Singlecell Analysis Systematic Biology 2018 05 04 1

Single-cell Analysis Systematic Biology 2018 -05 -04 1

BACKGROUND • Single cell gene expression Full length: Fluidigm C 1, Smart-seq Single end: Drop-seq, Genomics 10 X, Microwell-seq, SPLi. T-seq • Single cell chromatin accessibility sc-ATAC-seq, sci-ATAC-seq, sc-THS-seq • Sparse signal matrix Normalization, Dimension reduction 2
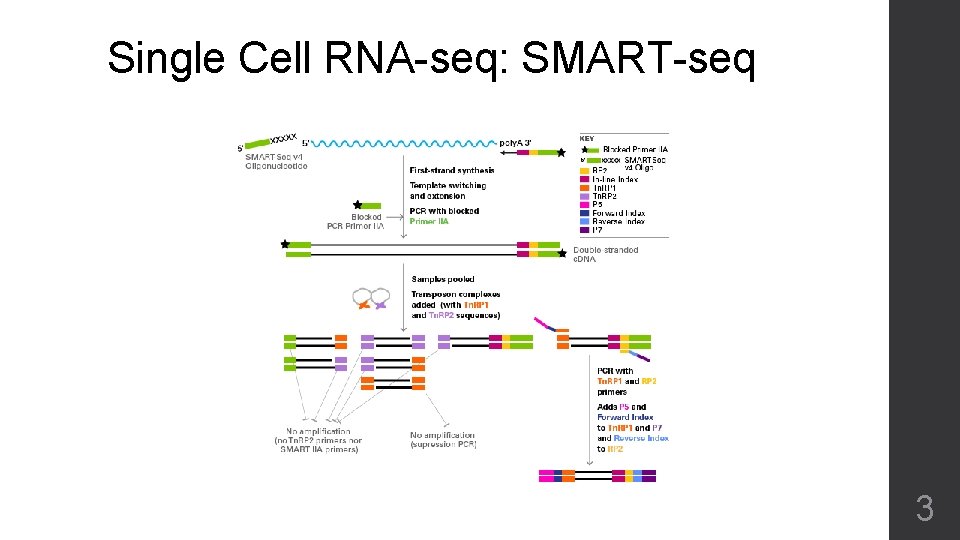
Single Cell RNA-seq: SMART-seq 3
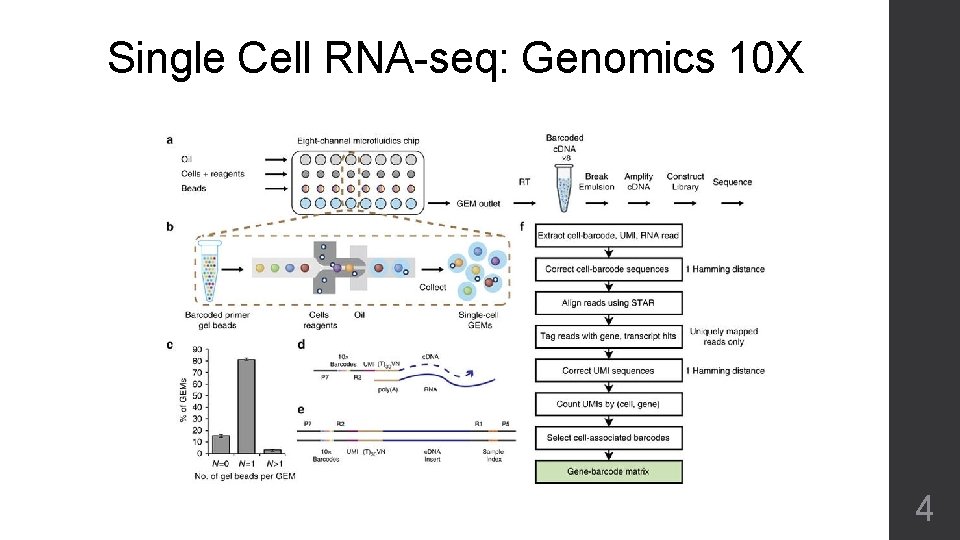
Single Cell RNA-seq: Genomics 10 X 4
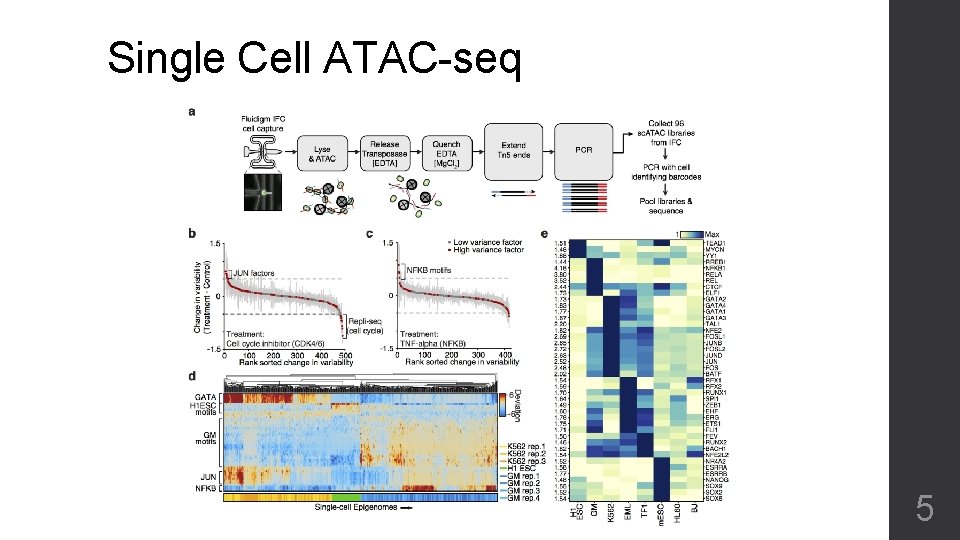
Single Cell ATAC-seq 5
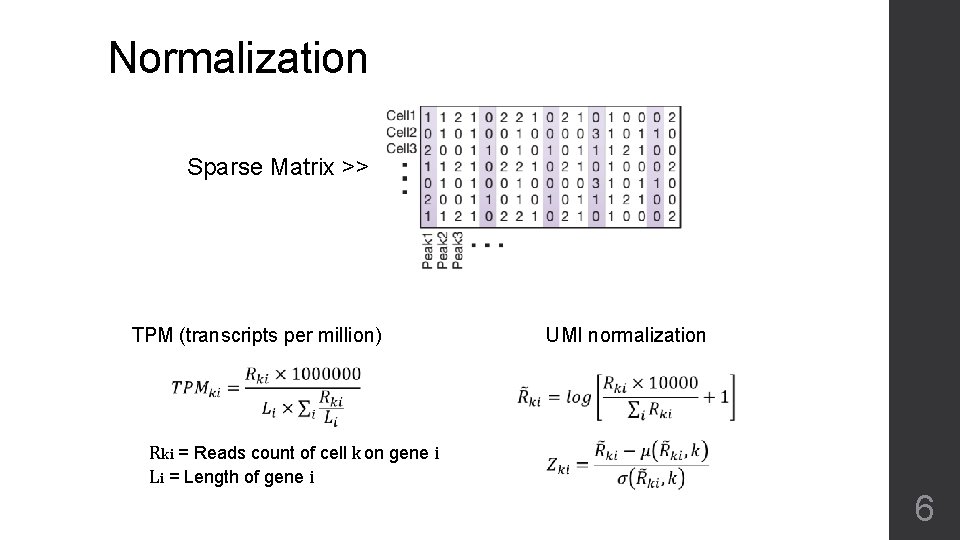
Normalization Sparse Matrix >> UMI normalization TPM (transcripts per million) Rki = Reads count of cell k on gene i Li = Length of gene i 6
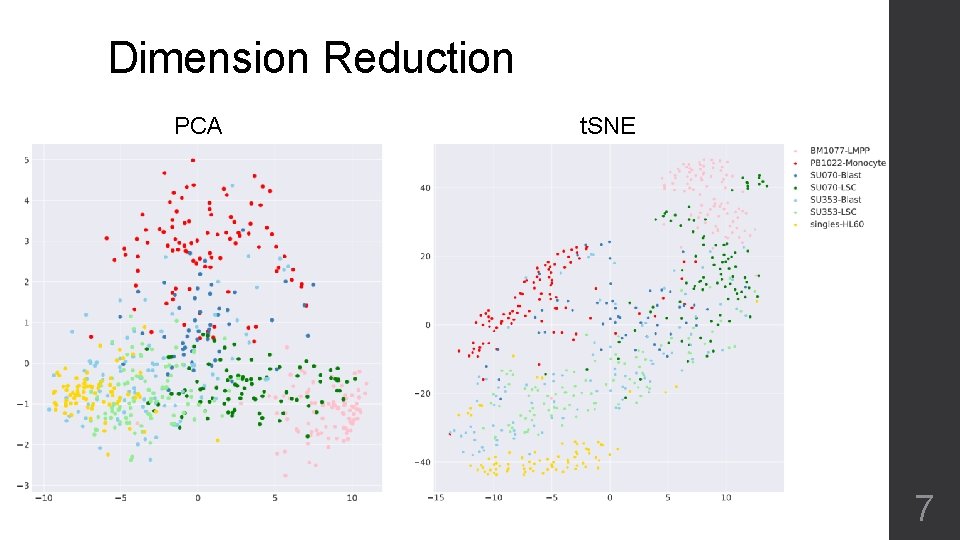
Dimension Reduction PCA t. SNE 7

