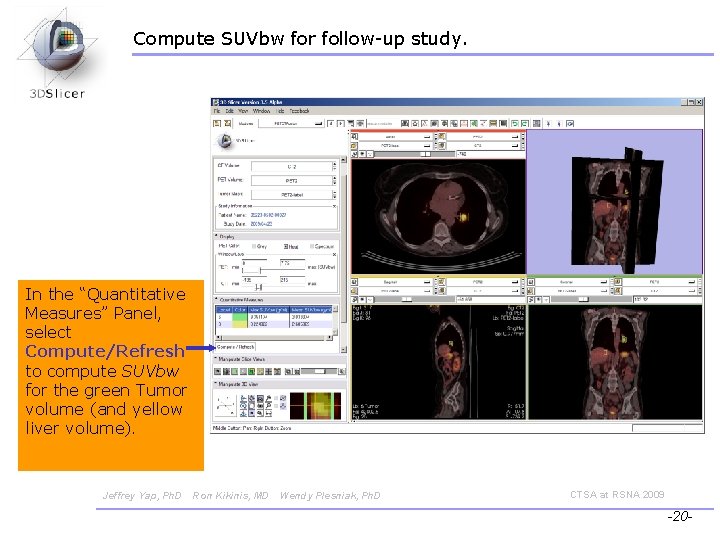
Compute SUVbw for follow-up study. In the “Quantitative Measures” Panel, select Compute/Refresh to compute SUVbw for the green Tumor volume (and yellow liver volume). Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -20 -

Assess response with respect to this VOI Pre-Treatment Max SUVbw = 8. 019048 g/ml Post-Treatment Max SUVbw = 9. 351174 g/ml +16. 61% (SD) Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -21 -

Summary This tutorial has demonstrated: • use of an interactive interface to load a scene • the combined display of PET and CT volumes in a single visualization • a workflow to make quantitative measurements of Standardized Uptake Value (SUV) within a volume of interest Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -22 -

Acknowledgments Harvard Clinical and Translational Science Center National Alliance for Medical Image Computing (NA-MIC) Neuroimage Analysis Center (NAC) National Center for Image-Guided Therapy (NCIGT) Surgical Planning Laboratory, Brigham and Women’s Hospital Jeffrey Yap, Ph. D Ron Kikinis, MD Wendy Plesniak, Ph. D CTSA at RSNA 2009 -23 -