Single Cell RNA-seq Analysis • Cell Ranger: Mapping and Preparation • Seurat: Filtering and KNN clustering • Scenic: Group Genes by Motif • SIMLR: Multiple Kernels • SC 3: Consensus Clustering with Multiple Transformations • Monocle: Pseudo-time Trajectory by Minimum Spanning Tree 8
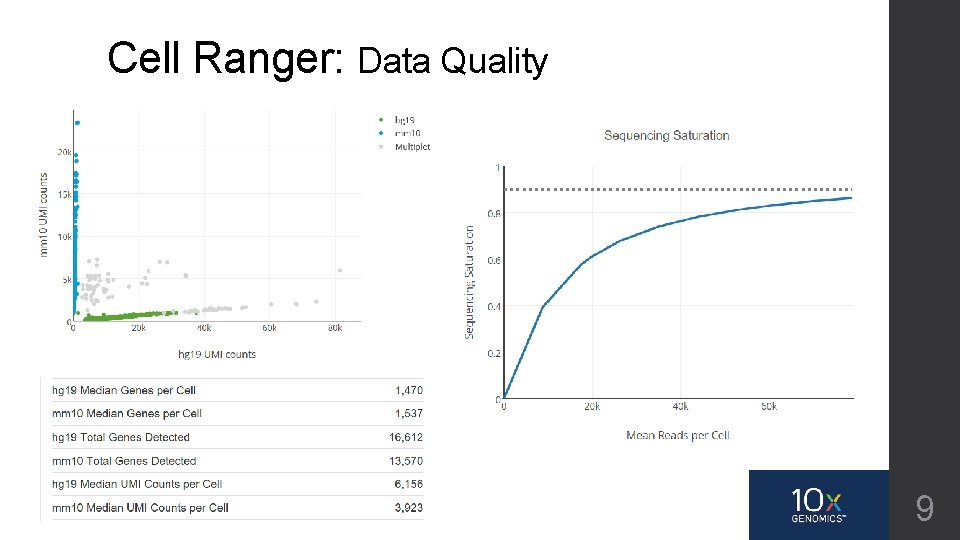
Cell Ranger: Data Quality 9
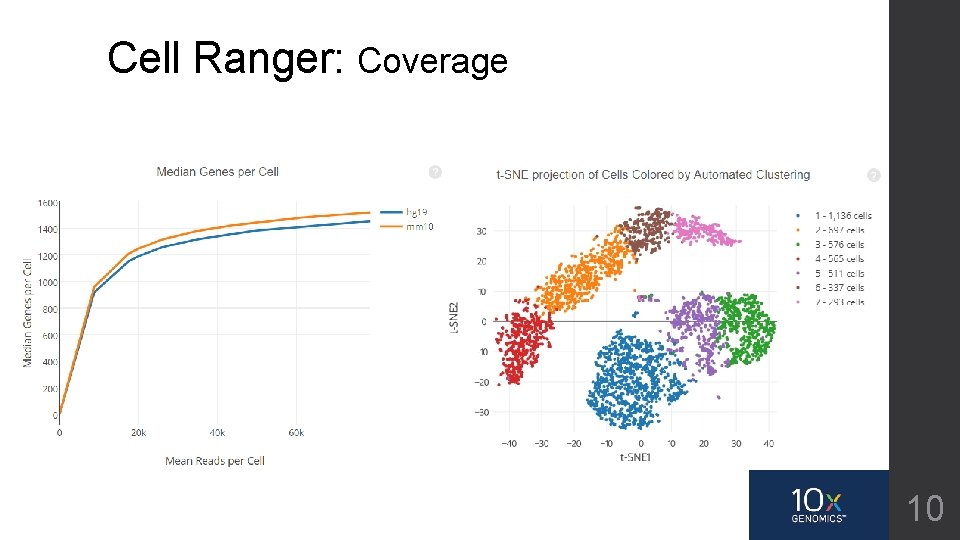
Cell Ranger: Coverage 10

Seurat: R toolkit for single cell genomics Satjia Lab: Nature Biotech. 2018: New York Genome Center NYU Center for Genomics and Systems Biology Integrating single-cell transcriptomic data across different conditions, technologies, and species 11
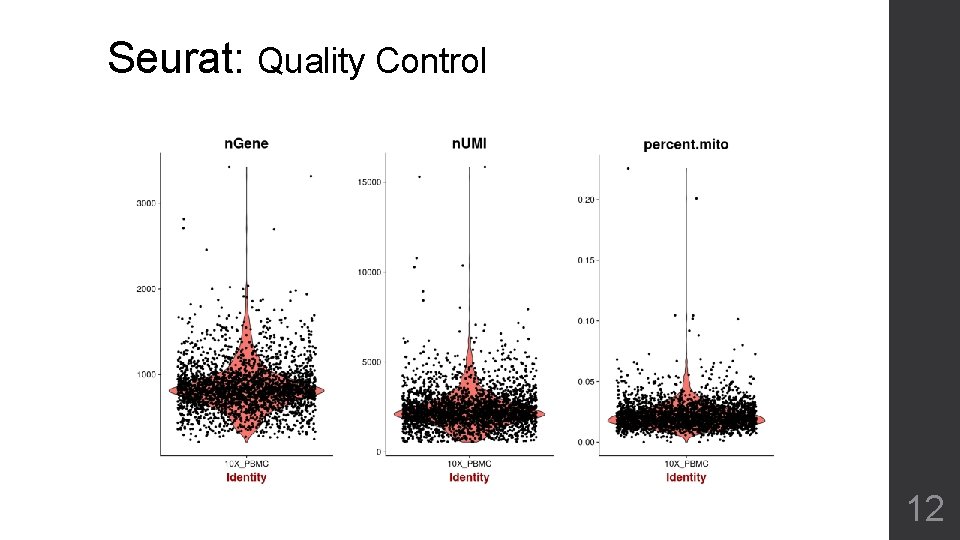
Seurat: Quality Control 12
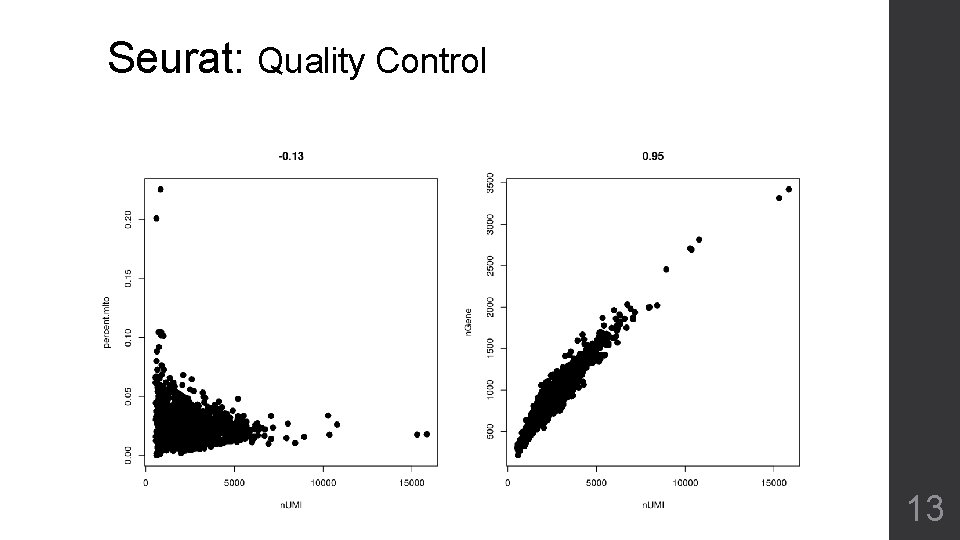
Seurat: Quality Control 13
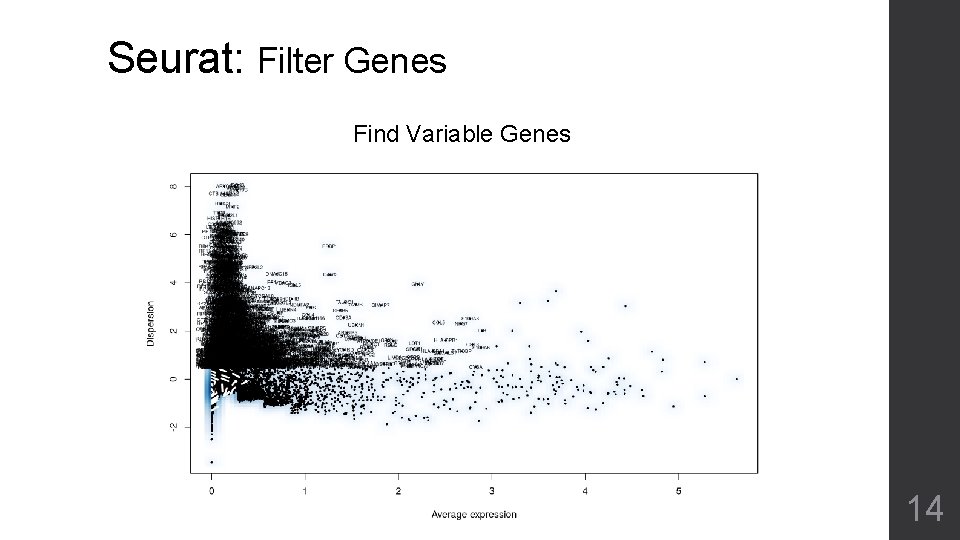
Seurat: Filter Genes Find Variable Genes 14
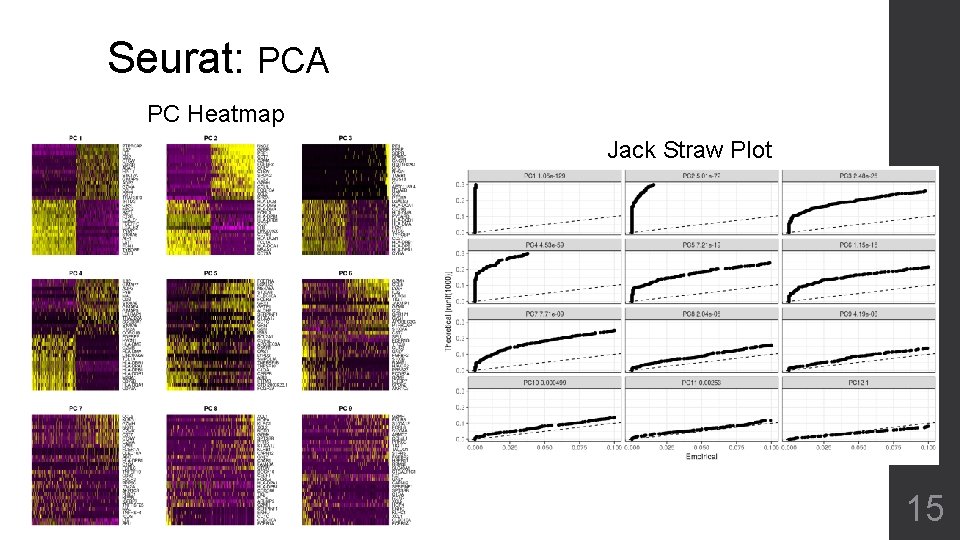
Seurat: PCA PC Heatmap Jack Straw Plot 15
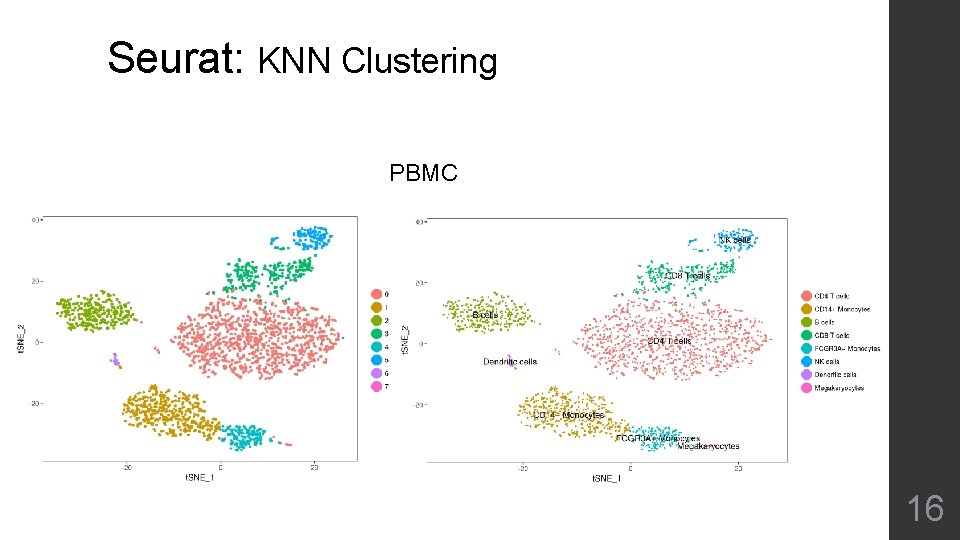
Seurat: KNN Clustering PBMC 16
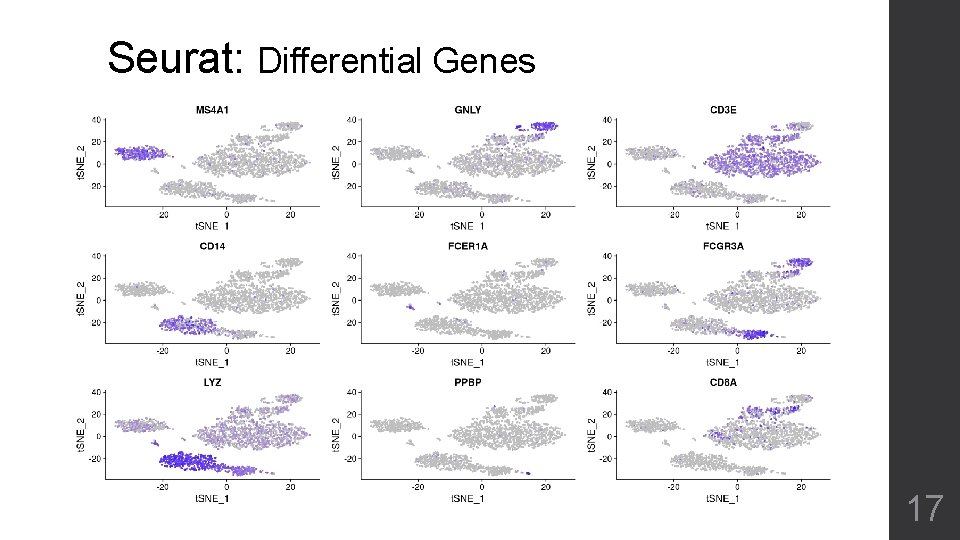
Seurat: Differential Genes 17
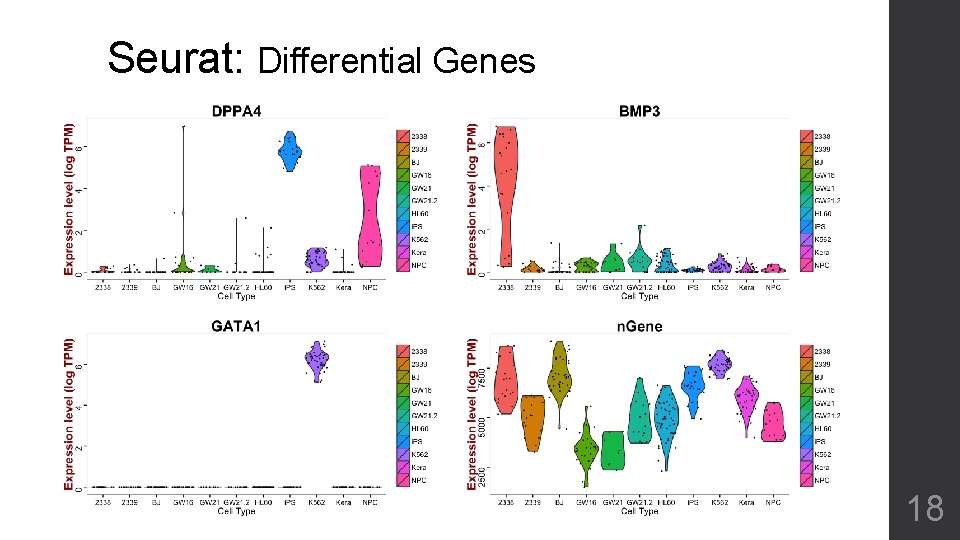
Seurat: Differential Genes 18
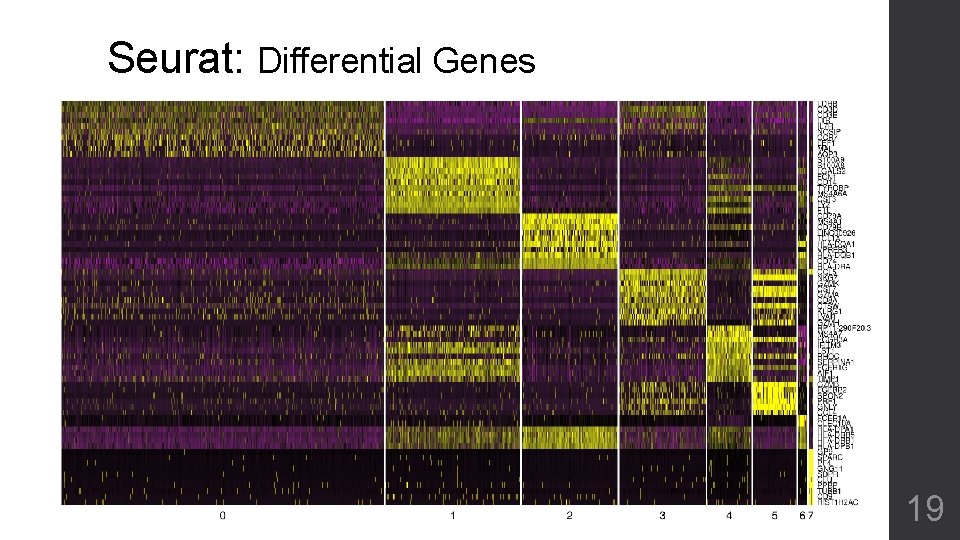
Seurat: Differential Genes 19
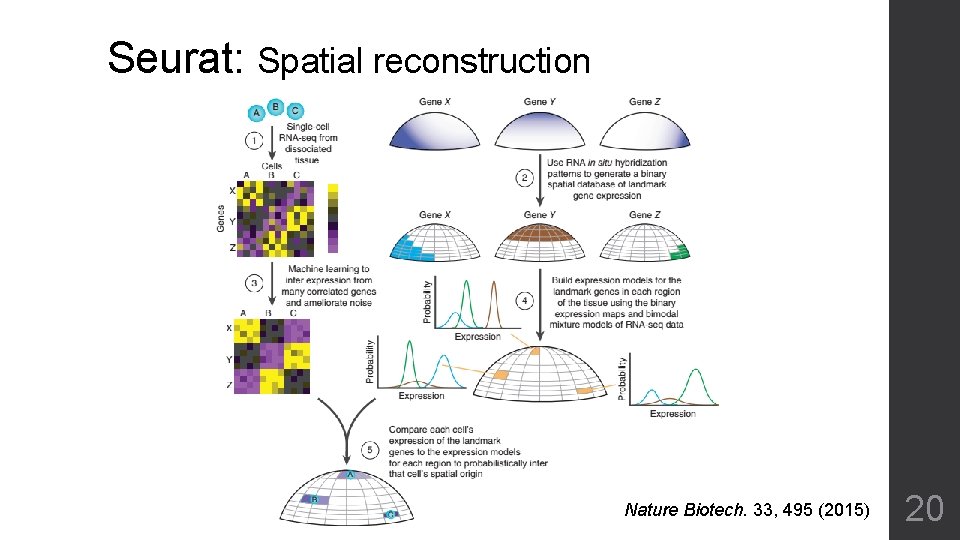
Seurat: Spatial reconstruction Nature Biotech. 33, 495 (2015) 20
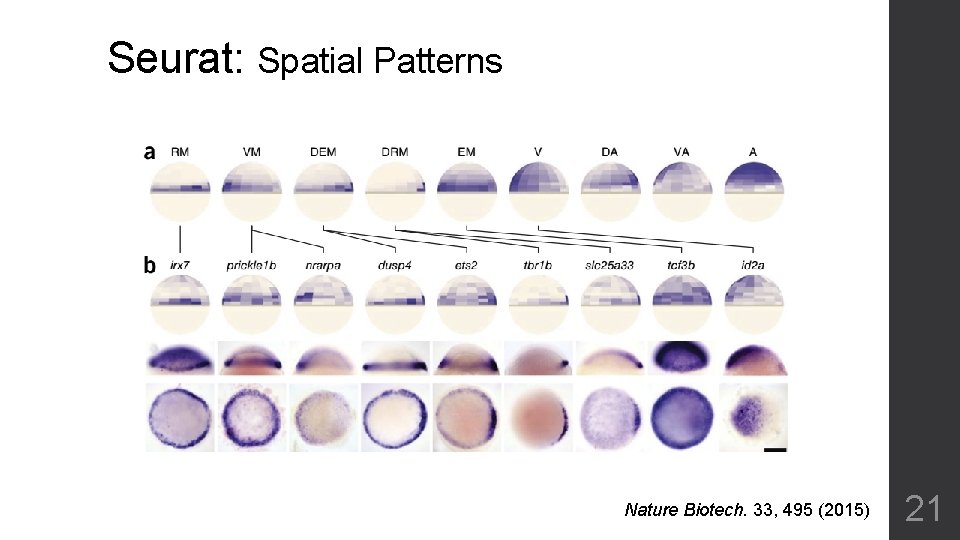
Seurat: Spatial Patterns Nature Biotech. 33, 495 (2015) 21

Seraut • Tools to filter low quality samples • Provides good clustering for significant different cell types • Detailed principle component analysis • Predict cluster number by Louvain algorithm • Ignore genes by a simple estimation of dispersion • Nor fit for expression matrix too sparse 22
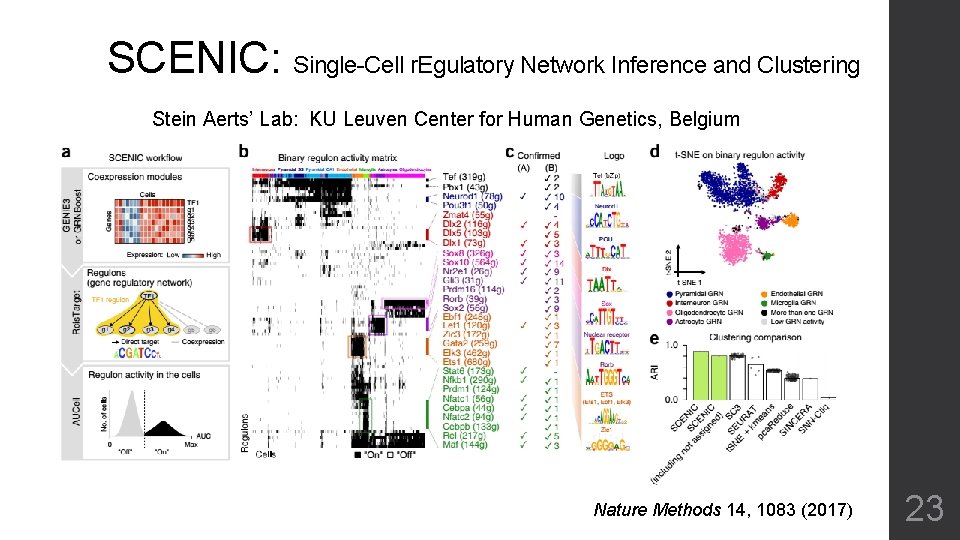
SCENIC: Single-Cell r. Egulatory Network Inference and Clustering Stein Aerts’ Lab: KU Leuven Center for Human Genetics, Belgium Nature Methods 14, 1083 (2017) 23
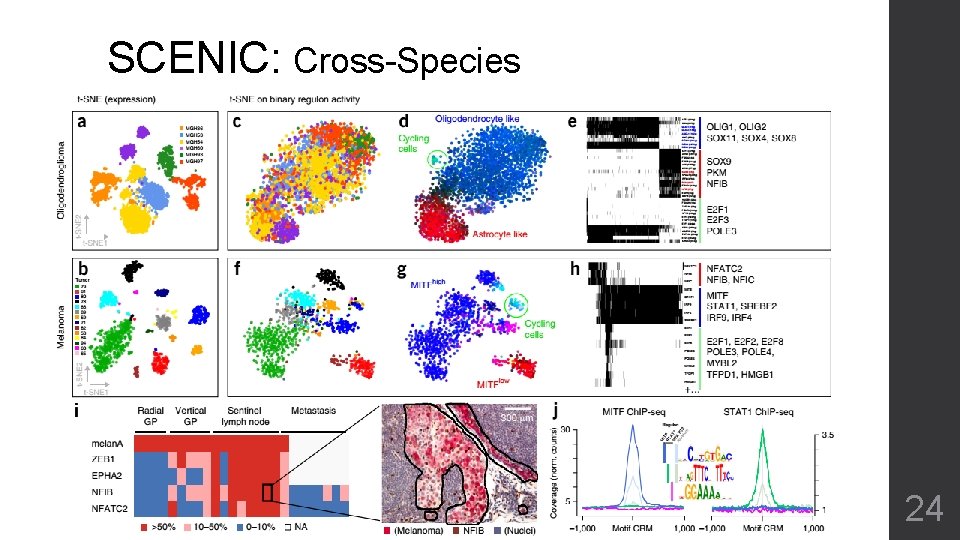
SCENIC: Cross-Species 24

SCENIC • Reveals the TFs that regulate expression patterns • Naturally merge sample from different origins • Avoid batch effect • Cannot provides proper sub-type clustering • Some patterns are concealed by the sum of motif-sites 25
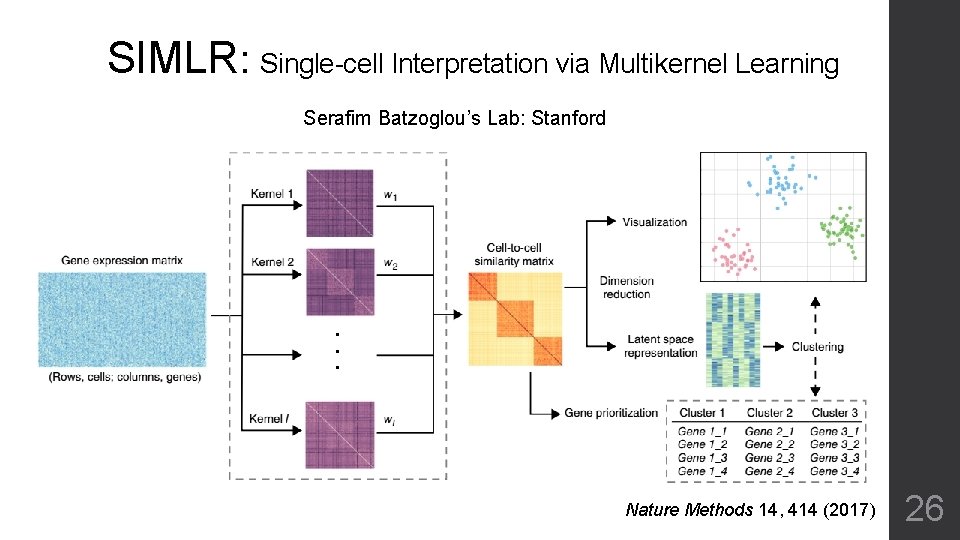
SIMLR: Single-cell Interpretation via Multikernel Learning Serafim Batzoglou’s Lab: Stanford Nature Methods 14, 414 (2017) 26
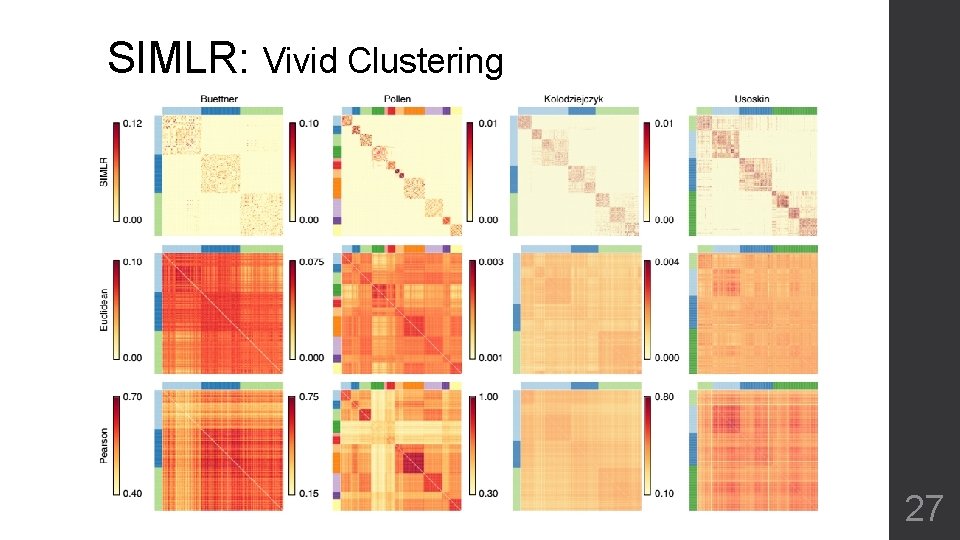
SIMLR: Vivid Clustering 27
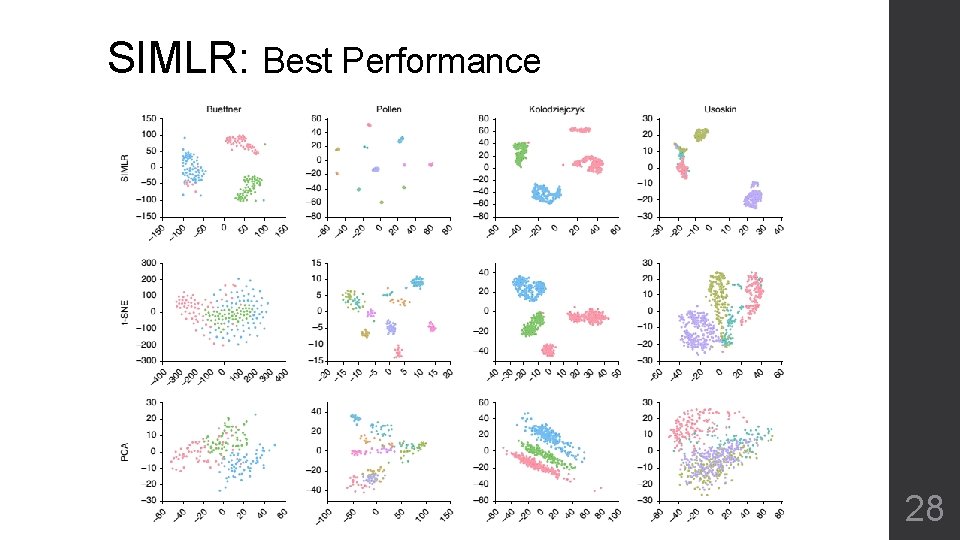
SIMLR: Best Performance 28

SIMLR: Algorithm 29

SIMLR • Very clear clustering of cell sub-types • Transformation that can be used for next step analysis • Artificial parameters (β, γ) for convergence • Have to define number of blocks (C) beforehand • Over clustering by itself 30
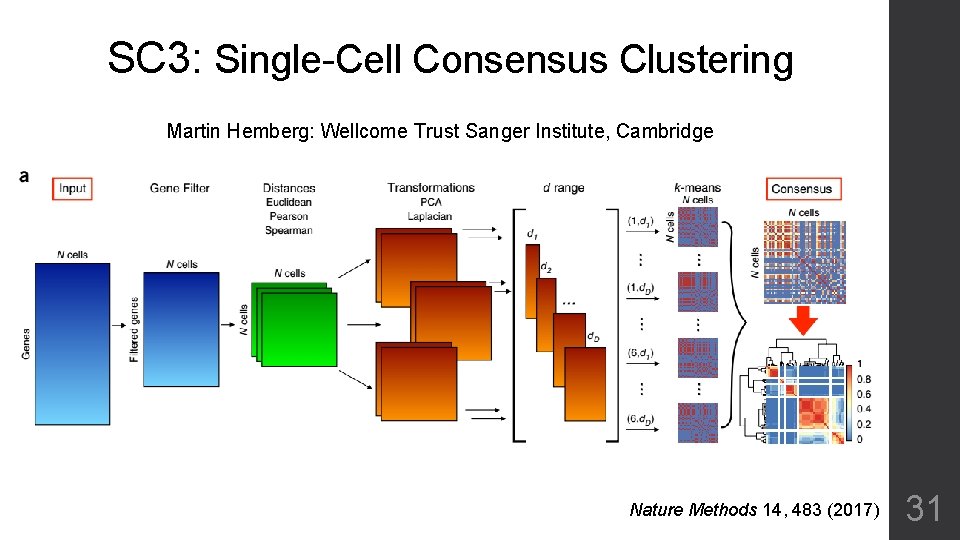
SC 3: Single-Cell Consensus Clustering Martin Hemberg: Wellcome Trust Sanger Institute, Cambridge Nature Methods 14, 483 (2017) 31
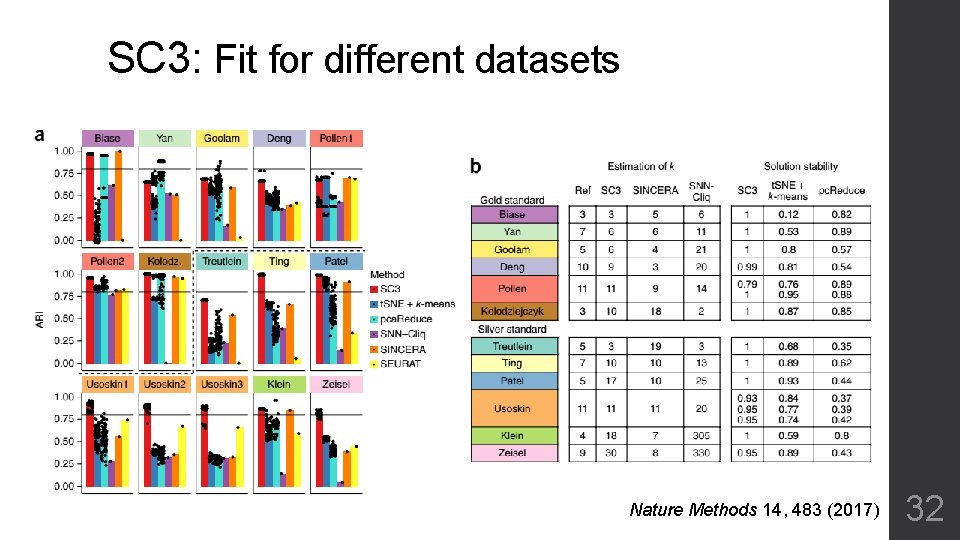
SC 3: Fit for different datasets Nature Methods 14, 483 (2017) 32
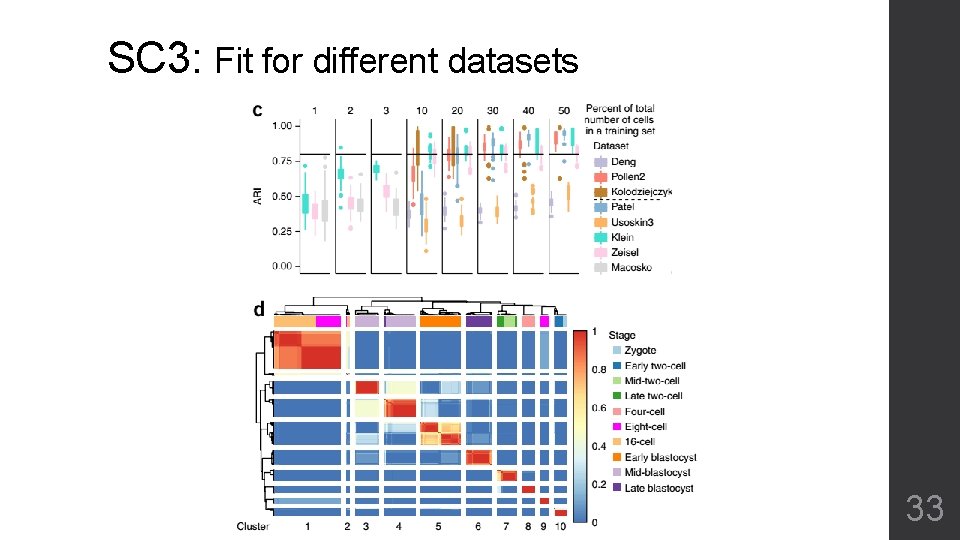
SC 3: Fit for different datasets 33
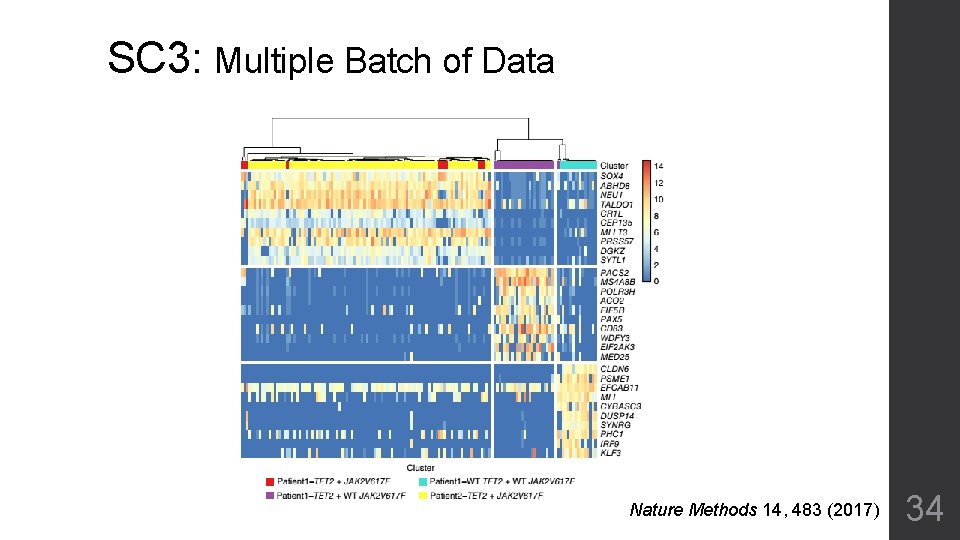
SC 3: Multiple Batch of Data Nature Methods 14, 483 (2017) 34

SC 3 • Stable, no many artificial parameters • Efficiency for huge datasets • Automatic predict cluster number • Limited considerable principle components (1~15) • Poor for subtly differentiated sub-types 35
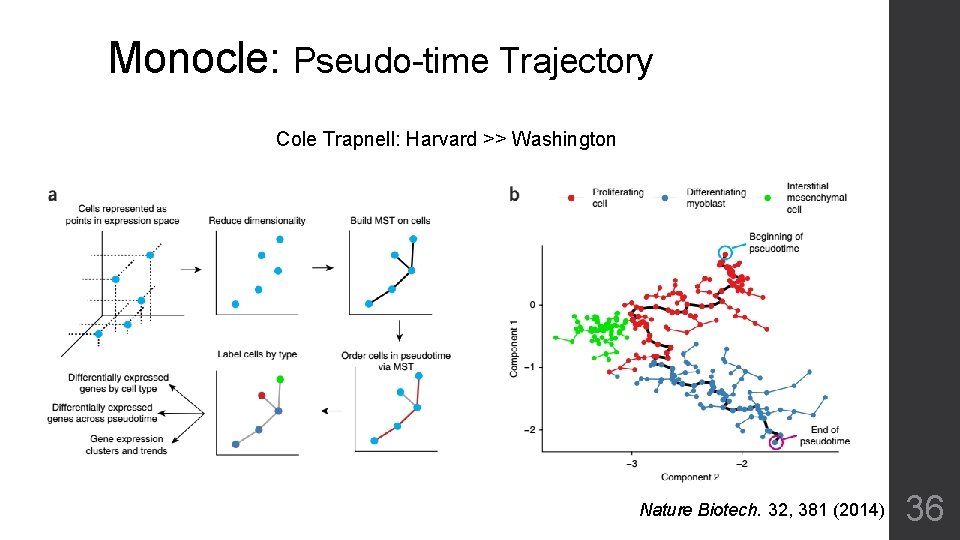
Monocle: Pseudo-time Trajectory Cole Trapnell: Harvard >> Washington Nature Biotech. 32, 381 (2014) 36
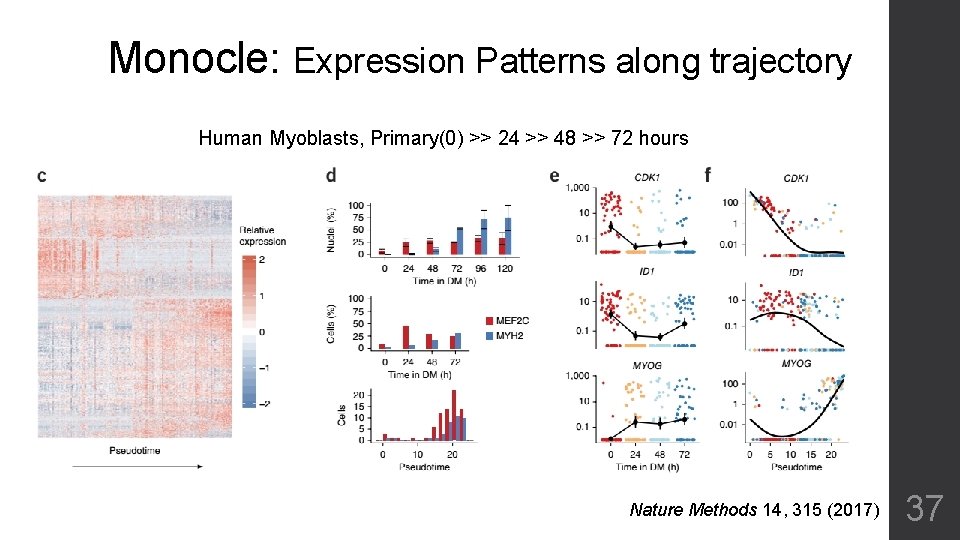
Monocle: Expression Patterns along trajectory Human Myoblasts, Primary(0) >> 24 >> 48 >> 72 hours Nature Methods 14, 315 (2017) 37
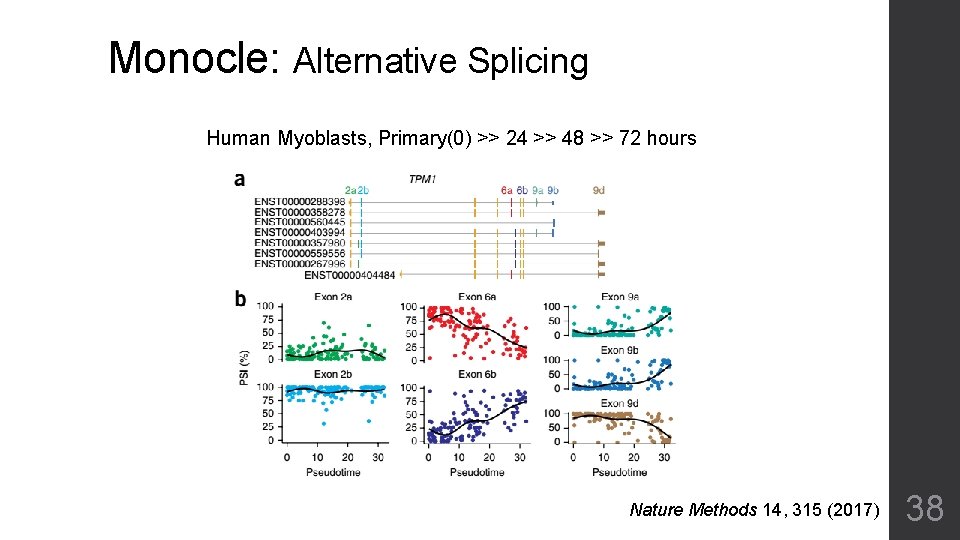
Monocle: Alternative Splicing Human Myoblasts, Primary(0) >> 24 >> 48 >> 72 hours Nature Methods 14, 315 (2017) 38
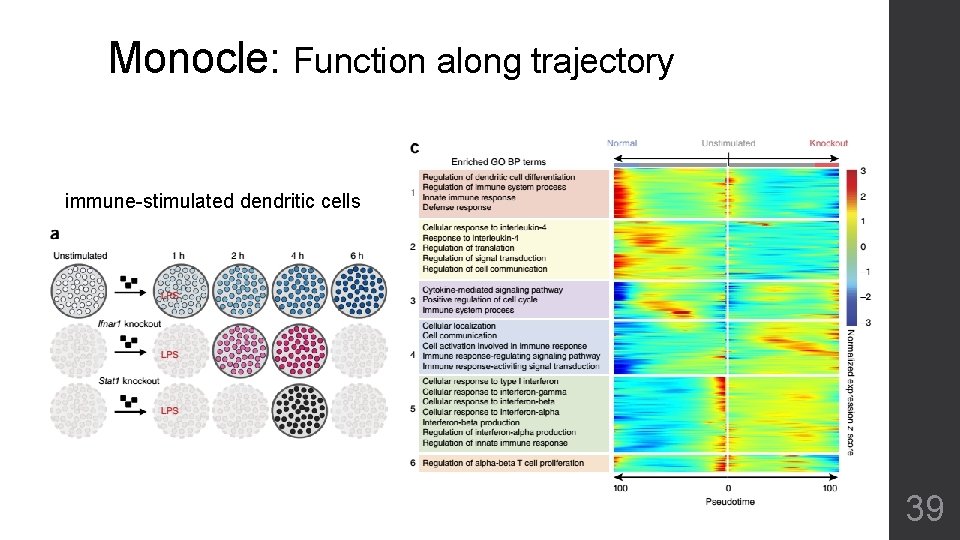
Monocle: Function along trajectory immune-stimulated dendritic cells 39

Monocle • Simulates time-dependent expression patterns • Predicts development/differentiation process • Blurring clustering of cell sub-types, using the algorithm same with Seurat • False positive prediction for time ordered cells 40

Single Cell ATAC-seq Analysis • More difficult: Even sparser than single cell RNA-seq data Unclear function for most signals • Chrom. VAR: Shrinks matrix by known motifs • Latent Semantic Indexing (LSI): Reduces dimension 41
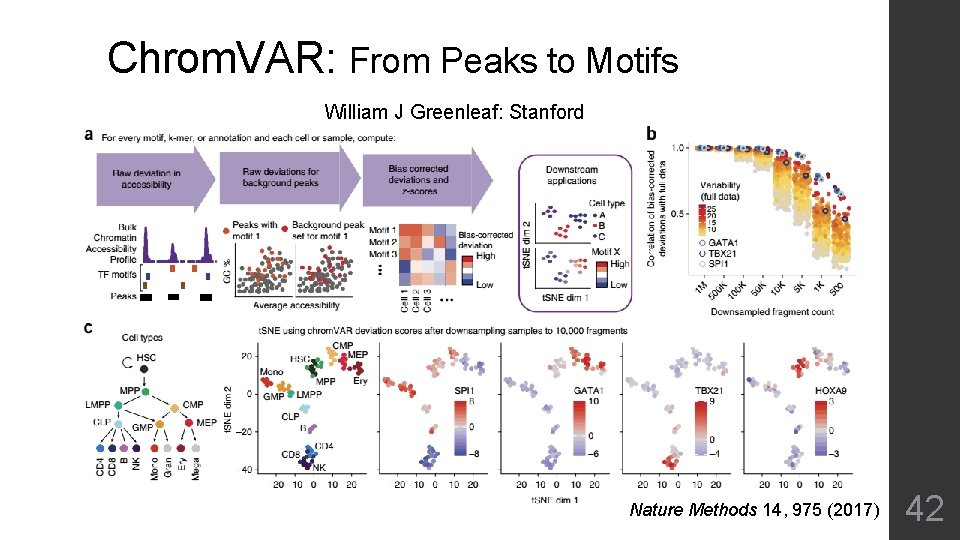
Chrom. VAR: From Peaks to Motifs William J Greenleaf: Stanford Nature Methods 14, 975 (2017) 42
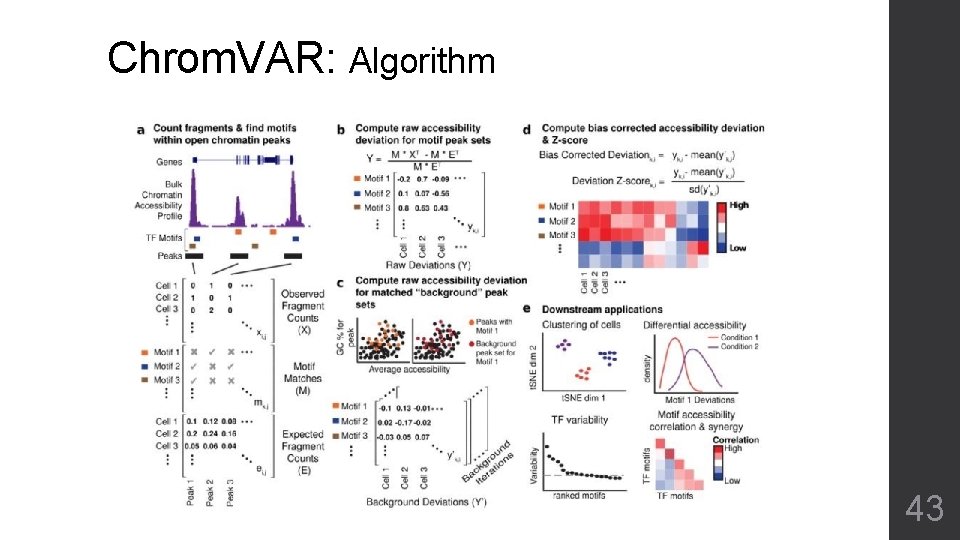
Chrom. VAR: Algorithm 43
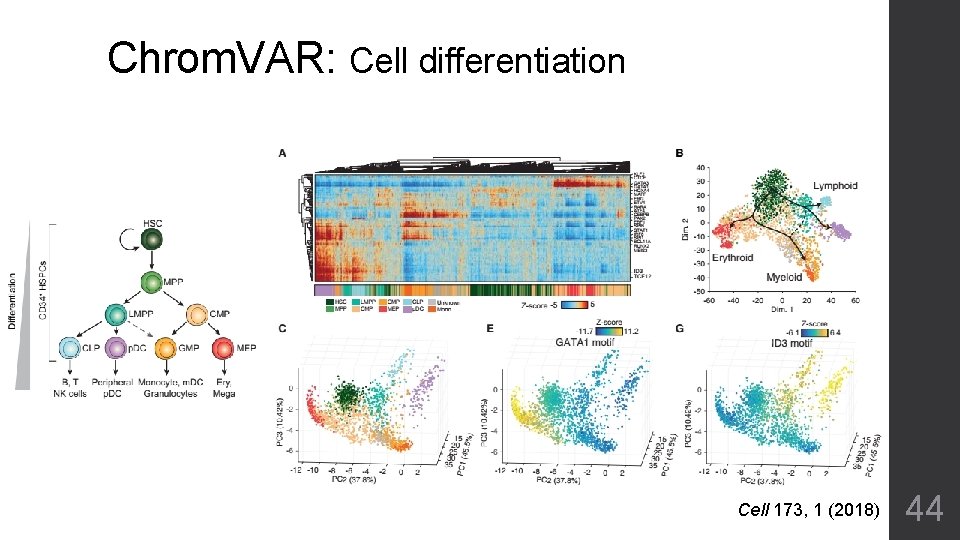
Chrom. VAR: Cell differentiation Cell 173, 1 (2018) 44
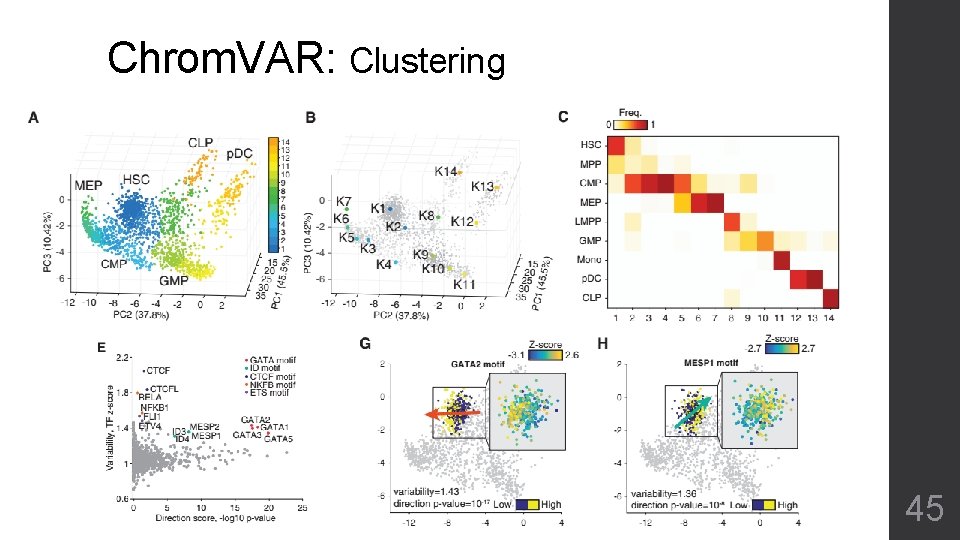
Chrom. VAR: Clustering 45
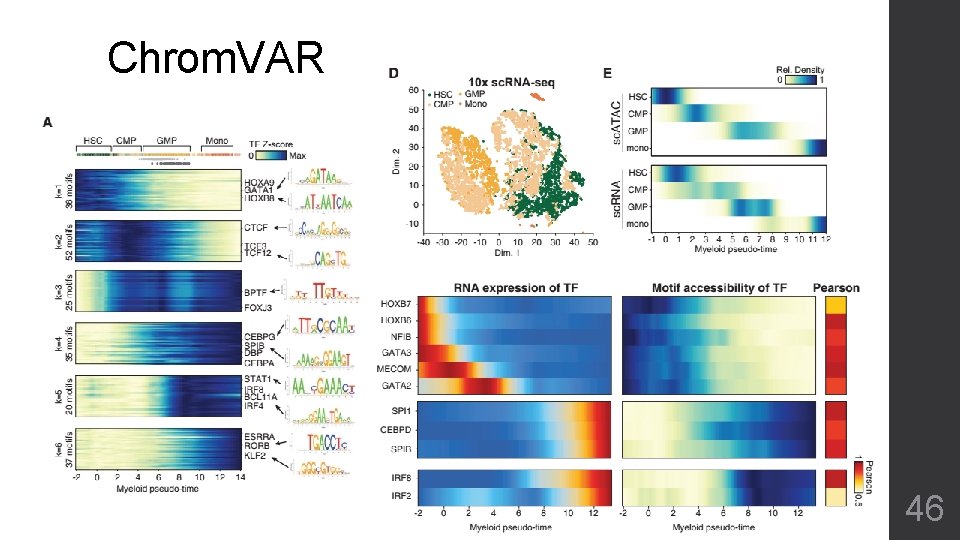
Chrom. VAR 46

Chrom. VAR • Clusters with Functional Motifs • Reveals development and/or linkage • Obscure clustering of sub-types • Ignore unknown bias of accessible regions except known motifs 47
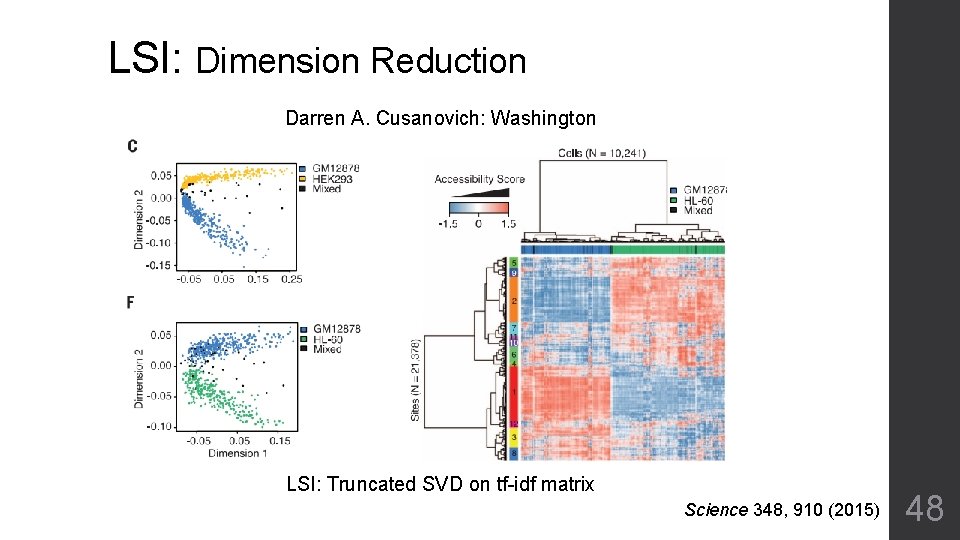
LSI: Dimension Reduction Darren A. Cusanovich: Washington LSI: Truncated SVD on tf-idf matrix Science 348, 910 (2015) 48

LSI: tf-idf matrix 49
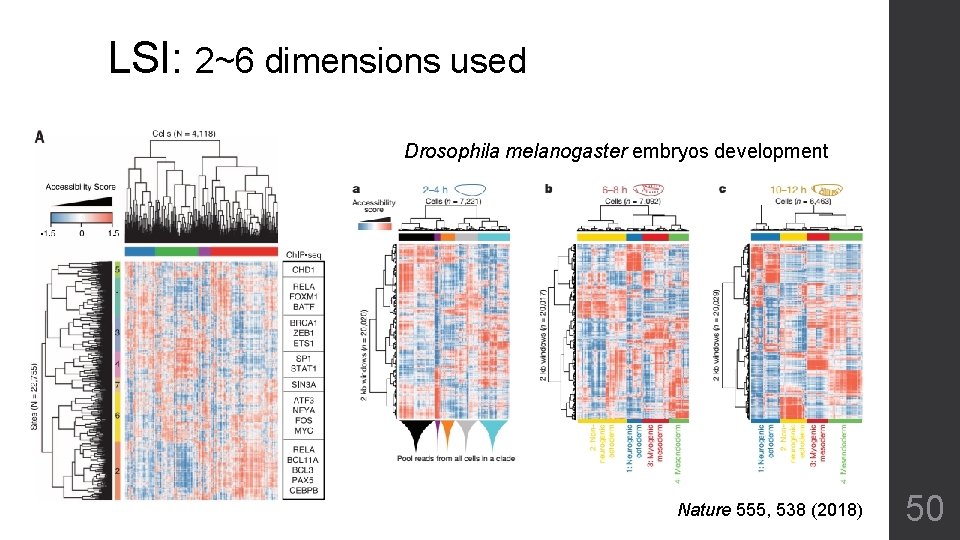
LSI: 2~6 dimensions used Drosophila melanogaster embryos development Nature 555, 538 (2018) 50
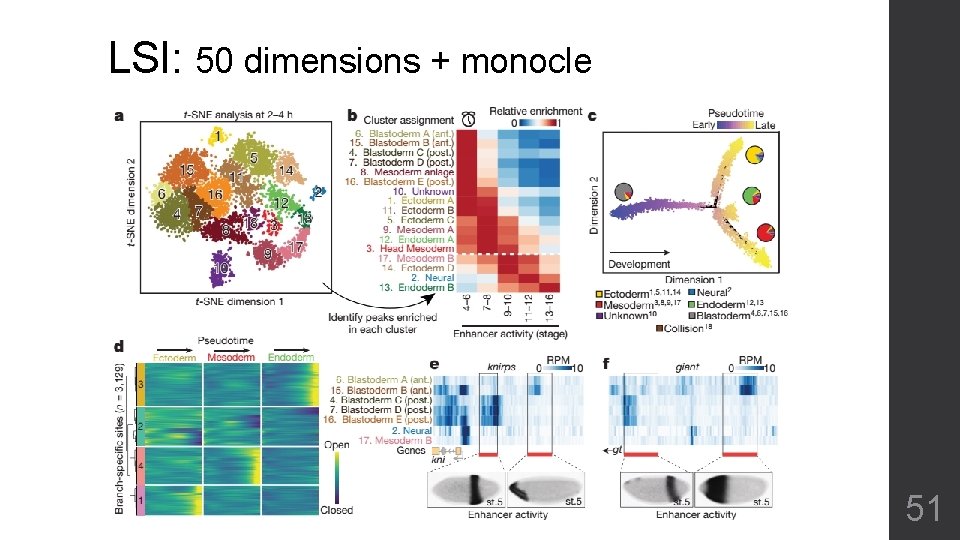
LSI: 50 dimensions + monocle 51

LSI • Convenient and cheap • Directly provide specific peaks • Very obscure clustering if only use LSI • Dangerous to dump first dimension • Not a proper replacement of PCA 52
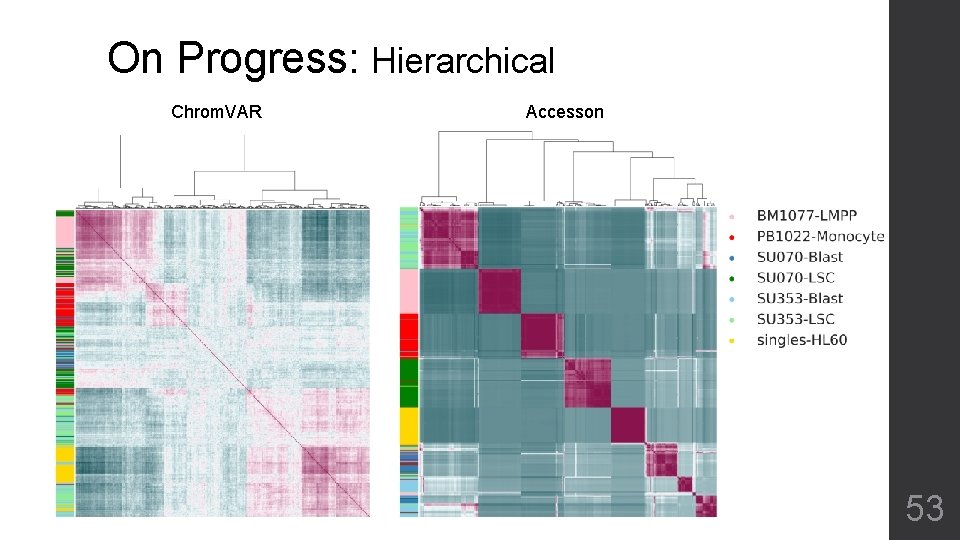
On Progress: Hierarchical Chrom. VAR Accesson 53
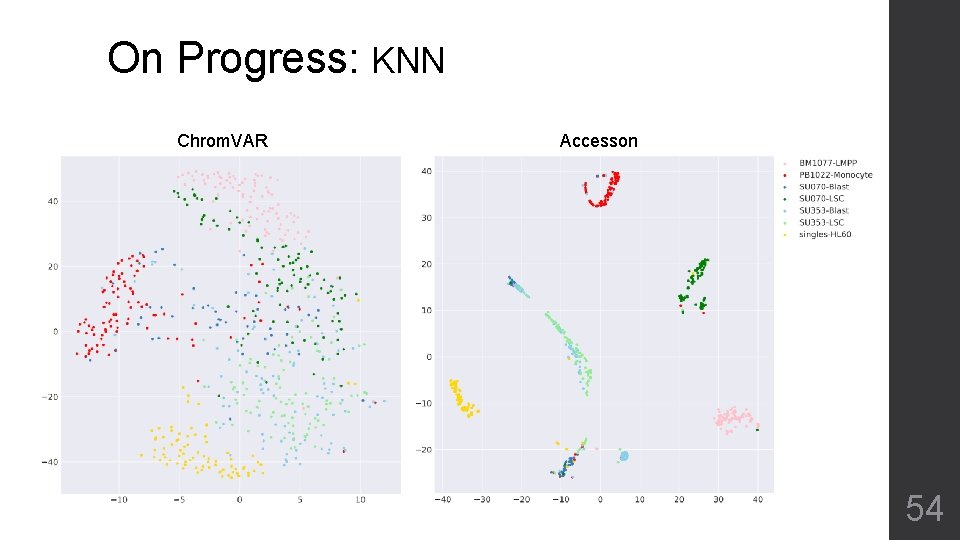
On Progress: KNN Chrom. VAR Accesson 54
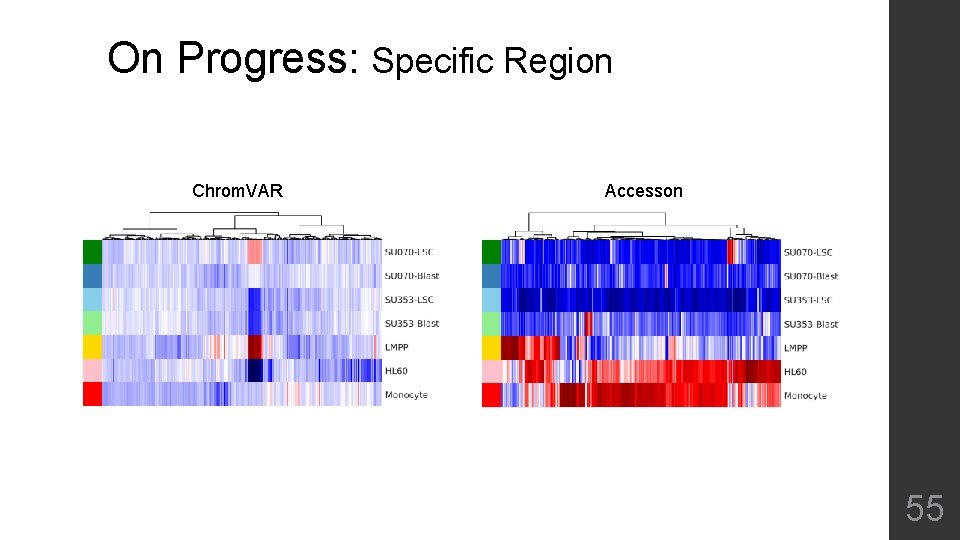
On Progress: Specific Region Chrom. VAR Accesson 55
- Slides: 55